Association between MATN-3 Gene Polymorphism and Primary Knee Osteoarthritis in Indian Population: A Community-Based Case-Control Study
Article Information
Vikas Trivedi, Ruchit Shah*, Shakeel Qidwai, A.N Mishra, Tasleem Raza, Sudhir Shyaam Kushwaha, Vipin Gupta, Afroz Khan, Nandan Mishra, Sharib Shamin, Abhishek Pandey, Robin Singh, Danish Khan, Priyank Sahoo, Mohd. Tabish
Department of Orthopaedics, Era’s medical college and hospital, Lucknow, Uttar Pradesh, India
*Corresponding Author: Ruchit Shah, Department of Orthopaedics, Era’s medical college and hospital, Lucknow, Uttar Pradesh, India
Received: 17 December 2019; Accepted: 02 January 2020; Published: 06 January 2020
Citation:
Vikas Trivedi, Ruchit Shah, Shakeel Qidwai, Mishra AN, Tasleem Raza, Sudhir Shyaam Kushwaha, Vipin Gupta, Afroz Khan, Nandan Mishra, Sharib Shamin, Abhishek Pandey, Robin Singh, Danish Khan, Priyank Sahoo, Mohd. Tabish. Association between MATN-3 Gene Polymorphism and Primary Knee Osteoarthritis in Indian Population: A Community-Based Case-Control Study. Fortune Journal of Rheumatology 2 (2020): 001-012.
View / Download Pdf Share at FacebookAbstract
Background and objectives: In the occurrence and development of primary knee osteoarthritis, the potential role of genetic factor has become an attraction for researches. Matrilin-3 (MATN-3), the newest identified gene in the pathomechanism of primary knee osteoarthritis, is a non-collagenous oligomeric extracellular matrix protein (ECM) and the smallest member of the matrilin family. This present study highlights the possible association of MATN-3 rs8176070 (SNP6) polymorphism with primary knee OA in the Indian population.
Material and Methods: In total, 50 patients with primary knee OA, aged between 40-75 years, enrolled in the case-control study conducted in North India. The other 50 demographically matched healthy individuals were considered as control groups. Determination of genotypes of MATN-3 SNP6 in cases as well as in controls was performed using polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP). Demographic details, medical history and other relevant information were obtained. The diagnosis was made by clinical examination and radiological grading using the Kellgren-Lawrence grade scale (KL).
Results: With regard to genotypes of MATN-3 gene SNP6 (rs8176070), 22 (78.57) patient with the B\b genotype had severe KL grades, yet no significant association compared with those B\B and b\b genotypes (p=0.485). Additionally, patients with b allele frequency had a severe KL grade, although no significant (p=0.180) association was found.
Conclusion: The verdicts obtained from the present study proposed that MATN-3 gene polymorphism was not associated with primary knee osteoarthritis in the Indian population.
Keywords
Matrilin-3, Primary Knee Osteoarthritis, Kellgren-Lawrence Grade, Restriction Fragment Length Polymorphism
Matrilin-3 articles, Primary Knee Osteoarthritis articles, Kellgren-Lawrence Grade articles, Restriction Fragment Length Polymorphism articles
Matrilin-3 articles Matrilin-3 Research articles Matrilin-3 review articles Matrilin-3 PubMed articles Matrilin-3 PubMed Central articles Matrilin-3 2023 articles Matrilin-3 2024 articles Matrilin-3 Scopus articles Matrilin-3 impact factor journals Matrilin-3 Scopus journals Matrilin-3 PubMed journals Matrilin-3 medical journals Matrilin-3 free journals Matrilin-3 best journals Matrilin-3 top journals Matrilin-3 free medical journals Matrilin-3 famous journals Matrilin-3 Google Scholar indexed journals Primary Knee Osteoarthritis articles Primary Knee Osteoarthritis Research articles Primary Knee Osteoarthritis review articles Primary Knee Osteoarthritis PubMed articles Primary Knee Osteoarthritis PubMed Central articles Primary Knee Osteoarthritis 2023 articles Primary Knee Osteoarthritis 2024 articles Primary Knee Osteoarthritis Scopus articles Primary Knee Osteoarthritis impact factor journals Primary Knee Osteoarthritis Scopus journals Primary Knee Osteoarthritis PubMed journals Primary Knee Osteoarthritis medical journals Primary Knee Osteoarthritis free journals Primary Knee Osteoarthritis best journals Primary Knee Osteoarthritis top journals Primary Knee Osteoarthritis free medical journals Primary Knee Osteoarthritis famous journals Primary Knee Osteoarthritis Google Scholar indexed journals Kellgren-Lawrence Grade articles Kellgren-Lawrence Grade Research articles Kellgren-Lawrence Grade review articles Kellgren-Lawrence Grade PubMed articles Kellgren-Lawrence Grade PubMed Central articles Kellgren-Lawrence Grade 2023 articles Kellgren-Lawrence Grade 2024 articles Kellgren-Lawrence Grade Scopus articles Kellgren-Lawrence Grade impact factor journals Kellgren-Lawrence Grade Scopus journals Kellgren-Lawrence Grade PubMed journals Kellgren-Lawrence Grade medical journals Kellgren-Lawrence Grade free journals Kellgren-Lawrence Grade best journals Kellgren-Lawrence Grade top journals Kellgren-Lawrence Grade free medical journals Kellgren-Lawrence Grade famous journals Kellgren-Lawrence Grade Google Scholar indexed journals Restriction Fragment Length Polymorphism articles Restriction Fragment Length Polymorphism Research articles Restriction Fragment Length Polymorphism review articles Restriction Fragment Length Polymorphism PubMed articles Restriction Fragment Length Polymorphism PubMed Central articles Restriction Fragment Length Polymorphism 2023 articles Restriction Fragment Length Polymorphism 2024 articles Restriction Fragment Length Polymorphism Scopus