Biosynthesis, Characterization and Antifungal Activity of Silver Nanoparticles by Aspergillus Niger Isolate
Article Information
Shrouaq Al-Zubaidi, Aisha Al-Ayafi, Hayam Abdelkader*
Microbiology Department, University of Jeddah, College of Science, Saudi Arabia
*Corresponding Author: Hayam Abdelkader, Microbiology Department, University of Jeddah, College of Science, Saudi Arabia
Received: 17 June 2019; Accepted: 27 June 2019; Published: 28 June 2019
Citation:
Shrouaq Al-Zubaidi, Aisha Al-Ayafi, Hayam Abdelkader. Biosynthesis, Characterization and Antifungal Activity of Silver Nanoparticles by Aspergillus Niger Isolate. Journal of Nanotechnology Research 2 (2019): 022-035.
View / Download Pdf Share at FacebookAbstract
A survey of fresh-market vegetables and fruits including orange (Citrus sinensis) lemon (Citrus lemon), tomatoes (Lycopersicon esculentum), grapes (Vitis vinifera), strawberries (Fragaria ananassa), cucurbita (Cucurbita pepo), cucumbers (Cucumis sativus), eggplants (Solanum melongena), Bell pepper (Capsicum annuum), and soft dates (Phoenix dactylifera Linn.) were conducted in Jeddah city to control the most common fungal infections by using biosynthesized Silver nanoparticles as antifungal agent. The results showed that Aspergillus Sp. had the highest occurrence in grapes, Onion, tomatoes and soft dates with a frequency of 70%, followed by Fusarium oxysporum with the frequency of occurrence of 31% in fruits and vegetables such as, strawberries, lemon, oranges, egg plants, cucumber and tomatoes while Penicillium digitatum had the least frequency of 10% each in orange, and lemon, respectively. The objective of this study was to biosynthesize silver nanoparticles (AgNPs) using A. niger fungal isolate and its evaluation as safe antifungal agent against plant pathogenic fungi. UV-visible spectroscopy, dynamic light scattering (DLS), and Fourier transform infra-red spectroscopy (FTIR) were used to characterize the biosynthesized AgNPs. The UV-visible spectra showed a characteristic peak at 430 nm, which correlate to the surface plasmon absorbance (SPA) of AgNPs. Size distribution pattern observation showed particles with sizes ranged from 10-100 nm. Mycosynthesized AgNPs showed considerable antifungal activity against pathogenic plant fungi.
Keywords
Mycosynthesized AgNPs, Antifungal, A. niger, UV-visible, FTIR, SPA, DLS
Mycosynthesized AgNPs articles Mycosynthesized AgNPs Research articles Mycosynthesized AgNPs review articles Mycosynthesized AgNPs PubMed articles Mycosynthesized AgNPs PubMed Central articles Mycosynthesized AgNPs 2023 articles Mycosynthesized AgNPs 2024 articles Mycosynthesized AgNPs Scopus articles Mycosynthesized AgNPs impact factor journals Mycosynthesized AgNPs Scopus journals Mycosynthesized AgNPs PubMed journals Mycosynthesized AgNPs medical journals Mycosynthesized AgNPs free journals Mycosynthesized AgNPs best journals Mycosynthesized AgNPs top journals Mycosynthesized AgNPs free medical journals Mycosynthesized AgNPs famous journals Mycosynthesized AgNPs Google Scholar indexed journals Antifungal articles Antifungal Research articles Antifungal review articles Antifungal PubMed articles Antifungal PubMed Central articles Antifungal 2023 articles Antifungal 2024 articles Antifungal Scopus articles Antifungal impact factor journals Antifungal Scopus journals Antifungal PubMed journals Antifungal medical journals Antifungal free journals Antifungal best journals Antifungal top journals Antifungal free medical journals Antifungal famous journals Antifungal Google Scholar indexed journals A. niger articles A. niger Research articles A. niger review articles A. niger PubMed articles A. niger PubMed Central articles A. niger 2023 articles A. niger 2024 articles A. niger Scopus articles A. niger impact factor journals A. niger Scopus journals A. niger PubMed journals A. niger medical journals A. niger free journals A. niger best journals A. niger top journals A. niger free medical journals A. niger famous journals A. niger Google Scholar indexed journals UV-visible articles UV-visible Research articles UV-visible review articles UV-visible PubMed articles UV-visible PubMed Central articles UV-visible 2023 articles UV-visible 2024 articles UV-visible Scopus articles UV-visible impact factor journals UV-visible Scopus journals UV-visible PubMed journals UV-visible medical journals UV-visible free journals UV-visible best journals UV-visible top journals UV-visible free medical journals UV-visible famous journals UV-visible Google Scholar indexed journals FTIR articles FTIR Research articles FTIR review articles FTIR PubMed articles FTIR PubMed Central articles FTIR 2023 articles FTIR 2024 articles FTIR Scopus articles FTIR impact factor journals FTIR Scopus journals FTIR PubMed journals FTIR medical journals FTIR free journals FTIR best journals FTIR top journals FTIR free medical journals FTIR famous journals FTIR Google Scholar indexed journals SPA articles SPA Research articles SPA review articles SPA PubMed articles SPA PubMed Central articles SPA 2023 articles SPA 2024 articles SPA Scopus articles SPA impact factor journals SPA Scopus journals SPA PubMed journals SPA medical journals SPA free journals SPA best journals SPA top journals SPA free medical journals SPA famous journals SPA Google Scholar indexed journals DLS articles DLS Research articles DLS review articles DLS PubMed articles DLS PubMed Central articles DLS 2023 