The Involvement of Leptospira Spp. in Swine Abortion and Susceptibility of the Pathogen to Antibiotics
Gemerlyn G Garcia, Marylaine Ivy M Dioses*
College of Veterinary Science and Medicine, Central Luzon State University 3120, Science City of Muñoz, Nueva Ecija, Philippines
*Corresponding Author: Marylaine Ivy M Dioses, Professor, Department of Pathobiology, College of Veterinary Medicine, Central Luzon State University, Munoz, Nueva Ecija, Philippines
Received: 07 September 2021; Accepted: 15 September 2021; Published: 30 September 2021
Article Information
Citation: Gemerlyn G Garcia, Marylaine Ivy M Dioses. The Involvement of Leptospira Spp. in Swine Abortion and Susceptibility of the Pathogen to Antibiotics. Archives of Veterinary Science and Medicine 4 (2021): 71-84.
View / Download Pdf Share at FacebookAbstract
The involvement of Leptospira spp. in thirty-five (35) out of fifty (50) cases of swine abortion was investigated which further required evaluation of pathogen susceptibility to a panel of antibiotics. Samples of urine from aborting sows were collected to recover the pathogen, amplify its 16S rRNA gene to confirm its identity and monitor cell density in McFarland Units (MFU) as a measure of susceptibility to antibiotics. The study permitted the isolation and evaluation of Leptospira spp. morphology, demonstration of 3 amplicon products of the 16S rRNA genes with molecular sizes of 1050 to 1080 bp and identification of 2 Leptospira spp. and L. hyos after gene sequencing. The susceptibility of the identified L. hyos to Cephalexin, Neomycin, Clindamycin, Ciprofloxacin and Sulfamethoxazole prepared at different concentrations (25 µg, 35 µg and 50 µg) and evaluated after 6 and 12 hour-exposure time was marked by lower cell densities of recovered L. hyos relative to the application of higher (35 µg and 50 µg) antibiotic concentrations. Significant reduction in cell densities at 12-hr compared to data obtained on the 6-hour were demonstrated and underscored the susceptibility of L. hyos as effects of antibiotic concentration and duration of exposure time. The bactericidal action of 25 µg Ciprofloxacin; and 25 and 35 µg Sulfamethoxazole against L. hyos were established at 6-hour exposure time and that this bactericidal action was maintained until the 12-hr with the application of Neomycin and Ciprofloxacin at 35 and 50 µg and Sulfamethoxazole at 50 µg. These data highlight usefulness of the antibiotics in the treatment of abortion-associated Leptospira infections when administered at the concentrations described.
Keywords
<p>Abortion; Antibiotic sensitivity; Cell density; Leptospira hyos; McFarland units</p>
Article Details
1. Introduction
Leptospira spp. are commonly encountered and identified in cases of abortion in livestock based on findings of a collaborative research undertaken by this laboratory and the Biotechnology Office, Philippine Department of Agriculture. The pathogen takes the kidneys of mammals, birds, reptiles, amphibians and rodents as reservoirs of infection. Leptospirosis outbreaks can happen during the rainy season when floods likely happen. In the Philippines, leptospirosis outbreaks cause alarm during the rainy season when flooded areas may get contaminated with the urine of rodents that carry the pathogenic Leptospira. Cases of abortion may cater to a wider dissemination of the pathogen in animal production systems, especially when farm premises, water supply and injured tissues, and genital mucosal surfaces get contaminated.
Leptospirosis often presents as an emerging zoonotic disease and identification of the pathogen in abortion cases of livestock is a challenging area for research. Researchers who deal with studies on Leptospira spp. often face difficulty in monitoring the growth of the organism with the use of ordinary culture media as the pathogen does not form colonies on the surface of a solid medium. The complex growth patterns of the pathogen and its zoonotic nature possibly hamper the interest of many in exploring Leptospira widely and this probably contributes to the paucity of literature that describes the susceptibility of animal-borne Leptospira spp. to antibiotics. In veterinary practice, penicillin and doxycycline are commonly reported as treatment for animal leptospirosis [1] and relevant information that define the effective levels of antibiotic treatment remains inadequate.