articles Restriction Fragment Length Polymorphism impact factor journals Restriction Fragment Length Polymorphism Scopus journals Restriction Fragment Length Polymorphism PubMed journals Restriction Fragment Length Polymorphism medical journals Restriction Fragment Length Polymorphism free journals Restriction Fragment Length Polymorphism best journals Restriction Fragment Length Polymorphism top journals Restriction Fragment Length Polymorphism free medical journals Restriction Fragment Length Polymorphism famous journals Restriction Fragment Length Polymorphism Google Scholar indexed journals Osteoarthritis articles Osteoarthritis Research articles Osteoarthritis review articles Osteoarthritis PubMed articles Osteoarthritis PubMed Central articles Osteoarthritis 2023 articles Osteoarthritis 2024 articles Osteoarthritis Scopus articles Osteoarthritis impact factor journals Osteoarthritis Scopus journals Osteoarthritis PubMed journals Osteoarthritis medical journals Osteoarthritis free journals Osteoarthritis best journals Osteoarthritis top journals Osteoarthritis free medical journals Osteoarthritis famous journals Osteoarthritis Google Scholar indexed journals etiology articles etiology Research articles etiology review articles etiology PubMed articles etiology PubMed Central articles etiology 2023 articles etiology 2024 articles etiology Scopus articles etiology impact factor journals etiology Scopus journals etiology PubMed journals etiology medical journals etiology free journals etiology best journals etiology top journals etiology free medical journals etiology famous journals etiology Google Scholar indexed journals mechanical misalignment articles mechanical misalignment Research articles mechanical misalignment review articles mechanical misalignment PubMed articles mechanical misalignment PubMed Central articles mechanical misalignment 2023 articles mechanical misalignment 2024 articles mechanical misalignment Scopus articles mechanical misalignment impact factor journals mechanical misalignment Scopus journals mechanical misalignment PubMed journals mechanical misalignment medical journals mechanical misalignment free journals mechanical misalignment best journals mechanical misalignment top journals mechanical misalignment free medical journals mechanical misalignment famous journals mechanical misalignment Google Scholar indexed journals non-collagenous oligomeric articles non-collagenous oligomeric Research articles non-collagenous oligomeric review articles non-collagenous oligomeric PubMed articles non-collagenous oligomeric PubMed Central articles non-collagenous oligomeric 2023 articles non-collagenous oligomeric 2024 articles non-collagenous oligomeric Scopus articles non-collagenous oligomeric impact factor journals non-collagenous oligomeric Scopus journals non-collagenous oligomeric PubMed journals non-collagenous oligomeric medical journals non-collagenous oligomeric free journals non-collagenous oligomeric best journals non-collagenous oligomeric top journals non-collagenous oligomeric free medical journals non-collagenous oligomeric famous journals non-collagenous oligomeric Google Scholar indexed journals Orthopaedics articles Orthopaedics Research articles Orthopaedics review articles Orthopaedics PubMed articles Orthopaedics PubMed Central articles Orthopaedics 2023 articles Orthopaedics 2024 articles Orthopaedics Scopus articles Orthopaedics impact factor journals Orthopaedics Scopus journals Orthopaedics PubMed journals Orthopaedics medical journals Orthopaedics free journals Orthopaedics best journals Orthopaedics top journals Orthopaedics free medical journals Orthopaedics famous journals Orthopaedics Google Scholar indexed journals metabolic derangements articles metabolic derangements Research articles metabolic derangements review articles metabolic derangements PubMed articles metabolic derangements PubMed Central articles metabolic derangements 2023 articles metabolic derangements 2024 articles metabolic derangements Scopus articles metabolic derangements impact factor journals metabolic derangements Scopus journals metabolic derangements PubMed journals metabolic derangements medical journals metabolic derangements free journals metabolic derangements best journals metabolic derangements top journals metabolic derangements free medical journals metabolic derangements famous journals metabolic derangements Google Scholar indexed journals
Article Details
Abbreviation:
OA-Osteoarthritis; MATN-3-Matrilin-3; ECM-Extracellular matrix protein; KL-Kellgren-Lawrence system; SNP-Single nucleotide polymorphism; PCR-RFLP-Polymerase chain reaction-restriction fragment length polymorphism
1. Introduction
Based on etiology, primary (idiopathic or non-traumatic) and secondary (usually due to trauma or mechanical misalignment) are two main types of osteoarthritis. Primary knee osteoarthritis (OA) is the second most common rheumatologic problem and one of the leading causes of disability. This degenerative and progressive joint disease affects nearly 250 million people worldwide [1] and 22% to 39% in India [2]. Clinically, joint pain, limitation of movement, tenderness, crepitus, occasional effusion, and variable degrees of local inflammation without systemic effects are the salient symptoms associated with primary knee OA [3, 4]. The severity of the disease can also be graded as per the radiographical findings by the Kellgren-Lawrence (KL) system, which was proposed by Kellgren et al. in 1957 [5] and later on accepted by WHO in 1961.