articles DLS 2024 articles DLS Scopus articles DLS impact factor journals DLS Scopus journals DLS PubMed journals DLS medical journals DLS free journals DLS best journals DLS top journals DLS free medical journals DLS famous journals DLS Google Scholar indexed journals livestock articles livestock Research articles livestock review articles livestock PubMed articles livestock PubMed Central articles livestock 2023 articles livestock 2024 articles livestock Scopus articles livestock impact factor journals livestock Scopus journals livestock PubMed journals livestock medical journals livestock free journals livestock best journals livestock top journals livestock free medical journals livestock famous journals livestock Google Scholar indexed journals Silver nanoparticles articles Silver nanoparticles Research articles Silver nanoparticles review articles Silver nanoparticles PubMed articles Silver nanoparticles PubMed Central articles Silver nanoparticles 2023 articles Silver nanoparticles 2024 articles Silver nanoparticles Scopus articles Silver nanoparticles impact factor journals Silver nanoparticles Scopus journals Silver nanoparticles PubMed journals Silver nanoparticles medical journals Silver nanoparticles free journals Silver nanoparticles best journals Silver nanoparticles top journals Silver nanoparticles free medical journals Silver nanoparticles famous journals Silver nanoparticles Google Scholar indexed journals anti-inflammatory articles anti-inflammatory Research articles anti-inflammatory review articles anti-inflammatory PubMed articles anti-inflammatory PubMed Central articles anti-inflammatory 2023 articles anti-inflammatory 2024 articles anti-inflammatory Scopus articles anti-inflammatory impact factor journals anti-inflammatory Scopus journals anti-inflammatory PubMed journals anti-inflammatory medical journals anti-inflammatory free journals anti-inflammatory best journals anti-inflammatory top journals anti-inflammatory free medical journals anti-inflammatory famous journals anti-inflammatory Google Scholar indexed journals
Article Details
1. Introduction
Many pathogenic fungi have harmful effects causing most of the diseases to fruits and vegetables that affect the quality of the crops causing high economic loss in Saudi Arabia [1]. Recently, consumption of fruit and vegetables has increased sharply in KSA by more than 40% during the past few years. Approximately 20% of all fruits and vegetables are lost every year because of spoilage [2]. Over direct contact with soil, dust, water during harvesting or post harvesting, fruits and vegetables exposed to contamination by microorganisms, especially plant fungi [3]. Fungi generate a huge amount of extracellular pectinases and hemicellulases that are important for fungal spoilage [4]. Some microbes can colonize and damage healthy plant tissue [5]. Management of fungal diseases of food crops and fruits is economically important. Recently, a higher effort has been given to develop secure management techniques that pose less risk to humans and livestock, with a focus on the consequences of synthetic fungicides. Kim et al. [6], Shams et al. [7] demonstrated that AgNPs had low toxicity, a broad spectrum of antimicrobial activity and also very effective against plant phytopathogenic fungi.
Silver nanoparticles are very attracting and interesting for several applications because of their unique and remarkable properties, enhanced permeability, retention effect and antimicrobial activity [8]. Silver nanoparticles are commonly used due to their wide antimicrobial activity against various micromicroorganisms and localized surface plasmon resonance effect [9, 10]. For many years, silver has been used as antimicrobial in the medical field in burn treatment [11]. Silver is more toxic to microorganisms compared to other metals, while it has a reduced toxicity for mammalian cells [12]. Silver nanoparticles have anti-fungal activity as reported by Wiley et al. [13] anti-inflammatory. Fungi are the best candidates for metal nanoparticle synthesis because they are able to secrete large amounts of enzymes and proteins which directly translate into higher nanoparticle productivity [14-16]. Fungi are also easy to handle, require simple nutrient, have a high wall binding capacity, as well as capable of intra cellular metal uptake [17]. Biological systems offer a novel concept for the manufacturing of nanomaterial compared to traditional synthetic techniques [18]. Many fungi like Verticilium sp [19], Aspergillus fumigates [20], Trichoderma asperellum [21], Fusarium oxysporum [22], Aspergillus niger [23], Phoma glomerata was widely used in nanoparticles production [24].
Silver nanoparticles’ mechanisms for antimicrobial effects are not yet understood, but several reports have shown that Silver nanoparticles may damage the cell wall and cell membrane as reported by Xia. Silver nanoparticles penetrated the cells causing damage to the organelles, including mitochondria and ribosome, resulting in condensation and margination of chromatin, a marker of apoptotic cell death. In addition, because of their tiny size, silver nanoparticles can bind to the cell surface and directly penetrate the cells without damaging the cell wall and trigger a cell death. The goals of the current study are, 1) mycosynthesis of silver nanoparticles using A. niger, 2) evaluation of biosynthesized AgNPs as antifungals against plant pathogenic fungi.