Researches currently undertaken involve diagnosis of animal infections like cases of abortion in swine. This work usually calls for an urgent need to come up with a simple method for the detection of Leptospira spp. and identify a research-generated information for a basis for treatment. In this study, contemporary methods of molecular biology like PCR and DNA sequencing were used to confirm the identity of the pathogen in integration with the application of basic technics on the cultivation of a fastidious microbe like Leptospira spp. to simplify assessment of bacterial cell densities relative to antibiotic susceptibility. Data which show the susceptibility of the identified Leptospira spp. that accompany abortion in sows will serve as an important reference in the administration of an effective treatment against Leptospira infec-tions.
2. Materials and Methods
2.1 Collection of urine samples
Urine samples were collected from sows that aborted within 24 hrs. A record that illustrated cases of abortion in sows during the period of study was used as reference to track animals as source of urine samples (Table 1). Urine was collected through the free-catch method which used clean plastic wide-mouth canisters that were connected to a 1.5 m flat aluminum rod which was held close to the vestibule of the urinating sow. The method of urine collection was made in compliance with the guidelines of the CLSU Institutional Animal Care and Use Committee (Registration Number LAF-0013/ LAF-007) in handling animals for research. The samples were held in polystyrene box packed with ice and transported to the Microbiology Laboratory of the CLSU College of Veterinary Science and Medicine for the succeeding experiments.
|
Species of aborted livestock |
Number of abortion cases |
Identified bacteria in recorded cases |
|
Caracows |
5 |
Brucella spp. (3) |
|
Cows |
0 |
- |
|
*Sows |
50 |
Leptospira hyos (25); Leptospira spp. (5); Leptospira spp. (5) |
|
Does |
15 |
Brucella spp. (2); Leptospira spp. (10) |
|
Total |
70 |
- |
*Highest number of abortion cases is recorded in sows.
Table 1: Record of abortion cases in livestock in the first 6 months of study.
2.2 Isolation, cultivation and microscopic evaluation of Leptospira spp.
Urine samples were placed in replicated tubes that contained freshly-prepared Fletcher’s medium supplemented with 8% rabbit serum. The samples were kept in an incubator at 31ºC for 24 hours. The growth of Leptospira spp. in the culture medium was indicated by the presence a dinger zone within the period of incubation. Small aliquots (15 to 20 µL) of samples were obtained from tubes and placed as smears on slides. The slides were stained applying the procedures of others [2] with slight modifications. The smears were fixed in 1% formalin before adding 5% Sodium bicarbonate and 10 drops of basic fuchsin, allowing the stain to adhere on the smear for 5 minutes at room temperature before washing the slides with water. The slides were allowed to dry before microscopic examination.
2.3 DNA extraction, PCR amplification and DNA sequencing of the Leptospira spp. Isolate
Samples of Leptospira spp. grown in Fletcher’s medium were centrifuged at 12,000 rpm for 2 minutes to collect the bacterial sediments. The nuclei lysis solution (600 µL) was added to the samples on tubes, mixed then incubated for 5 minutes at 80°C before cooling. A volume of 3 µL RNase solution was added to the samples before incubation at 37°C for 15 to 60 minutes. The samples were allowed to cool down after incubation then 200 µL protein precipitate solution was added. The samples were incubated on ice for 5 minutes then centrifuged at 12,000 rpm for 2 minutes. The supernatant samples were collected after centrifugation then transferred to clean micro-centrifuge tubes that contained 600 µL iso-propanol. The samples were again centrifuged at 12,000 rpm for 2 minutes to collect the sediments. A volume of 600 µL 70% ethanol was added to the samples then centrifuged again at 12,000 rpm for 2 min. The ethanol layer was discarded while the sediments were air-dried. Replicates of DNA samples from bacterial isolates were stored in aliquots at -80ºC and used as DNA for PCR evaluation. A pair of primers that recognize the 16S rRNA gene of bacteria (NF and NR) previously employed by other researchers [3] and [4] were used in the PCR amplification of the isolated Leptospira spp. (Table 2).
|
Primers |
Sequences |
|
NF |
5’ GGCGGCAKGCCTAYACATGCAAGT 3’ |
|
NR |
5’ GACGACAGCCATGCASCACCTGT 3’ |
(Carroll et al., 2000; and Garcia et al., 2013).