Primary knee OA was historically viewed as exclusively a degenerative disorder of the cartilage despite, recent evidence has shown that it is a multifactorial entity, involving multiple causative factors like trauma, mechanical forces, inflammation, biochemical reactions, and metabolic derangements [6]. Genetic factors come to light as affecting the risk of primary knee OA development [7]. Keeping this in mind, several genome-wide association studies (GWAS) have attempted to identify OA susceptibility genes and showed single-nucleotide polymorphisms (SNPs) in multiple genes associated with an increased risk of primary knee OA, including collagen genes (COL1A1, COL2A1, COL9A1, and COL11A2) as well as genes encoding interleukin-1 receptor (IL1R), transforming growth factor-β1 (TGFβ1), cartilage matrix protein1 (CMP1), tissue inhibitor of metalloproteinase 3 (TIMP3), insulin-like growth factor-1 (IGF1), bone morphogenetic protein (BMP), vitamin D receptor (VDR), aggrecan-1 (AGC1), secreted frizzled-related protein 3 (FRZB), growth differentiation factor 5 (GDF5), and cyclooxygenase-2 (COX-2) [8-10]. It is assumed that the genetic changes affecting the synthesis of cartilage may play a role in the development of primary knee OA.
Recently, Matrilin (MATNs) gene is identified as the newest causative agent for primary knee OA. The matrilin (MATNs) are a family of oligomeric extracellular matrix (ECM) proteins that have four members termed matrilin-1 through -4 consisting of von Willebrand factor A (vWFA) domains, epidermal growth factor (EGF) like domains, and a C-terminal coiled-coil domain [11]. Matrilin-3 (MATN-3) is the least complex member of the matrilin family, consisting of only one vWFA domain, four EGF-like domains, and a C-terminal coiled-coil domain and interacts with collagen fibrils, multiple proteoglycans, and other glycoproteins; thus, it plays a dominant role in the formation of a filamentous matrix network. It is encoded by the MATN-3 gene, which is present on the short arm of the chromosome 2 region 2p24-p23 [12]. The regulation of MATN-3 expression is essential for the maintenance of the cartilage extracellular matrix (ECM) microenvironment [13].
In the pathogenesis of many disorders, including hereditary microepiphyseal dysplasia and spondyloepimetaphyseal dysplasia, and predispose individuals to develop primary knee OA, MATN-3 gene mutations have been pointed out [14-16]. Many recently published studies in the past few years have been demonstrated that single nucleotide polymorphism (SNP) of MATN-3 gene is associated with primary knee OA in patients with different ethnicities, although the results were controversial [13, 17-19]. However, there are no reported studies that assess the association between MATN-3 gene polymorphism and primary knee OA in Indian patients so far. In the present case-control study, we only considered a subtype of OA, primary knee OA, and aimed to investigate the possible association and clinical significance of MATN-3 rs8176070 (SNP6) polymorphism in a series of Indian patients with primary knee OA.
2. Materials And Methods
2.1 Study participants
The study was conducted in the Department of Orthopedics with the collaboration of the Department of Biotechnology, Era’s Medical College and Hospital, Lucknow, Uttar Pradesh, India. It was conducted in accordance with the guidelines of the Declaration of Helsinki (World Medical Association, 2008), and the local ethics committee of the institute approved the study protocol. Informed written consent was obtained from all the participants before they were enrolled in the study. In the present case-control study, a total of 50 patients (Group I)with clinical symptoms along with radiographic signs of primary knee OA according to theKL grading, who attended or were admitted to the outpatients’ clinic and inpatients’ department of Era’s Lucknow Medical College and Hospital, from November 2017 to October 2019, together with 50 demographically matched healthy individuals (Group II) with no symptoms or signs on clinical examination or radiographic changes indicative of primary knee OA or other joint diseases were included.
2.2 Sample size estimation
Sample size estimation was done at the Department of Community Medicine, Era’s Medical College and Hospital. Itwas calculated based on the proportion of B allele of MATN-3 gene among cases and controls [20], using the following formula:
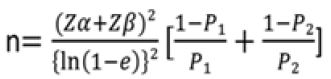
Where, p1= 0.46, the proportion of B allele in cases; p2= 0.68, the proportion of B allele in controls; error ratio e=0.35, considered to be clinically significant; type 1 errorα= 5%; type II errorβ= 20% for detecting results with the 80% power ofstudy; data loss Z= 10%.
By putting these values in the equation, the sample size found to be n=50 for each group.