2. Materials and Methods
2.1 Sample collection
A survey was conducted in Jeddah, extending from November 2018 to January 2019. Samples of infected vegetables and fruits were randomly collected from 10 locations and 15 markets in Jeddah city. The samples were transferred into sterile plastic bags and stored in laboratory conditions for further investigation. Tissues showing symptoms of post harvest disease were cultured to identify associated pathogens.
2.2 Isolation and identification of fungal pathogens
For isolation of fungal pathogens, small piece (1 cm) of infected fruit was cut, surface sterilized with 2% sodium hypochlorite solution for 1 min, washed twice with sterile water and cultured on PDA plates then incubated at 26-28°C for 24 h. Single hyphal tips were cultured on PDA medium and incubated at 26-28°C for 5 days [25]. Purified cultures were visually identified using laboratory manuals [26]. All fungal isolates were maintained in 4°C until used. Presumptive identifications were confirmed with ITS rDNA sequence analysis [27].
2.3 Molecular identification
Fungal DNA was extracted from freshly collected mycelium of 10- day-old cultures using the Microbial DNA Isolation Kit (Qiagen) according to manufacturer's instructions. The entire ITS region of the fungal isolates was amplified with the primer pair ITS1F (5’-CTTGGTCATTTAGAGGAAGTAA-3’) and ITS4 (5’-TCCTCCGCTTATTGATATGC-3’). PCR was performed as previously described by Nikolcheva et al. [28]. The PCR products were electrophoresed on 1% agarose gels, stained with 0.5 µg/ml ethidium bromide, and bands visualized with a UV illuminator. Negative control PCR reaction was also included. DNA sequencing PCR products were fractionated by electrophoresis in agarose gels and purified using a QIAquick Gel Extraction Kit (Qiagen). The recovered DNA was sequenced by the chain termination method using Sequencing Kit (BigDye terminator v2.0) and an ABI PRISM 377-96 automated sequencer.
2.4 Preparation of A. niger filtrate
200 ml of sterile YM (Yeast extract, Malt extract, Peptone, Glucose) broth media containing malt extract were inoculated with 5 mm fungal discs taken from A. niger plate (One weak-old). The inoculated media were incubated with shaking at 140rpm in the dark at 26°C for 7 days. After incubation, biomass of A. niger was filtrated on a filter paper, washed with sterile dH2O to remove any traces of media components, resuspended in 100 mL dH2O, incubated at 26°C for one day, and then filtered again.
2.5 Bioproduction of Ag-NPs
1 mM Silver nitrate (AgNO3) was added to the filtrate to promote the formation of AgNPs. The biomass filtrate and AgNO3 ratio was remained at 1 to 9 (v/v), and incubation of the reaction mixture was kept at 26°C for two days. Under the same conditions, controls (without addition of AgNO3) have been included.
2.6 Analysis of AgNPs using UV-visible spectrophotometer
UV-Vis spectroscopy was used to inspect the size and shape of biosynthesized nanoparticles in aqueous solutions [13]. Color change in the reaction mixture was the initial indicator of the formation of AgNPs. Three mL of the reaction mix was transferred to measure its absorbance when the color of the suspension was turned brown. UV-visible spectrophotometer (UV-2450; Shimadzu, Tokyo, Japan) was used for scanning the sample at 300- to 700-nm absorbance spectrum.
2.7 FTIR spectrophotometer
In order to separate the fungal biomass residues, the reaction mixture was centrifuged at 14,000 rpm for 25 minutes and the AgNPs pellets were resuspended in dH2O. Centrifugation and resuspension steps repeated for two to three times. Finally the samples were dried and suspended in sterile H2O and then analyzed by using FTIR instrument (Thermo Scientific Smart iTR™).
2.8 Antifungal test
The antifungal effect of AgNPs was examined against the fungal isolates Fusarium oxysporum, Aspergillus flavus and Penicillin digitatum. Fungal cells were grown in PDA liquid media at 26°C for 5 days, and then cells containing 1 × 106 colony-forming units/mL were cultured on fresh PDA solid media. Agar well diffusion method [29] was conducted. Wells of 8 mm diameter were punched into the PDA medium and filled with 50 μl containing different concentrations of AgNPs (2 µg/mL, 4 µg/mL, 6 µg/mL, 8 µg/mL, and 10 µg/mL) and incubated at 26°C for 5 days. Controls of Silver-free plates were incubated under the same conditions.
2.9 Data analysis
The radial growth of fungal mycelium was recorded on PDA plates containing AgNPs. When mycelial growth reached the edge of the petridish, radial inhibition was calculated. The following equation was used for the inhibition rate (%) calculation.
Inhibition rate 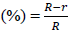
Where R is the fungal radial growth of on the control silver free plate and r is the radial of fungal growth on culture plate inoculated with AgNPs.
2.10 Measurement of MIC
The MIC of the antifungal effect for AgNPs was conducted corresponding to the National Committee for Clinical Laboratory Standards [29]. The antifungal activity was evaluated against the final fungal concentration of 106 colony-forming units/mL. Nystatin was used as a control. All experiments were performed thrice.
