In-Silico Analysis of Osmotic Stress Effects on Capsicum annuumTranscriptome
Sai Batchu
The College of New Jersey, Biology Department 2000 Pennington Rd. Ewing, NJ, USA
*Corresponding Author: Sai Batchu, The College of New Jersey, Biology Department 2000 Pennington Rd. Ewing, NJ, United States
Received: 18 February 2020; Accepted: 18 March 2020; Published: 20 March 2020
Article Information
Citation: Sai Batchu. In-Silico Analysis of Osmotic Stress Effects on Capsicum annuum Transcriptome. Journal of Bioinformatics and Systems Biology 3 (2020): 019-031.
View / Download Pdf Share at FacebookAbstract
Osmotic stress is a major abiotic limitation affecting yield and growth of crops worldwide, of which include one of the most cultivated bell peppers, Capsicum annuum. To understand the perturbations on gene expression in Capsicum annuum under osmotic stress, the present study compared RNA-sequencing data between osmotic stress-induced plants and mock controls. A total of 2879 genes were found significantly differentially expressed. Differentially expressed genes related to protein processing and hormone signaling exhibited increased expression in plants experiencing osmotic stress. In addition, differentially expressed genes associated with ribosomal proteins and ribosomal biogenesis showed decreased expression patterns under osmotic stress. These findings provide basis for further elucidating the coordinated transcriptional processes underlying response to abiotic stresses in Capsicum annuum.
Keywords
<p>Transcriptome; Capsicum annuum; Osmotic stress; In-silico</p>
Article Details
Introduction
Capsicum annuum L., an evergreen perennial originating from the Americas, is a widely cultivated crop from the Capsicum genus. The success of C. annuum as an economically important plant is attributed to its ability to be cultivated around the world as different varieties, contributing to distinct taste profiles, phenotypes, and colloquialisms (e.g. bell pepper, chili pepper, cayenne, poblano) [1].
Pepper cultivation is often influenced by various environmental factors, including temperature, drought, and salinity [2]. Of importance are drought and salinity, both of which contribute to osmotic stress. Osmotic stress, or osmotic shock, is a type of abiotic stress that induces decreased water potential in a plant. Sudden increases in solute concentrations in a plant’s environment, as observed in drought-affected and high-salinity environments, trigger movement of water out of the plant cell [3].
Global climate change, including severe droughts and increasing soil salinity, is introducing mounting challenges to food security [4]. Nonetheless, global warming poses as an imminent challenge for pepper development and maturation [5]. Although there have been recent advances in developing osmotic stress-resistant varieties C. annuum, the molecular mechanisms and gene regulation underlying response and resistance to osmotic stress is poorly understood [6, 7].
In this study, publicly available RNA-sequencing data of leaves from C. annuum experiencing osmotic shock were analyzed to investigate the transcriptional changes in the crop’s response to this abiotic stressor. This comparative transcriptomic analysis expands our knowledge and sets the stage for developing osmotic stress-robust varieties of C. annuum.
Materials and Methods
RNA-Sequencing Data Acquisition
Publicly available leaf transcriptome datasets were downloaded from the NCBI Sequence Read Archive under accession numbers SRR8692601, SRR8692602, SRR8692596 for three replicates of abiotic stress mock-induced replicates and SRR8692617, SRR8692630, SRR8692581 for three replicates of osmotic stress-induced replicates through mannitol treatment [8]. All replicates can be found under BioProject accession PRJNA525913.
Read Mapping and Normalization
Sequences were filtered with Trim Galore (https://github.com/FelixKrueger/TrimGalore) to remove low quality reads and adaptor sequences. FastQC [9] was used to determine final quality of reads. Cleaned reads were mapped to indexed Capsicum annuum genome assembly Pepper Zunla 1 version 1 [10] with HISAT2 aligner run with default parameters [11]. Alignment output was assembled and quantified using StringTie [12]. DESeq2 [13], an R software package [14], was used for gene count data normalization.