2.3 Inclusion criteria and exclusion criteria
Both cases and controls participants, with age group of 40-75 years, regardless of any gender bias were included in the study. Patients were completely excluded from the study if they had one or more of the following conditions: 1) isolated patellofemoral OA; 2) secondary OA due to trauma or deformity; 3) metabolic disorders such as diabetes mellitus, thyroid disorders, or Cushing syndrome; or renal failure or were undergoing dialysis; 4) infectious disorders such as septic arthritis, viral arthritis, or fungal arthritis; 5) inflammatory arthritis such as rheumatoid arthritis, systemic lupus erythematosus, and seronegative arthritis; 6) malignancy; 7) bilateral knee replacements.
2.4 Data collection
Detailed histories of demographic and other information including age, gender, occupation, medical history were obtained by using questionnaires. Only in cases, some blood investigations (CRP, RA factor) and X-ray of the bilateral knee in anteroposterior and lateral view on standing were also performed followed by graded according to theKL grading system [5].
2.5 Kellgrenand Lawrence (KL) grading system
Grade 0: No radiographic features of primary knee OA are present.
Grade 1: Doubtful joint space narrowing and possible osteophytic lipping.
Grade 2: Definite osteophytes and possible joint space narrowing on the anteroposterior weight
bearing radiograph.
Grade 3: Multipleosteophytes,definite joint space narrowing, sclerosis, possible bone deformity.
Grade 4: Largeosteophytes,marked joint space narrowing, severe sclerosis and definite bony deformity.
2.6 Procedure for DNA extraction
In total, 5 ml of venous blood was drawn and poured in EDTA containing tubes. The DNA samples will be isolated from peripheral blood lymphocytes by the standard phenol extraction method using the Thermo Scientific Gene JET Whole Blood Genomic DNA Purification Mini Kit (Thermo Fisher Scientific Inc, California, USA). The DNA concentration will be determined by the spectrometer and stored at -20degree Celsius till the time of DNA extraction.
2.7 MATN-3 genotyping
Polymerase chain reaction (PCR) was conducted in a reaction volume of 50 μL using Dream Taq Green PCR Master Mix (Thermo Fisher Scientific Inc, California, USA) with 25 μL dream taq green PCR master mix, 3 μL (5 pmol ) forward primer, 3 μL (5 pmol) reverse primer, 5 μL (10 ng) template DNA, 14 μL nuclease-free water. The primers used for PCR were: 5′d GGACAGGATCCCACAAAAAG 3′ as a forward primer and 5′d GAAAGAGGGGCTACAACAGG 3′ as a reverse primer. Here mentioned amplification protocol was used: initial denaturation at 95°C for 10 min, followed by 35 cycles of denaturation at 95°C for 1 min, annealing at 60°C for 1 min, extending at 72°C for 1 min, and final extension at 72°C for 10 min in a thermocycler. The resulting PCR products demonstrated a single fragment at 501 bp by gel electrophoresis. With 10 units of BSEYI restriction enzyme at 37°C for 10 h, in total, 10μLof the 501-bp product was digested. A 3% agarose gel containing 2% ethidium bromide was used for the visualization of digestion products. Restriction fragment length polymorphisms (RFLPs) were coded as Bb, where the uppercase letter indicated the absence of the restriction site and the lowercase letter implied the presence of the site. The wild-type genotype (CC), was coded as bb, which produced a double band at 149 and 352 bp; heterozygotes (CN), was coded as Bb, which produced three bands at 501, 149, and 352 bp; and homozygote polymorphic genotype (NN), was coded as BB, which produced only one band at 501 bp.Results of PCR-RFLP for MATN-3 gene are shown in Figure 1.
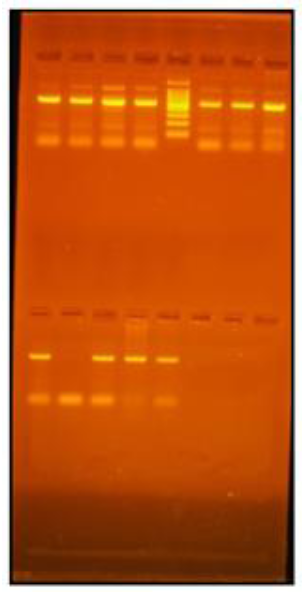
Figure 1: PCR-RFLP results of matrilin 3gene done on 3% agarose gel, Lane#6-100 bp molecular weight marker, Lane# 1,3,4,5,8 (Bb), Lane#2,7 (bb).
2.8 Statistical analysis
Statistical analyses were performed withStatistical Package for Social Sciences version 21.0. Software (SPSS Inc., Chicago, IL, USA). Calculation of Chi-square test for deviation from Hardy-Weinberg equilibrium (HWE) was performed using the online freely available calculator to compare the observed and expected frequencies of genotypes in the studied groups.
21Quantitative data were presented asmean ± standarddeviation, whereas, categorical data were shown as frequency (percentage). Unpaired T-test was used to compare continuous parametric variables, whileChi-squaretest was used for categoricalvariable
comparison.The differences in genotype and allele distributions were investigated by the chi-square (χ2) test and, represented by odds ratios (ORs) and 95% confidence intervals (CIs). In statistical comparison between two groups, the statistical significance was defined as p<0.05 (*), while a p-value of >0.05 was insignificant. A p-value of <0.001 was regarded as very highly significant (***).