3. Results and Discussion
3.1 Fungal identification using ITS rRNA gene sequencing
The isolated fungus appeared as blackish brown in color (Figure 1) was identified by PCR amplification of 18S rRNA gene using ITS primers. In this study, PCR techniques have been performed successfully for the detection of fungal species based on sequence analysis. A 600 bp DNA fragment was amplified by PCR from Aspergillus niger aggregate isolated from grapes with universal primer specific for distinguishing fungal species. The nucleotide sequences of the (ITS 1-5.8S-ITS 2) of 18 s rRNA gene region of Aspergillus isolates was aligned with other reference isolates by using the Blastn analyses, pairwise and multiple sequence alignment which revealed 98-100% identity with the sequences of A. niger strains. This result is consistent with reports by several workers [30-32].
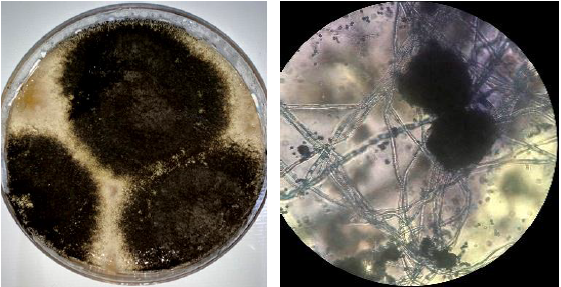
Figure 1: Colony morphology of Aspergillus niger isolated from the Vitis vinifera (A) Macroscopic morphology (PDA, 26°C, 7 days) and (B) Microscopic morphology.
3.2 Biosynthesis of silver nanoparticles (AgNPs)
In the present study, A. niger isolate was successfully incorporated in biosynthesis of silver nanoparticles by reducing silver nitrate as described by Sagar et al. [23]. A change to an intense brown color of the culture filtrate with silver nitrate solution has been noticed after 24 hour of incubation, whereas, the control has not shown any color change (Figure 2). The reduction of silver ions (Ag+) into AgNPs (Ag0) was indicated by a color change. This color change is mainly due to the Plasmon resonance surface of deposited silver nanoparticles, i.e., the color of the nanoparticles was due to coherent and collective surface electrons oscillations [33]. It has been reported that AgNPs exhibit a dark brown color in aqueous solution as a result of surface plasmon resonance (SPR). Similarly, metal nanoparticles posses well-formed irradiated colors related to the localized surface plasmon resonance (SPR) [34, 35].

Figure 2: (a). Light yellow color of the reaction solution containing 1 mM AgNO3 and fungal filtrate; (b) 1 mM of AgNO3 solution; (c) Color change of the reaction solution from light yellow to dark brown indicating the formation of AgNPs.
3.3 UV-visible spectra analysis
The UV-visible spectra of synthesized AgNPs (Figure 3) showed characteristic SPR peak at 430 nm and clearly indicates the increased intensity of the silver nitrate solution after 48 hours, suggesting the formation of an increased number of silver nanoparticles. The SPR peak at 430 nm was detected to be a characteristic of variant metabolites and proteins existing in the extracellular fungal filtrate, which have a vital role in the metal ions reduction by fungal induced nanoparticles synthesis. Our findings are in agreement with the results obtained by Sagar et al. [23].
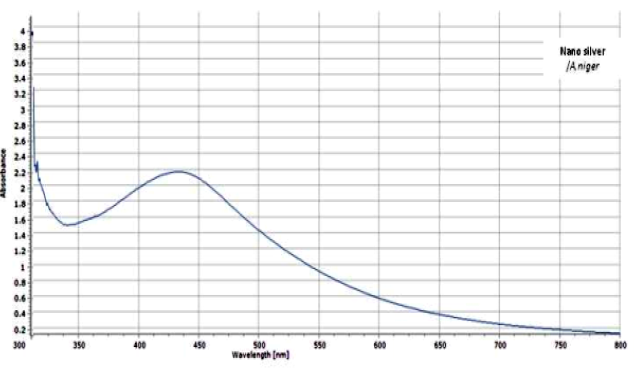
Figure 3: UV-visible absorption spectrum of AgNPs biosynthesized by the reduction of AgNO3 solution with the fungal filtrate of A. niger after 2h.
3.4 Size distribution pattern of synthesized nanoparticles
Results obtained from dynamic light scattering (DLS) pattern measured using (Malvern Zetasizer Nano) reveal that the biosynthesized silver nanoparticles have an average diameter of 30 nm with polydispersity index (PDI) of 0.144 (Figure 4). Particle size range of the nanoparticles in addition to its polydispersity was calculated. This analysis tool is very suitable for determining the mean size of nanoparticles inside the sample. Particle size was determined based on dynamic light scattering of laser light.
3.5 FTIR spectra analysis
FTIR analysis was carried out to characterize the potential biomolecules responsible for the reduction of Ag+ ions and the stabilization of the bioreduced silver nanoparticles to prevent nanoparticles clustering and capping in the aqueous medium Basavaraja et al. [36]. The FTIR analysis has displayed a peak range from 500 to 4000 cm-1. Intensive peaks at 3329, 2358, 1638, 1401, 1249, 1056 and 556 cm-1 (Figure 5) were corresponding to various functional groups and demonstrated the existence of protein stabilizing molecules. The peak at 3329 cm-1 corresponding to N-H stretching of the secondary amide of the protein and the peak at 2358 cm-1 corresponding to several functional groups [37]. Peak at 1638 cm-1 is associated with stretch vibration of -C=C- [38] (characteristic of amino acids containing NH2 groups, amide I band), which is assumed for the amide I bond of proteins [39].