Table 2: Pair of primers applied in the amplification of the 16S rRNA gene.
A volume of 100 µL DNA rehydration solution was added to DNA samples before PCR. For the amplification of Leptospira spp., each reaction mixture contained 1 µL 10X PCR Buffer (Takara, Japan), 0.5 µL MgCl2, 1 µL dNTP (Takara, Japan), 0.1 µL Taq polymerase (Takara, Japan), 0.5 µL forward primer, 0.5 µL reverse primer, 5.4 µL sterile deionized water (Thermo Fisher Scientific, USA) to make up to a total volume of 10 µL before adding 1 µL DNA.
The PCR conditions involved initial denaturation at 94°C for 3 minutes, 35 cycles of denaturation at 94°C for 30 seconds, annealing at 54°C for 30 seconds, extension at 72°C for 1 minutes, final extension at 72°C for 8 minutes and a final holding temperature of 4°C. The PCR products were purified and used as a sample for DNA sequencing. The sequences were aligned and used to search for identical Leptospira spp. in the gene bank (NCBI). A request for the annotation of the aligned sequences in NCBI was made thereafter.
2.4 In-vitro evaluation on the susceptibility of Leptospira hyos to a panel of antibiotics
The identified L. hyos isolate was used as the test organism in the evaluation of antibiotic susceptibility. Stock solutions of the five antibiotics (Cephalexin, Neomycin, Clindamycin, Ciprofloxacin and Sulfamethoxazole) were prepared in sterile distilled water at concentrations of 1 mg/mL at the start of the experiment. Antibiotic concentrations at 25 µg, 35 µg and 50 µg per mL of each of the five antibiotics were separately obtained from the stock solutions. For each antibiotic concentration described, the antibiotic was added correspondingly to three replicated vials that contained 1 mL Fletcher’s medium with a fresh culture of L. hyos suspension (4.68 + 0.47 MFU).
The inability of Leptospira spp. to form colonies on the surface of a medium required the need to evaluate cell density by means of McFarland standards. At the start of this experiment, cell suspensions of L. hyos were obtained from replicated tubes and centrifuged at 12,000 x g for 2 minutes. The sediments were saved, re-suspended in 5 mL sterile deionized water (Thermo Fisher Scientific, USA) and McFarland readings were obtained before application of the antibiotics. After the 6- and 12-hrs interaction period with the antibiotics, cell suspensions of L. hyos were obtained from replicated tubes and centrifuged at 12,000 x g for 2 minutes. The sediments were saved, re-suspended in 5 mL sterile deionized water (Thermo Fisher Scientific, USA) before taking the McFarland readings. Comparison of McFarland readings taken before antibiotic application were used as reference in comparing the cell density of L. hyos recovered after exposure to the antibiotics.
2.5 Statistical analysis
McFarland readings relative to the application of the 3 different concentrations of the 5 different antibiotics were expressed as mean McFarland units (MFU) of 3 replicates. Differences in MFUs as an effect of antibiotic concentrations were statistically analyzed using LSD (Least significant differences) while differences in MFUs across time intervals (6 and 12 hr) within treatments were analyzed by Students’ T-test (P< 0.01).
3. Results and Discussion
3.1 Morphology of the Leptospira spp. Isolate
The isolate of Leptospira spp. from aborted sows appeared short with S-shaped or curved ends that stain dark pink to purple on microscopic evaluation (Figure 1). The length ranged from 5.0 to 10.0 microns (mean measurement 7.91 + 1.70 microns) and the width ranged from 0.297 to 0.445 micron (mean measurement 0.376 + 0.05 micron).