3. Results
Out of the 100 subjects enrolled in the study, a total of 50 (50%) were patients of primary knee OA considered as the case group of study, while remaining 50 (50%) were demographically matched healthy individuals served as the control group of the study. Analysis of HWE showed that genotypes of the MATN-3 gene SNP6 (rs8 176070) in both cases and control subjects were not in HWE. There was evidence to reject the assumption of HWE in the sample (p<0.05 for each).
|
Characteristics |
Group I: Cases (n=50) |
Group II: Control (n=50) |
p-value |
|
Age/years, Mean ± SD (range)# |
55.90 ± 7.04 (45-70) |
56.00 ± 7.34 (42-72) |
0.945 |
|
Age groups, n (%) |
|||
|
≤50 Years |
16 (32) |
13 (26) |
0.102 |
|
51-60 Years |
21 (42) |
26 (52) |
|
|
61-70 Years |
13 (26) |
8 (16) |
|
|
71-80 Years |
- |
3 (6) |
|
|
Gender, n (%) |
|||
|
Male |
30 (60) |
31 (62) |
0.838 |
|
Female |
20 (40) |
19 (38) |
|
|
Occupation, n (%) |
|||
|
Unskilled worker |
7 (14) |
15 (30) |
0.314 |
|
Skilled worker |
7 (14) |
6 (12) |
|
|
Service |
4 (8) |
2 (4) |
|
|
Business/ Profession |
12 (24) |
13 (26) |
|
|
Housewife |
20 (40) |
14 (28) |
|
|
Medical history, n (%) |
|||
|
Diabetes |
- |
- |
0.806 |
|
Hypertension |
11 (22) |
10 (20) |
|
|
Other systemic illnesses |
- |
- |
|
¥ SD, Standard Deviation. #Unpaired T-test was used to compare continuous parametric variables. Chi-square test was used for the categorical variable comparison. p>0.05 was statistically insignificant.
Table 1: Demographic characteristics of the studied groups.
|
Genotype, n(%) |
Group I: Cases |
Group II: Control |
p-value |
OR (95% CI) |
|
n=50 |
n=50 |
|||
|
bb |
10(20) |
10(20) |
- |
- |
|
Bb |
40(80) |
39(78) |
0.807 |
1.128(0.430-2.956) |
|
BB |
- |
1(2) |
0.317 |
0.333(0.013-8.379) |
|
Allele, n(%) |
n=100 |
n=100 |
||
|
b |
60(60) |
59(59) |
0.886 |
1.042(0.593-1.833) |
|
B |
40(40) |
41(41) |
0.886 |
0.959(0.545-1.687) |
Table 2: Comparison of MATN-3 gene SNP6 (rs8176070) genotypes and allele frequencies between case and control groups.
|
KL Grade |
Total No. |
bb, n(%) |
Bb, n(%) |
BB, n(%) |
Statistics |
|
Mild (I-II) Severe (III-IV) Grade I Grade II Grade III Grade IV |
22 28 6 16 18 10 |
4 (18.18) 6 (21.43) - 4 (25.00) 4 (22.22) 2 (20.00) |
18 (81.82) 22 (78.57) 6 (100.00) 12 (75) 14 (77.78) 8 (80.00) |
- - - - - - |
X2=0.08, p=0.485 X2=1.81, p=0.614 |
¥ p>0.05 was statistically insignificant.
Table 3: Association of genotype with KL Grade in cases (n=50).
|
KL Grade |
Total No. |
b, n(%) |
B, n(%) |
Statistics |
|
Mild (I-II) Severe (III-IV) Grade I Grade II Grade III Grade IV |
44 56 12 32 36 20 |
20 (45.45) 34 (60.71) - 20 (62.50) 22 (61.11) 12 (60) |
24 (54.55) 22 (39.29) 12 (100) 12 (37.50) 14 (38.89) 8 (40.0) |
X2=1.80, p=0.180 X2=16.04, p=0.001*** |
¥ p<0.001 was considered highly significant***
Table 4: Association of allele frequency with KL Grade in cases (n=100).