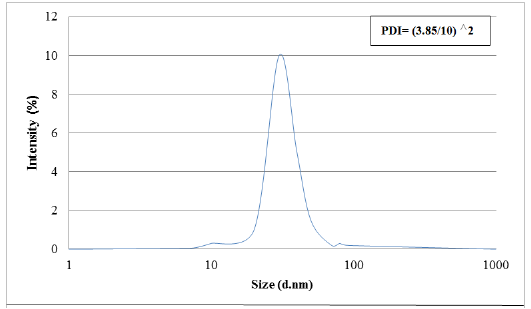
Figure 4: Graph showing particle size distribution for AgNPs.
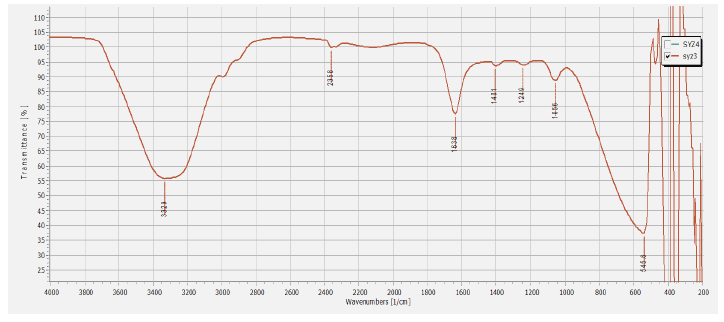
Figure 5: FTIR analysis of biosynthesized AgNPs.
The sharp peak at 1401 cm-1 can be assigned to associated with benzene ring stretching of C=C and C-C bonds [40, 41]. The peak at 1249 corresponding to in plane deformation of C-H bonds in the pyridine ring. The peak and 1056 cm-1 can be attributed to C-OH of the phenols, supporting the reduction of Ag+ into Ag0 through the participation of polyphenols, such as flavanoids and triterpenoids [42, 43]. The peak at 556 may correspond to torsional deformation of CH bonds. The reduction of silver ions Ag+ into nanosize silver particles Ag0 could be due to the presence of extracellular proteins in the fungal filtrate. These proteins have a powerful ability to bind silver nanoparticles, acting as capping agents and hence provide the stability to them. These results are consistent with the earlier reports for the fungal mediated synthesis of AgNPs [34, 44, 45]. FTIR analysis revealed that polyphenols could function as bioreducing agents, while proteins could play a dual role as bioreducing and stabilizing agents.
3.6 Antifungal activity of AgNPs
The antimicrobial activity of AgNPs against various pathogenic fungi was investigated (Figure 6). Compared with the control, the inhibition of fungal growth increased for all the tested fungi. The biothensized AgNPs inhibitd the growth of three different pathogenic fungi, including Fusarium oxysporum, Aspergillus flavus and Penicillin digitatum, as previously reported. Thus, AgNPs could be considered as excellent broad-spectrum antifungal agents. Since the biosynthesized AgNPs showed considerable antifungal activity, they could potentially be used widely in clinical applications.
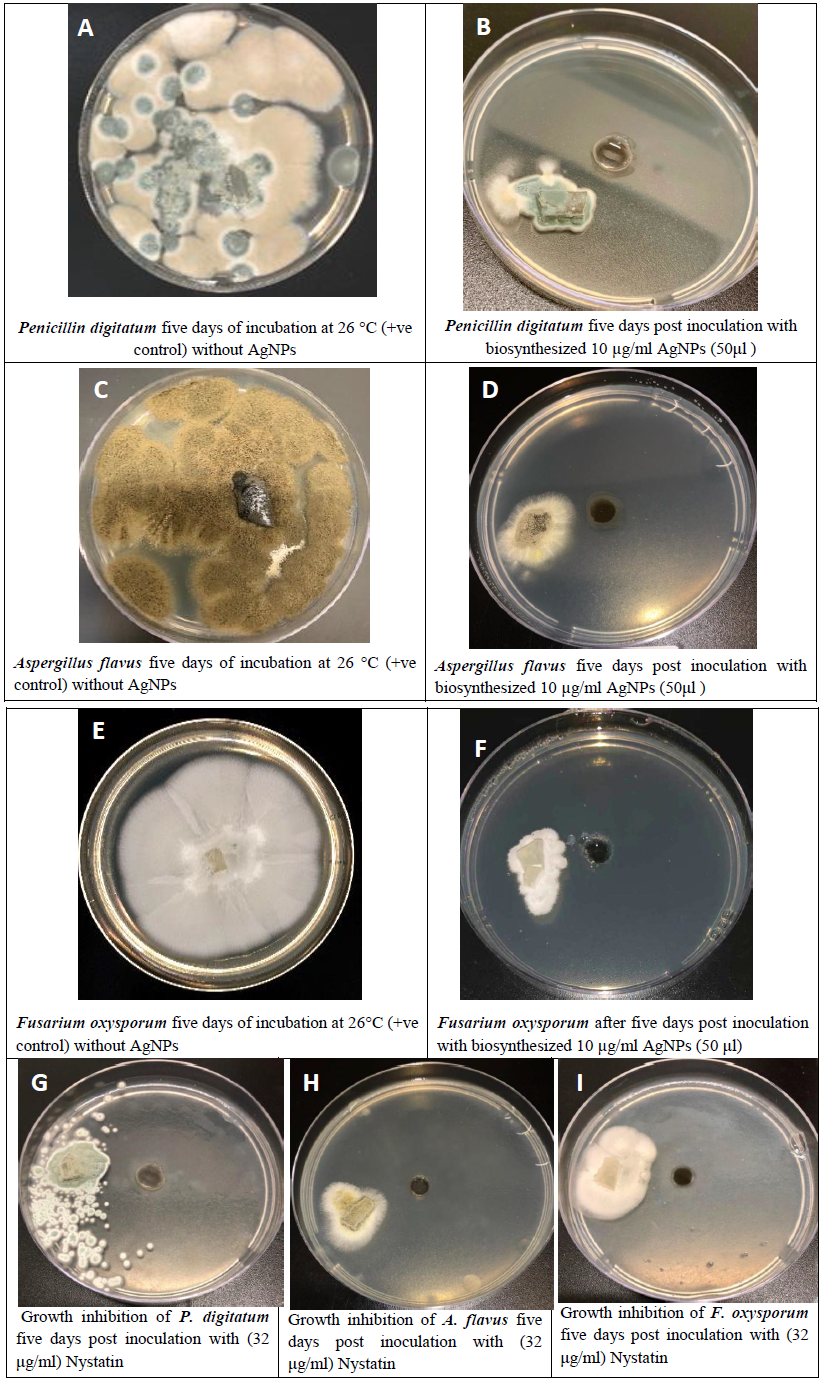
Figure 6: Antifungal activity of the synthesized silver nanoparticles against different pathogenic fungi (A, C and E). Growth inhibition of fungal isolates produced by biosynthesized silver nanoparticles (B, D, F) five days post inoculation with 10 µg/ml AgNPs. Positive control of (A) Penicillin digitatum, (C) Aspergillus flavus, and (F) Fusarium oxysporum. G, H and I: Growth inhibition of fungal isolates by commercial antifungal (Nystatin).
3.7 MICs of silver nanoparticles against isolated fungi