Differential Expression Analysis
Genes were considered differentially expressed if log2 fold change < -1 or log2 fold change > 1 with an adjusted p value < 0.05. Gene enrichment analysis for biological processes was performed using available Arabidopsis orthologs of tobacco genes with R package ‘KEGGREST’ through the Kyoto Encyclopedia of Genes and Genomes (KEGG) with Wilcoxon rank-sum test to evaluate the statistical significance of annotated pathways [15]. Pathways were visualized with ‘pathview’ package [16]. Scatterplot matrix and volcano plot were generated with R packages ‘vidger’ [17] and ‘EnhancedVolcano’ [18], respectively. Additional figures were generated with R package ggplot2 [19].
Results
Quality checks
To confirm the quality of RNA-sequencing data, gene abundances of independent individual runs from each sample were compared (Figure 1B). Further quality checks provided gene density (Figure 1A) and a scatterplot matrix (Figure 1C). The percentage of mapped reads to the C. annuum genome was determined from HISAT2 alignment summaries (Figure 1D). These sequencing and mapping quality checks suggested that the data qualified for further analysis.
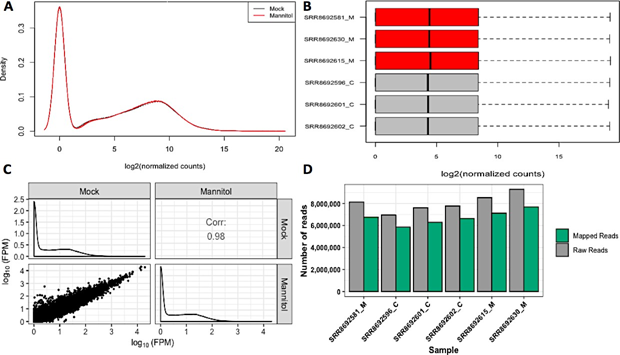
Figure 1: RNA-sequencing quality check and mapping efficiency with C. annuum genome.
Panels A, B, and C show data amongst replicates in terms of gene density, distribution of read counts, and non-summarized read counts for all genes. Panel D displays mapped reads compared to raw sequencing reads where x axis indicates samples used in the study and y axis indicates number of reads.
Differential expression analyses
Normalization resulted in 2879 genes that were differentially expressed between mock control samples and plants induced for osmotic shock (Figure 2B). Plotting the top two principal components revealed a clustering trend (Figure 2A).
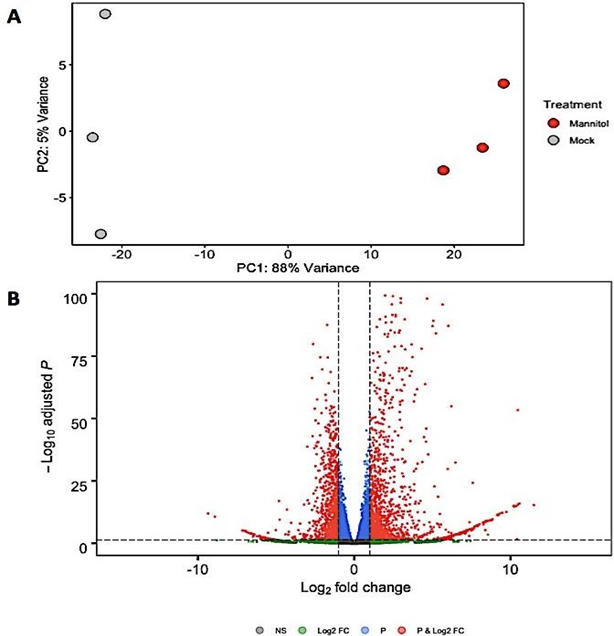
Figure 2(A): Principle component analysis showing the distance of variance among all 6 samples. (B) Volcano plot depicting number of differentially expressed genes based on their adjusted p values and log2 fold change in the analysis between osmotic stress-induced and mock samples. Horizontal dashed black line signals statistical significance threshold (adjusted P £ 0.05). Two vertical dashed black lines show the threshold of fold change (log2FC > 1 or < -1).
Further analysis revealed decreased expression of genes encoding late embryogenesis abundant (LEA) proteins (Figure 3).