|
Author, year |
Findings |
|
Belluoccio et al.,1998 [23] |
Matrix MATN-3 molecule was expressed in every cartilage type and it is a beneficial indicator of the state of differentiation of cells obtained from the articular cartilage. |
|
Pullig et al., 2002 [24] and Pullig et al., 2007 [25] |
MATN-3 is a fundamental component of human articular cartilage matrix, which is remarkably up-regulated in OA cartilage, may be a cellular response to the modified microenvironment in this condition. Its expression levels are associated with disease severity. |
|
Min et al., 2006 [26] |
Polymorphism in MATN-3 associated with risk of thumb carpometacarpal (CMC1) OA. |
|
Van der Weyden et al., 2006 [27] |
The preclinical study revealed that the MATN-3 deletion in mice causes an imbalance in the maintenance of structural integrity with increased inflammation and degradation of ECM components. As a result, MATN-3 null mice are predisposed to the initiation and progression of OA. This way, MATN-3 plays a dominant role in maintaining the structural integrity of cartilage ECM. Apart from this, MATN-3 null mice also display an increased bone mineral density. |
|
Vincourt et al., 2008 [28] |
In the structure of MATN-3, the first EGF-like domain is a decisive determinant of its regulatory activity toward SOX9. The actions of MATN-3 propose that its enhanced expression in OA might contribute to the degeneration of articular cartilage. |
|
Klatt et al., 2009b [29] |
The role of MATN-3 as a matrix degradation regulator in human cartilage is crucial and potent MATN-3-dependent induction of the pro-inflammatory cytokines TNFa, IL-1b, IL-6, and IL-8, which are up-regulated in OA. These pro-inflammatory cytokines boost the process of inflammation and induce different matrix-degrading enzymes, are therefore vital regulators of cartilage degradation. Apart from these attributes, MATN-3 induces iNOS and COX-2 gene; both are involved in the pathogenesis of OA, in cartilage. At last, MATN-3 induces the expression of MMP1, MMP3, and MMP13 in PHCs, which implies that MATN-3 acts as an intermediary and regulate ECM degradation. |
|
Vincourt et al., 2008 [28] and Klatt et al., 2009b [29] |
In vitro characterization of MATN-3identified anti-anabolic and pro-catabolic functions, suggesting that it may actively participate in OA by an unknown mechanism. |
|
Vincourt et al.,2012 [30] |
Revealed higher levels of MATN-3 in both the serum and synovial fluid of OA patients. |
|
Jayasuriya et al.,2012 [31] |
MATN-3 plays not only a structural role in cartilage ECM, but also a regulatory role in cartilage homeostasis due to its capacity to induce IL-1Ra, to up regulate gene expression of the major cartilage matrix components, and to down regulate the expression of OA-associated matrix-degrading proteinases in chondrocytes. |
|
Minafra et al.,2014 [32] |
Potential statistically significant associations of polymorphisms in the foremost OA susceptibility genes including, GDF5 rs143383 & DVWA rs11718863 genes and KL grading was hypothesized. The DVWA rs11718863 gene had a predictive role in OA progression, as associated with a more severe radiographic grade. |
Table 5: Historical evidence of clinical studies claimed the role of MATN-3 gene in osteoarthritis.
4. Discussion
Rising prevalence of primary knee OA in India, compromised lifestyle of affected individuals due to morbidity, polymorphisms and mutations of multiple genes involved in the pathogenesis of primary knee OA and a little while back attraction on new identified MATN-3 gene as a predisposing factor in primary knee OA inspired us to carry out this research. MATN-3 is a non-collagenous oligomeric ECM mainly involved in cartilage development, ossification and potential osteoarthritis pathomechanisms. Moreover, it interacts directly or indirectly with transforming growth factor β (TGF-β), and bone morphogenetic protein 2 (BMP2) and, ultimately regulates chondrocyte proliferation and hypertrophic differentiation. Interestingly, MATN-3 raises interleukin receptor antagonists (IL-Ra) in chondrocytes, indicating its role in suppression of inflammatory activity mediated by IL-1β. It also down regulates the expression of matrix-degrading enzymes, such as a disintegrin metalloproteinase with thrombospondin motifs (ADAMTS 4 and ADAMTS 5), matrix metalloproteinase 13 (MMP13), and collagen X, a hypertrophy marker during growth and inflammatory conditions. Expression of collagen II and aggrecan is significantly improved by MATN-3, which is needed to maintain the tensile strength and elasticity of cartilage, respectively. Apart from these attributes, MATN-3 also induces OA-associated markers in chondrocytes in a concentration-dependent manner is also blamed for pathogenesis. In summary, MATN-3 plays a dominant role in inflammation, matrix degradation, and matrix formation in cartilage development and at last, in OA occurrence [22]. Historical evidence in favour of the role of MATN-3 in OA was summarized in Table V and, in a simple and straightway, they proved that MATN-3 is a cornerstone in the skeletal development and its mutation predisposes individuals to develop OA [23-32].
It is already well known that any disorder occurs due to genetic aetiology may lead to ethnic or racial differences in health. To date, two reported studies observed an association between MATN-3 gene polymorphism and OA in different ethnicity group [13, 33]. The most interesting part is to compare demographic features and obtained findings of these studies that may clinch the clue regarding the fluctuating role of MATN-3 gene in the development of OA with an ethnic variation. Gu et al [13] studied on 732 OA cases with a mean age of 57.4± 9.5 years from china revealed that the B\B genotype increased the risk of OA (OR=1.724, 95% CI=1.071-2.770; p=0.025) especially knee OA. Additionally, they stated B allele might have associated with increased knee OA (OR=3.143, 95% CI=2.283-4.328; p=0.000) compared with control subjects.