3.2 Amplification of the 16S rRNA gene and identification of the Leptospira spp. from aborted sows
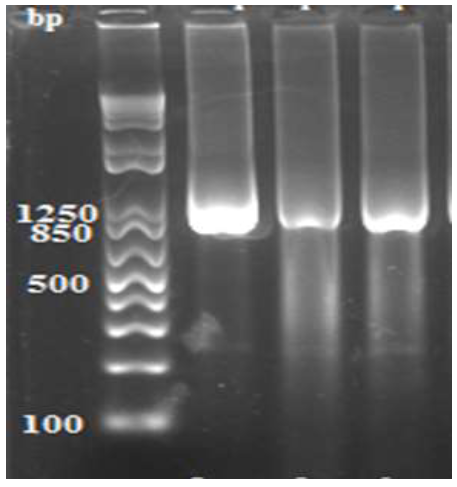
Data in Figure 2 demonstrate the amplified products of the 16S rRNA genes of three (3) Leptospira spp. isolates recovered from aborted sow. The 3 amplified products registered a molecular size that ranged from 1050 to 1080 bp.
Figure 2: Amplified products of the 16S rRNA gene of 3 isolates of Leptospira spp. from aborted sows showing DNA fragments with sizes that vary from 1050 to 1080 bp. Lane 1 (Molecular weight marker). Lane 2 (Leptospira spp. from aborted sow, isolate MK629955), Lane 3 (Leptospira spp. from aborted sow, isolate MK629956), Lane 4 (Leptospira spp. from aborted sow, isolate MK629957).
Tables 3, 4 and 5 show the DNA sequences of the 16S rRNA genes of 3 sow-borne Leptospira organisms. The identified pathogens possess 16S rRNA genes that are composed of 1,082 (Accession number MK629955) and 1,058 nucleotides (Accession numbers MK629956 and MK629957) and are currently annotated in the gene bank as L. hyos (Accession number MK629955) and 2 Leptospira spp. (Accession numbers MK629956 and MK629957).
Aligned sequence of the 16S rRNA gene of Leptospira hyos with the given accession number
Table 3: Genomic sequence of the 16S rRNA gene of Leptospira hyos.
Aligned sequence of the 16S rRNA gene of Leptospira spp. with the given accession number.
Table 4: Genomic sequence of the 16S rRNA gene of Leptospira spp.
Aligned sequence of the 16S rRNA gene of Leptospira spp. with the given accession number.
Table 5: Genomic sequence of the 16S rRNA gene of Leptospira spp.
3.3 In-vitro susceptibility of the isolated Leptospira hyos to antibiotics
Table 6 gives a summary of the cell density of L. hyos expressed as McFarland units (MFU) in response to the applied concentrations of antibiotics. In the first 6 hours, higher L. hyos MFU was associated with the application of 25 µg cephalexin compared to lower MFUs obtained with higher (35 µg and 25 µg) concentrations. The susceptibility of L. hyos to the cell membrane inhibitor neomycin was also marked by higher MFU relative to the application of 25 µg neomycin in contrast to the comparably lower MFUs derived from the application of 35 and 50 µg. Higher MFU of L. hyos was also associated with the application of 25 µg protein synthesis inhibitor clindamycin in contrast to the significantly lower MFUs obtained with the application of 35 and 50 µg. Higher MFU of L. hyos was linked to the application of nucleic acid synthesis inhibitor ciprofloxacin at 25 µg concentration and nil MFUs related to the application of 35 and 50 µg concentration. Higher MFU of L. hyos was attained with the use of 25 µg folate co-factor synthesis inhibitor sulfamethoxazole contrary to significantly lower MFUs associated with the application of 35 and 50 µg concentrations. The foregoing data explain the significant effect of antibiotic concentrations on the susceptibility of L. hyos to cephalexin, neomycin, clindamycin, ciprofloxacin and sulfamethoxazole measured in terms of MFU after 6 hours exposure time.