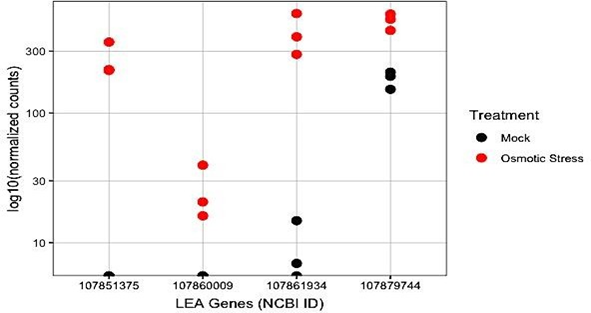
Figure 3: Normalized counts of genes encoding late embryogenesis abundant (LEA) proteins between the two treatments. Genes are labeled by their NCBI (National Center for Biotechnology Information) identifications.
Gene enrichment analysis revealed ribosome and ribosomal biogenesis as significantly down- regulated pathways (Figure 4; Figure 5; Figure 6). The pathway for protein processing in the endoplasmic reticulum was annotated for a plethora of differentially expressed genes and showed diverse differential expression patterns (Figure 7). Plant hormone signal transduction pathways were also affected in the osmotic stress condition, with many genes in different signaling pathways being up-regulated (Figure 8).
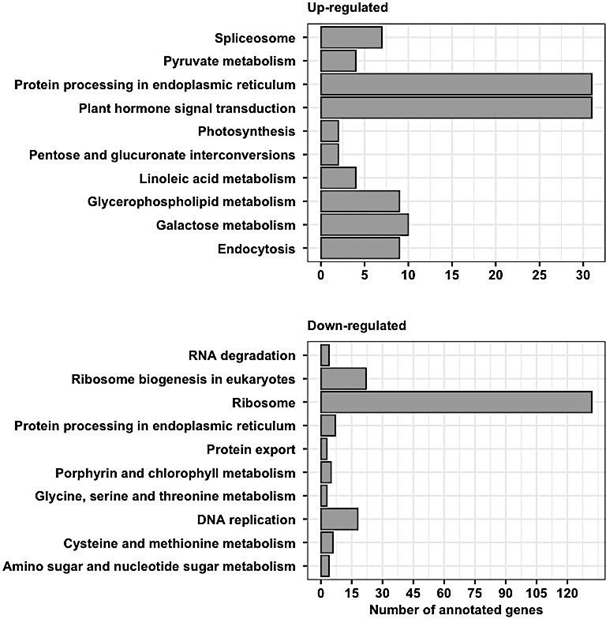
Figure 4(A): Gene ontology pathways enriched with over-expressed genes in osmotic stress induction. (B) Gene ontology pathways enriched with under-expressed genes in osmotic stress induction.
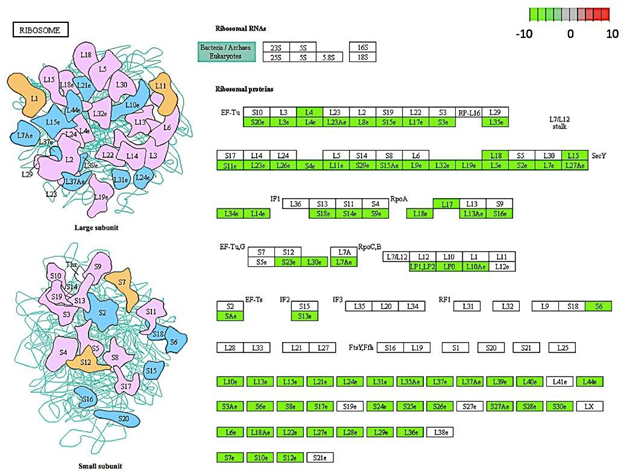
Figure 5: Genes encoding for ribosomal proteins down-regulated in C. annuum undergoing osmotic stress. Scale is log2 fold change compared to mock control samples.