MICs of silver nanoparticles against isolated fungi are shown in Table 1. The MIC of silver nanoparticles was estimated to be 0.5-10.0 µg/mL, which were lower than those of Nystatin. The results suggest that the mycosynthesized AgNPs are capable of inhibiting isolated fungi Penicillin digitatum, Aspergillus flavus, and Fusarium oxysporum and the MICs of AgNPs were 6.7-9.62 µg/ml (Table 1).
|
Isolate |
MIC (µg/mL) |
|
|
AgNPs |
Nystatin |
|
|
P. digitatum |
6.75 ± 0.24 |
8.1 ± 0.30 |
|
A. flavus |
7.45 ± 0.18 |
9.2 ± 0.21 |
|
F. oxysporum |
9.62 ± 0.14 |
10.4 ± 0.06 |
Data are presented as mean ± standard deviation.
Table 1: Minimum inhibitory concentration (MIC) of silver nanoparticles and other common antifungal drug against plant pathogenic fungi.
In addition, results indicate that a higher inhibition rate was observed at the 10 µg/ml concentration of AgNPs in comparison to Nystatin (Figure 3). Similarly, a recent publication also showed that biosynthesized silver nanoparticles had antimicrobial effects against eighteen plant pathogenic fungi-isolated from spoiled fruits and vegetables [6]. Inhibition effect of AgNP Data on inhibition effect of silver nanoparticles (AgNPs) against A. flavus, F.oxysporum, and P. digitatum on PDA in vitro was shown in Table 2. Inhibition (97.3%) was obtained against A. flavous treated with a 10 µg/ml concentration of silver nanoparticles. The minimal level of inhibition was observed against P. digitatum and F. oxysporium with 2 µg/ml concentrations of AgNPs. Accordingly, the results demonstrated that in the vast majority of cases, the inhibition effect increases with the increment of AgNP concentration. This could be a result of the high density at which the solution was able to saturate and adhere to fungal hyphea deactivating plant pathogenic fungi. Several reports on the mechanism of silver ions inhibitory action on microorganisms have shown that DNA loses its ability to replicate upon treatment with Ag+ [46], resulting in inactivated expression of ribosomal subunit proteins, as well as certain other cellular proteins and enzymes essential to ATP production [47].
|
Isolate |
Inhibition rate (%) |
||||
|
2 μg/ml |
4 μg/ml |
6 μg/ml |
8 μg/ml |
10 μg/ml |
|
|
P. digitatum |
40.0 |
73.3 |
80.0 |
86.6 |
93.75 |
|
A. flavus |
56.25 |
62.5 |
68.75 |
87.5 |
97.3 |
|
F. oxysporum |
43.75 |
50.0 |
56.25 |
62.5 |
91.0 |
Inhibition rates were determined based on five replicates of each experiment, inhibition rate of control=0%.
Table 2: Inhibitory rate (%) of silver nanoparticles against plant pathogenic fungi.