|
Antibiotics |
Concentrations (µg/mL) |
Duration of Interaction |
|
|
6 hrs |
12 hrs |
||
|
Cephalexin |
25 |
4.00 (0.00) a,y |
3.33 (0.47) a,z |
|
35 |
3.00 (0.00) b,y |
0.83 (0.23) b,z |
|
|
50 |
0.83 (0.23) c,y |
0.50 (0.00) c,z |
|
|
Neomycin |
25 |
0.83 (0.23) a,y |
0.66 (0.23) a,y |
|
35 |
0.66 (0.23) b,y |
0.00 (0.00) b,z |
|
|
50 |
0.50 (0.00) bc,y |
0.00 (0.00) b,z |
|
|
Clindamycin |
25 |
2.33 (0.47) a,y |
1.00 (0.00) a,z |
|
35 |
1.66 (0.47) b,y |
1.00 (0.00) a,z |
|
|
50 |
0.66 (0.23) c,y |
0.50 (0.00) b,yz |
|
|
Ciprofloxacin |
25 |
2.00 (0.00) a,y |
1.00 (0.00) a,z |
|
35 |
0.00 (0.00) b,y |
0.00 (0.00) b,y |
|
|
50 |
0.00 (0.00) b,y |
0.00 (0.00) b,y |
|
|
Sulfamethoxazole |
25 |
3.00 (0.82) a,y |
2.00 (0.00) a,z |
|
35 |
1.33 (0.47) b,y |
0.83 (0.23) b,z |
|
|
50 |
0.00 (0.00) c,y |
0.00 (0.00) c,y |
|
|
Fletcher’s medium (Reference control) |
4.66 (0.47) y |
4.33 (0.47) y |
|
Values represent the mean (± standard deviation) cell density of Leptospira hyos expressed as McFarland units (MFU) recovered after the application of different concentrations of antibiotics at specified duration of interaction. MFU of 0.5 is equivalent to 1.5 x 108 CFU/ml; 1.0 MFU/3.0 x 108 CFU/ml; 2.0 MFU/6.0 x 108 CFU/ml; 3.0 MFU/9.0 x 108 CFU/ml; 4.0 MFU/12.0 x 108 CFU/ml; 5.0 MFU/15 x 108 CFU/ml; 6.0 MFU/18.0 x 108 CFU/ml; 7.0 MFU/21.0 x 108 CFU/ml; 8.0 MFU/24.0 x 108 CFU/ml; 9.0 MFU/27.0 x 108 CFU/ml; and 10.0 MFU/30.0 x 108 CFU/ml. a, b, c (significant differences in cell density as an effect of the concentrations of each antibiotic in a given time point, P<0.01). y, z (significant differences in cell density as an effect of the duration of interaction between pathogen and the drugs (6- and 12-hour, P<0.01).
Table 6: Cell density of Leptospira hyos (MFU) in response to its exposure to the different antibiotic concentrations at indicated durations of interaction.
Data in Table 6 also show the responses of L. hyos exposed to the different concentrations of antibiotics in the next 6 hours (12th hour). Higher L. hyos MFU was associated with the application of 25 µg of cephalexin compared to lower MFUs obtained with higher cephalexin (35 µg and 25 µg) concentrations. The susceptibility of L. hyos to neomycin was also marked by higher MFU relative to the application of 25 µg neomycin in contrast to the nil MFUs derived from the application of 35 and 50 µg. Comparably higher MFUs of L. hyos were also linked with the application of 25 and 35 µg clindamycin in contrast to the significantly lower MFU obtained with the application of 50 µg. Higher MFU of L. hyos was associated with the application of ciprofloxacin at 25 µg concentration and nil MFUs related to the application of 35 and 50 µg. Higher MFU of L. hyos was attained with the use of 25 µg sulfamethoxazole contrary to significantly lower MFU associated with the application of 35 µg and nil MFU with 50 µg sulfamethoxazole.