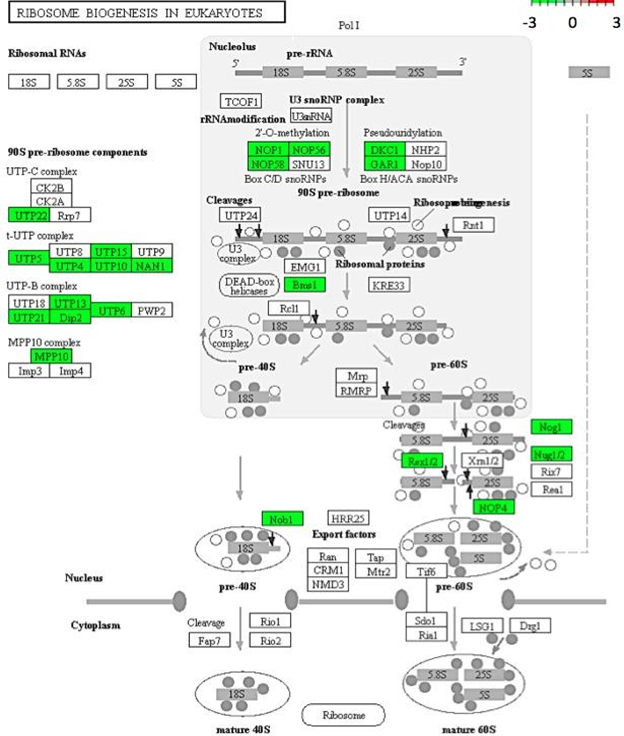
Figure 6: Genes encoding for ribosome biosynthesis down-regulated in C. annuum undergoing osmotic stress. Scale is log2 fold change compared to mock control samples.
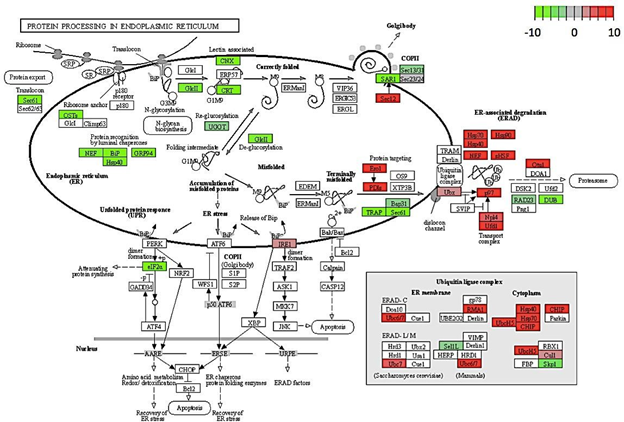
Figure 7: Genes encoding for protein processing showing differential expression in C. annuum undergoing osmotic stress. Scale is log2 fold change compared to mock control samples.
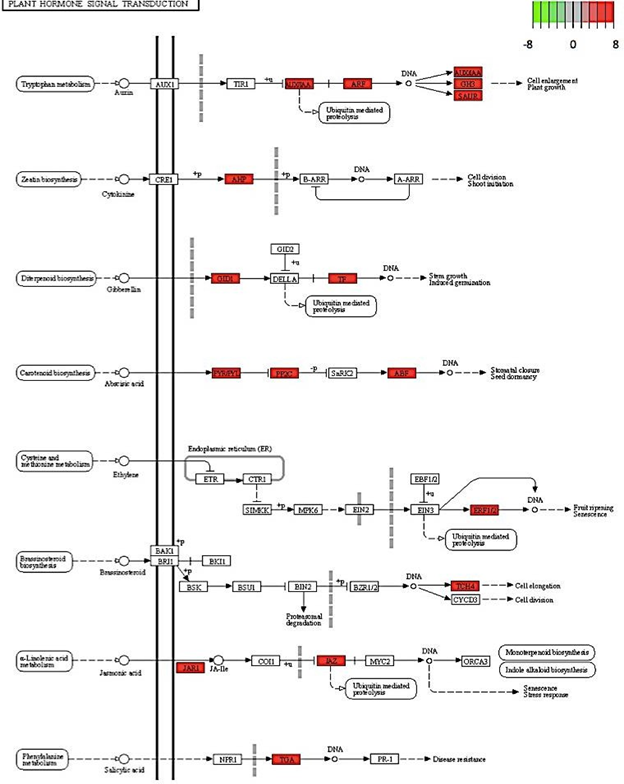
Figure 8: Genes involved in various plant hormone signal transduction pathways down- regulated in C. annuum undergoing osmotic stress. Scale is log2 fold change compared to mock control samples.