4. Conclusion
The biosynthesis of nanoparticles by microorganisms is feasible, safe, cheap and less time consuming, it provides effective satisfactory results without any hazardous chemicals involvement. In the present study, A. niger was exploited to biosynthesize silver nanoparticles by reducing silver nitrate. These mycosynthesized AgNPs exerted powerful antifungal effects on the in vitro tested fungi, likely by destroying membrane integrity. With the application point of view, a suitable pharmaceutical formulation using these nanoparticles, as well as studies on different biological activities in different fields should be strengthened in future studies.
References
- Al-Najada AR and Gherbawy YA. Molecular Identification of Spoilage Fungi Isolated from Fruit and Vegetables and Their Control with Chitosan. Food Biotechnology 29 (2015): 166-184.
- Droby S. Improving quality and safety of fresh fruits and vegetables after harvest by the use of biocontrol agents and natural materials. Acta Horticulura 709 (2006): 45-51.
- Eni AO, Ibokunoluwa O, and Oranusi U. Microbial quality of fruits and vegetables. African Journal of Food Science 4 (2010): 291-296.
- Miedes E and Lorences EP. Apple (Malus domestica) and Tomato (Lycopersicum esculentum) Fruits Cell-Wall Hemicelluloses and Xyloglucan Degradation during Penicillium expansum Infection. Journal of Agricultural and Food Chemistry 52 (2005): 7957-7963.
- Tournas VH. Spoilage of Vegetable Crops by Bacteria and Fungi and Related Health Hazards. Critical Review of Microbiology 31 (2005): 33-44.
- Kim SW, Jung JH, Lamsa K, et al. Antifungal Effects of Silver Nanoparticles (AgNPs) against Various Plant Pathogenic Fungi. Mycobiology 40 (2012): 53-58.
- Shams-Ghahfarokhi M, Aghaei-Gharehbolagh S, Aslani N, et al. Investigation on distribution of airborne fungi in outdoor environment in Tehran, Iran. J Environ Health Sci Eng 12 (2014): 54.
- Pellegri FN, Nicastro D and de Sanctis O. Effect of amine groups in the synthesis of Ag nanoparticles using aminosilanes. Mater Chem Phys 94 (2005): 148-152.
- Shen J, Shi M, Li N, et al. Facile synthesis and application of of Ag-chemically converted graphene nanocomposite. Nano Res 3 (2010): 339-349.
- Chernousova S and Epple M. Silver as antibacterial agent: ion, nanoparticle, and metal. Angew Chem Int Ed Engl 52 (2013): 1636-1653.
- Maiti S, Krishnan D, Barman G, et al. Antimicrobial activities of silver nanoparticles synthesized from Lycopersicon esculentum extract. Journal of Analytical Science and Technology 5 (2014): 40.
- Zhao W, Wolfendar JL, Hostettmann K, et al. Antifungal alkaloids and limonoid derivatives from Dictamnus dasycarpus. Phytochemistry 47 (1998): 7-11.
- Wiley BJ, McLellan J, Siekkkinen A, et al. Maneuvering the surface Plasmon resonance of silver nanostructures through shape-controlled synthesis. J Phy Chem B 110 (2006): 15666.
- Slawson RM, Van Dyke MI, Lee H, et al. Germanium and silver resistance, accumulation and toxicity in microorganisms. Plasmid 27 (1992): 73.
- Bae D, Kim E, Bang J, et al. Synthesis and Characterization of Silver Nanoparticles by a Reverse Micelle Process. Met Mater Int 4 (2005): 291-294.
- Mohanpuria P, Nisha R and Sudesh Y. Biosynthesis of nanoparticles: Technological concepts and future applications. Journal of Nanoparticle Research 10 (2008): 507-517.
- Prameela Devi T, Kulanthaivel S, Kamil D, et al. Biosynthesis of silver nanoparticles from Trichoderma species. Indian Journal of Experimental Biology 51 (2013): 543-547.
- Narayanan BK and Sakthivel N. Biological synthesis of metal nanoparticles by microbes. Advances in colloid and interface science 156 (2010): 1-13.
- Mukherjee P, Ahmad A, Mandal D, et al. Bioreduction of AuCl4-ions by the fungus Verticillium sp and surface trapping of the gold nanoparticles formed, Angewante Chemie International Edition 40 (2001): 3585.
- Bhainsa KC and D’Souza SF. Extracellular biosynthesis of silver nanoparticles using the fungus Aspergillus fumigates. Colloids and Surfaces B, Biointerface 47 (2006): 160.
- Mukherjee P, Roy M, Mandal BP, et al. Green synthesis of highly stabilized nanocrystalline silver particles by a nonpathogenic and agriculturally important fungus asperellum, Nanotechnology 19 (2008): 103.
- Verma VC, Singh SK, Solanki R, et al. Biofabrication of anisotropic gold nanotriangles using extract of endophytic Aspergillus clavatus as a dual functional reductant and stabilizer. Nanoscale Res Lett 6 (2011): 16.
- Sagar G and Ashok B. Green Synthesis of Silver Nanoparticles Using Aspergillus niger and Its Efficacy Against Human Pathogens. European Journal of Experimental Biology 2 (2012): 1654-1658.
- Birla SS, Tiwari VV, Gade AK, et al. Fabrication of silver nanoparticles by Phoma glomerata and its combined effect against Escherichia coli Pseudomonas aeruginosa and Staphylococcus aureus, Lett Applied Microbio 27 (2009): 76.