A little while back in 2017 the published report by Diab et al [33] postulated that prevalence of B\b genotype (p=0.046) and b allele (p= 0.042) was more evident in OA patients with a high risk of OA development. Patients with the B\b genotype had worse clinical and radiological findings than those with B\B and b\b genotypes. This dilemma regarding outcome variation may be explained by ethnic differences related to geographic distribution. Among Indian patients with the primary knee OA, possible association and clinical significance of MATN-3 gene rs8176070 (SNP6) were investigated in the present study. Regarding genotype (bb, Bb and BB) and allele (b and B) distribution of MATN-3 gene, similarities were found between primary knee OA groups and control groups in the current study. Procured data from this study concluded that none genotype and allele frequency were contributing to primary knee OA development. Notably, genotype and allele frequency were not significantly associated with the severity of the disease in the Indian population.
In the present study, HWE was estimated in our genotypes for both cases and controls. According to HWE theorem, allele and genotype frequencies in a population will remain constant from generation to generation in the absence of other evolutionary influences include genetic drift, mate choice, assortative mating, natural selection, sexual selection, mutation, gene flow, meiotic drive, genetic hitchhiking, population bottleneck, founder effect and inbreeding. In genetic association studies, HWE evaluation is an essential parameter which is usually used to detect genotyping errors. Previously HWE was estimated only in the control group but this trend has been changed now. Recently, a likelihood ratio test for testing HWE in the study population using both case and control samples were widely accepted [34]. HWE findings from our data revealed there was a deviation in HWE in both case and control groups. One of the reasons behind this is the small sample size involved in the study. And another reason may be already the existence of MATN-3 gene mutation in the Indian population, which is supported by our findings that we did not obtain any association of MATN-3 with primary knee OA.
There are some drawbacks to be addressed in the present study. Firstly, our sample size was comparatively small, that may restrict statistical power to detect the possible association between the MATN-3 gene and primary knee OA. Secondly, here we have only focused on primary Knee OA, as we can't predict outcomes for other subtypes. Thirdly, the restricted sample size may result in the distribution of genotype and allele might be deviated from the actual condition. Furthermore, all the patients were from India's north-eastern region, as obtained results might not represent the entire situation of India. At last, deviation in HWE findings may also interfere with the final conclusion.
As concluded, in the Indian population, the examined polymorphism in the MATN-3 gene may not illuminate the risk and severity of knee OA. For more conclusive statement for association of MATN-3 gene polymorphism in primary knee OA patients, we recommended future studies with large sample size in the Indian population. If possible, the involvement of other subtypes of OA can be helpful to support the final justification and also assist to observe type related variation in the existing association. A study involving multiple ethnic group and comparison of association between them may become the most exciting matter for further research.
Acknowledgement
We are wholeheartedly thankful to patients who participate in the study. We are also grateful to Mrs Feral Daruwala to support in manuscript writing work.
Sources of Funding
Authors report no source of funding.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Lim SS, Vos T, Flaxman AD, et al. A comparative risk assessment of burden of disease and injury attributable to 67 risk factors and risk factor clusters in 21 regions, 1990-2010: a systematic analysis for the Global Burden of Disease Study 2010. Lancet 380 (2012): 2224-2260.
- Pal CP, Singh P, Chaturvedi S, et al. Epidemiology of knee osteoarthritis in India and related factors. Indian J Orthop 50 (2016): 518-522.
- Molloy ES, McCarthy GM. Eicosanoids, osteoarthritis, and crystal deposition diseases. Curr Opin Rheumatol 17 (2005): 346-350.
- Hawley DJ. Psycho-educational interventions in the treatment of arthritis. Bailliere’s Clinical Rheumatology 9 (1995): 803-823.
- Kellgren JH, Lawrence JS. Radiological assessment of osteoarthrosis. Ann Rheum Dis 16 (1957): 494-502.
- Ayhan E, Kesmezacar H, Akgun I. Intraarticular injections (corticosteroid, hyaluronic acid, platelet rich plasma) for the knee osteoarthritis. World J Orthop 5 (2014): 351-361.
- Arya RK, Jain V. Osteoarthritis of the knee joint:an overview. JIACM 14 (2013): 154-162.
- Evangelou E, Kerkhof HJ, Styrkarsdottir U, et al. A meta-analysis of genome-wide association studies identifies novel variants associated with osteoarthritis of the hip. Ann Rheum Dis 73 (2014): 2130-2136.
- Aigner T, Dudhia J. Genomics of osteoarthritis. Curr Opin Rheumatol 15 (2003): 634-640.
- Raisz LG. Prostaglandins and bone: physiologyand pathophysiology. Osteoarthr Cartil 7 (1999): 419-421.
- Déak F, Wagener R, Kiss I, et al. The matrilins:a novel family of oligomeric extracellular matrix proteins. Matrix Biol 18 (1999): 55-64.
- Chapman KL, Mortier GR, Chapman K, et al. Mutations in the region encoding the von Willebrand factor A domain of matrilin- 3 are associated with multipleepiphyseal dysplasia. Nat Genet 28 (2001): 393-396.
- Gu J, Rong J, Guan F, et al. Matn gene polymorphism is associated with osteoarthritis in Chinese Han population: a community- based case-control study. Sci World J 2012 (2012): 1-6.