Discussion
Scrutinizing up regulated genes in osmotic stress treatment group showed that numerous genes encoding late embryogenesis abundant proteins were significantly increased in their expression levels. LEA proteins were first characterized during the maturation drying process of embryo development. Many of these proteins have been classified as ‘hydrophilins,’ a family of hydrophilic proteins [20]. LEA proteins have been shown to accumulate in vegetative tissue of plants inhabiting water-deficit environments [20]. The have also been implicated with desiccation tolerant structures and stages of plant development [21,22]. Under water deficits, crucial enzymes suffer conformational changes that lead to decreased activity or denaturation. It is hypothesized that LEA proteins prevent the conformational changes of critical enzymes during these stress conditions [23, 24]. Genes encoding these proteins may serve as an attractive basis for developing varieties resilient in osmotic stress-affected environments.
Gene enrichment analysis was used to acquire a global view of gene expression perturbations. Of the top significantly down-regulated pathways, ribosomal proteins and ribosomal biogenesis were two that were annotated with the most genes (Figure 4). Although the literature focused on these pathways in plants under stress is limited, it has been documented in yeast that exposure to abiotic stress factors causes a reduction the ribosome abundance. This is thought to be a survival mechanism since the major source of toxicity derived from misfolding of newly synthesized proteins, such as ribosomes. Decreasing ribosomal production would decrease the amount of misfolded proteins and act as an effective stress response [25]. A similar mechanism may play a part in the response of C. annuum leaf cells against osmotic stress.
This hypothesis may be further supported by the fact that certain genes in the endoplasmic reticulum (ER) protein-processing pathway were up regulated (Figure 7). Genes encoding heat shock proteins (Hsp), such as Hsp70, Hsp90, and Hsp40, showed increased expression under osmotic stress. These proteins play important roles in proper folding and degradation of proteins [26].
In addition, several hormone-signaling pathways were up regulated, including carotenoid biosynthesis and tryptophan metabolisms. These findings are consistent with recent literature results. For example, in model plant species, Arabidopsis thaliana, carotenoid biosynthesis was up regulated in response to osmotic stress [27]. It is crucial to note that an important product of the carotenoid biosynthetic pathway is hormone abscisic acid (ABA) and it is well documented that ABA protects the xanthophyll cycle and the photosynthetic systems from oxidative stress [28]. In addition, ABA triggers the closing of stomata when soil water is insufficient for transpiration [28].
This study analyzed the leaf transcriptome of Capsicum annuum undergoing osmotic stress. The results conclude that there is an increased regulation of major stress response pathways.
Although this study was limited to an in silico perspective, the putative findings establish a basis for additional investigations and validation experiments to clarify the transcriptomic changes in response to abiotic stress.
Acknowledgements
Special thanks to Electronic Laboratory for Science and Analysis (ELSA) High Performance Computing cluster provided by The College of New Jersey for enabling the analysis of this high- throughput data.
Conflicts of Interest
The author declares no conflict of interest.
References
- Moscone EA, Scaldaferro MA, Grabiele M, Cecchini NM, Sánchez García Y, et al. The evolution of chili peppers (Capsicum – Solanaceae): A cytogenetic perspective. Acta Hortic 745 (2007): 137–170.
- Erickson AN, Markhart AH. Flower developmental stage and organ sensitivity of bell pepper (Capsicum annuum L.) to elevated temperature. Plant Cell Environ 25 (2002): 123–130.
- Hoffmann S. Osmotic stress signaling and osmoadaptation in yeasts. Microbiol Mol Biol Rev 66 (2002): 300–372.
- Ashour EK, Al-Najar H. The Impact of Climate Change and Soil Salinity in Irrigation Water Demand in the Gaza Strip. J Earth Sci Climate Change 3 (2012): 120.
- Hedhly A, Hormaza JI, Herrero M. Global warming and sexual plant reproduction. Trends Plant Sci 14 (2009): 30–36.
- Bojórquez-Quintal E, Velarde-Buendía A, Ku-González Á, Carillo-Pech M, Ortega- Camacho D, et al. Mechanisms of salt tolerance in habanero pepper plants (Capsicum chinense Jacq.): Proline accumulation, ions dynamics and sodium root-shoot partition and compartmentation. Frontiers in Plant Science 5 (2014): 605.
- Sahitya UL, Krishna MSR, Suneetha P. Integrated approaches to study the drought tolerance mechanism in hot pepper (Capsicum annuum L.). Physiol Mol Biol Plants 25 (2019): 637.