The foregoing data further show the significant effect of antibiotic concentrations on the susceptibility of L. hyos to cephalexin, neomycin, clindamycin, ciprofloxacin and sulfamethoxazole measured in terms of MFU after the 12-hour exposure time.
Data taken on the 12-hour also demonstrate a significant difference in the reduction of cell densities of L. hyos relative to the application of the 3 concentrations of Cephalexin compared to those data obtained in the 6th hour-exposure time. The application of 25 µg Neomycin caused significant reduction of L. hyos MFU noted after 12-hour in contrast to data obtained on the 6-hour while nil MFUs were associated with the higher neomycin concentrations (35 and 50 µg). Significant declines in MFUs of L. hyos were comparably associated with the application of 25 and 35 µg Clindamycin after 12 hours exposure compared to the 6-hour data while comparably low MFUs were related with 50 µg Clindamycin noted after 6- and 12-hour exposure time. The application of 25 µg Ciprofloxacin caused significant reduction in the MFU of L. hyos after 12 hours compared to data recorded on the 6 hours while comparable nil MFUs were associated with the application of 35 and 50 µg Ciprofloxacin recorded after 6- and 12-hour exposure time.
The use of 25 µg Sulfamethoxazole contributed to significant reduction in MFU of L. hyos after 12-hour exposure time compared to the data after 6 hours. The application of the 35 µg Sulfamethoxazole also caused significant reduction in MFU of L. hyos after 12 hours exposure time compared to the data after 6 hours while nil MFUs were related with the use of 50 µg Sulfamethoxazole in the two periods of obser-vation.
4. Discussion
The use of contemporary technics in molecular biology undertaken by other researchers to identify pathogens in various infections of humans and animals [3] and [4] were applied to validate the involvement of the pathogen in abortion cases of swine. Basic techniques in bacterial cultivation were also applied to come up with a special medium for the fastidious Leptospira spp. which made isolation congruent with other intended methodologies. The identification of Leptospira spp. during the period of study prompted the need to screen for its susceptibility to antibiotics that can be recommended for treatment or as a control strategy against further infections that may set in after an incidence of abortion.
The study involved evaluation of the responses of the identified abortion-associated L. hyos to a panel of antibiotics composed of cephalexin, neomycin, clindamycin, ciprofloxacin and sulfamethoxazole by monitoring bacterial cell densities post-exposure with the antibiotics. It has been established that the susceptibility of L. hyos to antibiotics is an effect of both antibiotic concentration and duration of exposure to the antibiotics. It has also been demonstrated that high antibiotic concentrations were related to low cell densities pointing toward a correspondingly greater microbial death as a result of antibiotic application.
The antibacterial effects of the antibiotics varied as demonstrated by the bactericidal action of 25 µg Ciprofloxacin; and 25 and 35 µg Sulfamethoxazole against L. hyos attained as early as 6-hour exposure time while higher Neomycin, Ciprofloxacin (35 and 50 µg) and Sulfamethoxazole (50 µg) concentrations were required to sustain their bactericidal action against the pathogen until 12 hours. These data altogether define the minimum inhibitory concentrations (MICs) of Cephalexin, Neomycin, Clindamycin, Ciprofloxacin and Sulfamethoxazole at 25 µg concentration.
Other researchers have attempted to screen the sensitivity of Leptospira spp. to antibiotics. The efficacy of first generation cephalosporins against Leptospira spp. was evaluated through microdilution method by others [5]. The efficacy of neomycin as an antimicrobial was tested by another researcher [6] in a study where the antibiotic was applied in combination with furazolidone as the control of contaminants in cultures of Leptospira spp.. Combined furazolidone and neomycin each at 5 µg/mL concentrations in Fletcher‘s medium or in implanted sensitivity disks at 50 µg and 10 µg respectively, was reportedly effective in inhibiting contaminants and did not interfere with the growth of different serotypes of Leptospira. In a separate study conducted by another group of researchers [7] in 2015, it was reported that Leptospira spp. are susceptible to ciprofloxacin and clindamycin on as a simple screening method for epidemiological surveillance of Leptospira spp.