- Pullig O, Tagariello A, Schweizer A, et al. MATN3 (matrilin-3) sequence variation (pT303M) is a risk factor for osteoarthritis of the CMC1 joint of the hand, but not for knee osteoarthritis. Ann Rheum Dis 66 (2007): 279-280.
- Borochowitz ZU, Scheffer D, Adir V, et al. Spondylo- epi-metaphyseal dysplasia (SEMD) matrilin 3 type: homozygote matrilin 3 mutation in a novel form of SEMD. J Med Genet 41 (2004): 366-372.
- Stefánsson SE, Jónsson H, Ingvarsson T, et al. Genomewide scan for hand osteoarthritis: A novel mutation in matrilin-3. Am J Hum Genet 72 (2003): 1448-1459.
- Diab SM, Kamal HM, Mansour AI, et al. Clinical significance of Matrilin-3 gene polymorphism in Egyptian patients with primary knee osteoarthritis. Eur J Rheumatol Link 4 (2017): 200-204.
- Minafra L, Bravatà V, Saporito M, et al. Genetic, clinical and radiographic signs in knee osteoarthritis susceptibility. Arthritis Res Ther 16 (2014): 91.
- Sydorchuk LP, Serdulets YI, Fediv OI, et al. The polymorphism of matrilin-3 (rs77245812) and interleukin-10 (rs1800872) genes in osteoarthritis patients with arterial hypertension, obesity and type 2 diabetes mellitus. Arch Balk Med Union 52 (2017): 422-429.
- Fresquet M, Jowitt TA, Ylöstalo J, et al. Structural and functional characterization of recombinant matrilin-3 A-domain and implications for human genetic bone diseases. J Biol Chem 282 (2007): 34634.
- Calculation of Chi-square test for deviation from Hardy-Weinberg equilibrium.
- Muttigi MS, Han Inbo, Hun-Kuk Park, et al. Matrilin-3 Role in Cartilage Development and Osteoarthritis Published online. Int J Mol Sci 17 (2016): 590.
- Belluoccio D, Schenker T, Baici A, et al. Characterization of human matrilin-3 (MATN3). Genomics 53 (1998): 391-394.
- Pullig O, Weseloh G, Klatt AR, et al. Matrilin-3 in human articular cartilage: increased expression in osteoarthritis. Osteoarthr Cartil 10 ( 2002): 253-263.
- Pullig O, Tagariello A, Schweizer A, et al. MATN3 (matrilin-3) sequence variation (pT303M) is a risk factor for osteoarthritis of the CMC1 joint of the hand, but not for knee osteoarthritis. Ann Rheum Dis 66 (2007): 279-280.
- Min JL, Meulenbelt I, Riyazi N, et al. Association of matrilin-3 polymorphisms with spinal disc degeneration and with osteoarthritis of the CMC1 joint of the hand. Ann Rheum Dis 65 (2006): 1060-1066.
- Van der Weyden L, Wei L, Luo J, et al. Functional knockout of the matrilin-3 gene causes premature chondrocyte maturation to hypertrophy and increases bone mineral density and osteoarthritis. Am J Pathol 169 (2006): 515-527.
- Vincourt JB, Vignaud JM, Lionneton F, et al. Increased Expression of matrilin-3 Not Only in Osteoarthritic Articular Cartilage but Also in Cartilage-Forming Tumors, and Down-Regulation of SOX9 via Epide. Arthritis Rheum 58 (2008): 2798-2808.
- Klatt AR, Klinger G, Paul-Klausch B, et al. Matrilin-3 activates the expression of osteoarthritis-associated genes in primary human chondrocytes. FEBS Lett 583 (2009): 3611-3617.
- Vincourt JB, Etienne S, Grossin L, et al. Matrilin-3 switches from anti- to pro-anabolic upon integration to the extracellular matrix. Matrix Biol 31 (2012): 290-298.
- Jayasuriya CT, Goldring MB, Terek R, et al. Matrilin-3 induction of IL-1 receptor antagonist is required for up-regulating collagen II and aggrecan and down-regulating ADAMTs-5 gene expression. Arthritis Res Ther 14 (2012): 197.
- Minafra L, Bravatà V, Saporito M, et al. Genetic, clinical and radiographic signs in knee osteoarthritis susceptibility. Arthritis Res Ther 16 (2014): 91.
- Diab SM, Kamal HM, Mansour AI, et al. Clinical significance of Matrilin-3 gene polymorphism in Egyptian patients with primary knee osteoarthritis. Eur J Rheumatol 4 (2017): 200-204.
- Yu C, Zhang S, Zhou C, et al. A likelihood ratio test of population Hardy-Weinberg equilibrium for case-control studies. Genet Epidemiol 33 (2009): 275-280.

 Impact Factor: * 1.7
Impact Factor: * 1.7 CiteScore: 2.9
CiteScore: 2.9  Acceptance Rate: 11.01%
Acceptance Rate: 11.01%  Time to first decision: 10.4 days
Time to first decision: 10.4 days  Time from article received to acceptance: 2-3 weeks
Time from article received to acceptance: 2-3 weeks 