- Carisse O and Van Der Heyden H. Relationship of Airborne Botrytis cinerea Conidium Concentration to Tomato Flower and Stem Infections: A Threshold for De-leafing Operations. Plant Dis 99 (2015): 137-142.
- Dugan FM. The identification of fungi: an illustrated introduction with keys, glossary, and guide to literature. St. Paul, Minn.: American Phytopathological Society (2006).
- Raja HA, Miller AN, Pearce CJ, et al. Fungal Identification Using Molecular Tools: A Primer for the Natural Products Research Community. J Nat Prod 80 (2017): 756-770.
- Nikolcheva LG, Cockshutt AM and barlocher F. Determining 741 diversity of freshwater fungi on decaying leaves: comparison of traditional and molecular 742 approaches. Appl Enviro. Microbiol 69 (2003): 2548-2554.
- National Committee for Clinical Laboratory Standards. Reference method for broth dilution antifungal susceptibility testing of yeasts. Approved standard M27eA2. Wayne, PA: Clinical and Laboratory Standards Institute (2002).
- Accensi F, Cano J, Figuera L, et al. New PCR method to differentiate species in the Aspergillus niger aggregate. FEMS Microbiology Letters 180 (1999): 191-196.
- Perrone G, Susca A, Stea G, et al. PCR assay for identification of Aspergillus carbonarius and Aspergillus japonicus. European Journal of Plant Pathology 110 (2004): 641-649.
- Gonzalez-Salgado A, Patino B, Vazquez C, et al. Discrimination of Aspergillus niger and other Aspergillus species belonging to section Nigri by PCR assays.FEMS Microbiol Lett 245 (205): 353-361.
- Link S and El-Sayed MA. Optical properties and ultrafast dynamics of metallic nanocrystals. Annu Rev Phys Chem 54 (2003): 331.
- Jaidev LR and Narasimha G. Fungal mediated biosynthesis of silver nanoparticles, characterization and antimicrobial activity. Colloids. Surf B Biointerfaces 81 (2010): 430-433.
- Narasimha G Janardhan, Alzohairy M, Khadri H, et al. Extracellular synthesis, characterization and antibacterial activity of Silver nanoparticles by Actinomycetes isolative. Int J Nano Dimens 4 (2012): 77-83.
- Basavaraja S, Balaji S, Lagashetty K, et al. Extracellular biosynthesis of silver nanoparticles using the fungus Fusarium semitectum. Mater Res Bull 43 (2008): 1164-1170.
- Bozanic DK, Trandafilovic LV, Luyt AS, et al. Green synthesis and optical properties of silver-chitosan complexes and nanocomposites. React Function Polym 70 (2010): 869-873.
- Sivaraman SK, Elango I, Kumar S, et al. A green protocol for room temperature synthesis of silver nanoparticles in seconds. Current Sciences 97 (2009): 1055.
- Fayez AM, Balaji K, Kalaichelvan PT, et al. Fungal based synthesis of silver nanoparticles-an effect of temperature on the size of particles. Colloids Surf B: Biointerfaces 74 (2009): 123-126.
- Gipson K, Stevens K, Brown P, et al. Infrared Spectroscopic Characterization of Photoluminescent Polymer Nanocomposites. Journal of Spectroscopy (2015): 9.
- Khandelwal N, Abhijeet S, Devendra J, et al. Green synthesis of silver nanoparticles using Argimone mexicana leaf extract and evaluation of their antimicrobial activities. Dig J Nanomat Biostruct 5 (2010): 483-489.
- Litvin VA and Minaev BF. Spectroscopy study of silver nanoparticles fabrication using synthetic humic substances and their antimicrobial activity. Spectrochim Acta Part A Mol Biomol Spectrosc 108 (2013): 115-122.
- Litvin VA, Galagan RL and Minaev BF. Kinetic and mechanism formation of silver nanoparticles coated by synthetic humic substances. Colloids and surf A: Physicochem Engg Asp 414 (2012): 234-243.
- Rafie MH, Shaheen TI, Mohamed AA, et al. Bio-synthesis and applications of silver nanoparticles onto cotton fabrics. Carbohydr Polym 90 (2012): 915-920.
- Asad S, Supriya S, Gopal CK, et al. Biological synthesis of silver nanoparticles using the fungus Humicola sp. and evaluation of their cytotoxicity using normal and cancer cell lines. Spectrochim Acta A Mol Biomol Spectrosc 114 (2013): 144-147.
- Feng QL, Wu J, Chen GQ, et al. A mechanistic study of the antibacterial effect of silver ions on Escherichia coli and Staphylococcus aureus. J Biomed Mater Res 52 (2000): 662-668.
- Yamanaka M, Hara K and Kudo J. Bactericidal actions of a silver ion solution on Escherichia coli, studied by energy filtering transmission electron microscopy and proteomic analysis. Appl Environ Microbiol 71 (2005): 7589-7593.

 Impact Factor: * 1.2
Impact Factor: * 1.2 CiteScore: 2.9
CiteScore: 2.9  Acceptance Rate: 11.01%
Acceptance Rate: 11.01%  Time to first decision: 10.4 days
Time to first decision: 10.4 days  Time from article received to acceptance: 2-3 weeks
Time from article received to acceptance: 2-3 weeks 