Another in-vitro study reportedly evaluated the susceptibility of 46 Leptospira strains isolated from rats in the Philippines to 14 antimicrobial agents [8] which identified Leptospira strains that were resistant to amphotericin B, 5-fluorouracil, fosfomycin, sulfamethoxazole and trimethoprim. The study of another group of reseachers [9] tested antimicrobial susceptibility of Leptospira spp. isolated from environmental, human and animal sources in Malaysia and reported resistance of the pathogen to trimethoprim and sulfamethoxazole.
The present study describes the isolation of Leptospira spp. obtained from clinical cases of abortion in swine, different from the varied sources of Leptospira spp. described and evaluated to serve the purposes of the aforementioned researchers. This report upholds the importance of the data that define the susceptibility of the abortion-associated L. hyos and related Leptospira isolates to antibiotics, as this serves as a relevant information for veterinary practitioners to identify an effective line of treatment against Leptospira infections.
5. Conclusion
The identity of the abortion-associated L. hyos and the data that defined the susceptibility of the pathogen to the antibiotics with different mechanisms of action have been addressed. The significant reduction of cell densities relative to the application of antibiotics at described time points, and identification of the MICs of each antibiotic provide a basis in recommending a line of treatment against L. hyos infections in swine.
Acknowledgement
This work was undertaken with the assistance of the Biotechnology Office of the Philippine Department of Agriculture and the College of Veterinary Science and Medicine, Central Luzon State University, Philippines.
Conflict of Interest
No conflict of interest exists among the authors.
References
- Suputtamongkol Y, Niwattayakul K, Suttinont C, et al. An open, randomized, controlled trial of penicillin, doxycycline and cefotaxime for patients with severe leptospirosis. Clinical Infectious Diseases 10 (2015):1417-1424.
- Ryu E. A simple method for staining Leptospira and Treponema. Japanese Journal of Microbiology 2 (1963): 81-85.
- Carroll NM, Jaeger EEM, Choudhury S, et al. Detection and discrimination between Gram-positive and Gram-negative bacteri in intra-ocular samples using nested PCR. Journal of Clinical Microbiology 38 (2000): 1753-1757.
- Garcia GG, Belotindos L, Mingala CN. Molecular characterization of respiratory bacterial pathogens in large and small ruminants. The Thai Journal of Veterinary Medicine 4 (2013): 483-489.
- Harris B, Blatz P, Hinkle M, et al. In vitro and in vivo activity of first generation cephalosporins against Leptospira. The American Journal of Tropical Medicine and Hygiene 85 (2011): 905-908.
- Myers DM. Efficacy of combined furazolidone and neomycin in the control of contamination in Leptospira cultures. Antimicrobial Agents and Chemotherapy 5 (1975) 666-671.
- Wuthiekanun V, Amornchai P, Langla S, et al. Antimicrobial disk susceptibility testing of Leptospira spp. using Vanaporn Wuthiekanun (LVW) Agar. American Journal of Tropical Medicine and Hygiene 2 (2015): 241-243.
- Chakraborty A, Miyahara S, Villanueva SY, et al. In vitro sensitivity and resistance of 46 Leptospira strains isolated from rats in the Philippines to 14 antimicrobial agents. Antimicrobial Agents and Chemotherapy 12 (2010): 5403-5405.
- Benacer D, Zain SNM, Ooi PT, et al. Antimicrobial susceptibility of Leptospira spp. isolated from environmental, human and animal sources in Malaysia. Indian Journal of Medical Microbiology 35 (2017): 124-128.


 Impact Factor: * 1.1
Impact Factor: * 1.1 Acceptance Rate: 80.20%
Acceptance Rate: 80.20%  Time to first decision: 10.4 days
Time to first decision: 10.4 days  Time from article received to acceptance: 2-3 weeks
Time from article received to acceptance: 2-3 weeks 