- Kang W, Sim YM, Koo N, Nam JY, Lee J, Kim N, Jang H, Kim YM, Yeom SI. Transcriptome profiling of abiotic responses to heat, cold, salt, and osmotic stress of Capsicum annuum L. Scientific Data 7 (2020):17.
- Andrews S. FastQC: a quality control tool for high throughput sequence data (2010). Available online at: http://www.bioinformatics.babdraham.ac.uk/projects/fastqc.
- Qin C, Yu C, Shen Y, Fang X, Chen L, et al. Whole-genome sequencing of cultivated and wild peppers provides insights into Capsicum domestication and specialization. Proc Natl Acad Sci USA 111 (2014): 5135-5140.
- Kim D, Langmead B, Salzberg SL. HISAT: a fast spliced aligner with low memory requirements. Nature Methods 12 (2015): 357-60.
- Pertea M, Pertea GM, Antonescu MA, Chang T, Mendell JT, Salzberg SL. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nature Biotechnology 33 (2015): 290-295.
- Love M, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biology 15 (2014): 550.
- R Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria (2013).
- Tenenbaum D. KEGGREST: Client-side REST access to KEGG (2019).
- Luo W, Brouwer C. Pathview: an R/Bioconductor package for pathway-based data integration and visualization. Bioinformatics 29 (2013): 1830-1831.
- Monier B, McDermaid A, Jing Z, Qin M. vidger: Create rapid visualizations of RNAseq data in R. (2019) : https://github.com/btmonier/vidger.
- Blighe K. EnhancedVolcano: Publication-ready volcano plots with enhanced colouring and labeling. R package version 1.2.0 (2019).
- Wickham H. ggplot2: Elegant Graphics for Data Analysis. Springer-Verlag New York (2016).
- McCubin WD, Kay CM, Lane BG. Hydrodynamics and optical properties of the wheat Em protein. Can J Biochem Cell Biol 63 (1985): 803–811.
- Hundertmark M, Hincha DK. LEA (Late Embryogenesis Abundant) proteins and their encoding genes in Arabidopsis thaliana. BMC Genomics 9 (2008): 118–139.
- Shih M, Hoekstra FA, Hsing YC. Late Embryogenesis Abundant Proteins. Advanc Bot Res 48 (2008):211–255.
- Goyal K, Tisi L, Basran A, Browne J, Burnell A, Zurdo J. Transition from natively unfolded to folded state induced by desiccation in an anhydrobiotic nematode protein. J Biol Chem. 12 (2003): 977–12984.
- Reyes JL, Rodrigo MJ, Colmenero-Flores JM, Gil JV, Salamini F, Bartels D. Hydrophilins from distant organisms can protect enzymatic activities from water limitation effects in vitro. Plant Cell Environ. 28 (2005): 709–718.
- Guerra-Moreno A, Isasa M, Bhanu MK, et al. Proteomic Analysis Identifies Ribosome Reduction as an Effective Proteotoxic Stress Response. J Biol Chem 50 (2015): 29695– 29706.
- Lackie RE, Meciejewski A, Ostapchenko V, Marque-Lopes J, Choy WY, Duennwald ML, Prado VF, Prado MAM. The Hsp70/Hsp90 Chaperone Machinery in Neurodegenerative Diseases. Frontiers in Neuroscience 11 (2017): 254.
- Meier S, Tzfadia O, Vallabhaneni R, Gehring C, Wurtzel ET. A transcriptional analysis of carotenoid, chlorophyll and plastidial isoprenoid biosynthesis genes during development and osmotic stress responses in Arabidopsis thaliana. BMC Syst Biol. 5 (2011): 77.
- Barickman TC, Kopsell DA, Sams CE. Abscisic Acid Increases Carotenoid and Chlorophyll Concentrations in Leaves and Fruit of Two Tomato Genotypes. Journal of American Society for Horticultural Science 139 (2014): 261-266.


 Impact Factor: * 4.2
Impact Factor: * 4.2 Acceptance Rate: 77.66%
Acceptance Rate: 77.66%  Time to first decision: 10.4 days
Time to first decision: 10.4 days  Time from article received to acceptance: 2-3 weeks
Time from article received to acceptance: 2-3 weeks 