Aerobic Bacterial Community of the Rice Striped Stem Borer: A Step towards Finding Bacterial Candidates for Paratransgenesis and RNAi Control
Abbas Heydari1, Mohammad Ali Oshaghi2*, Alireza Nazari1, Mansoureh Shayeghi2, Elham Sanatgar1, Nayyereh Choubdar2, Mona Koosha2, Fateh Karimian2
1Department of Entomology, Arak Branch, Islamic Azad University, Arak, Iran
2Department of Medical Entomology and Vector Control, School of Public Health, Tehran University of Medical Sciences, Tehran, Iran
*Corresponding Author: Mohammad Ali Oshaghi, Department of Medical Entomology and Vector Control, School of Public Health, Tehran University of Medical Sciences, Tehran, Iran
Received: 26 July 2021; Accepted: 03 August 2021; Published: 25 August 2021
Article Information
Citation: Abbas Heydari, Mohammad Ali Oshaghi, Alireza Nazari, Mansoureh Shayeghi, Elham Sanatgar, Nayyereh Choubdar, Mona Koosha, Fateh Karimian. Aerobic Bacterial Community of the Rice Striped Stem Borer: A Step towards Finding Bacterial Candidates for Paratransgenesis and RNAi Control. International Journal of Plant, Animal and Environmental Sciences 11 (2021): 485-502.
View / Download Pdf Share at FacebookAbstract
It is recognized that insects have close associations with a wide variety of microorganisms, which play a vital role in the insect's ecology and evolution. The rice striped stem borer, Chilo suppressalis, has economic importance at the global level. With the development of insecticide resistance, it is widely recognized that control of this pest is likely to need new tools that are not available today. Here, we focus on the aerobic bacterial community of the pest, to seek candidates for paratransgenesis or RNAi biocontrol of C. suppressalis. Culture-dependent PCR-direct sequencing was used to characterize the midgut bacterial communities of C.suppressalis at different life stages, collected in northern Iran, both from rice plants and from weeds on which the insect feeds. Our results show that the predominant genera of the Firmicutes and Proteobacteria phyla were Bacillus, Enterobacter and Klebsiella in all life stages, and they displayed differences in abundance. The high levels of B. subtilis and Enterobacter found in C. suppressalis suggests their potential utility for paratransgenesis and RNAi approaches to control this pest.
Keywords
Chilo suppressalis; Paratransgenesis; RNAi; Bacteria; 16SrRNA; Iran
Article Details
1. Introduction
Classified amongst various stem borers in the order Lepidoptera, the striped stem borer, Chilo suppressalis (Walker), in the family Crambidae, is an important insect pest of rice in southern Europe and Asia, including Iran [1-5]. It infests rice plants from the seedling stage to maturity and may cause dead hearts and whiteheads during the vegetative and reproductive stages, respectively. In addition to rice, this insect harms other crops and weeds, including water bamboo, water oat, corn, and sugarcane [6, 7]. Current efforts to control the rice striped stem borer rely heavily on the application of chemical insecticides; however, this pest still causes economically important damage to crops in many countries [8]. The emergence of pest strains resistant to insecticides, the impact of insecticides on the environment, the toxicity to animals and humans, and the costs to farmers have led to renewed calls for the search for, and development of, new sustainable and cost-effective pest control tools.
Paratransgenesis and RNA interference (RNAi) are among possible alternative control strategies, where commensal or symbiont bacteria found in the gut of the insect pest are engineered to reduce insect fitness and to suppress the pest population [9-15]. The principal and essential step in paratransgenesis is the identification of suitable bacteria in the insect. The characteristics required for a candidate include being non-pathogenic to humans, host plants, and non-target animals; being a dominant species within the insect microflora; being cultivable in cell-free media, being malleable to transformation with foreign DNA, and having a wide distribution [16, 17]. However, so far, most of the research on paratransgenesis or RNAi has been focused on vectors of human diseases, and only a few crop pests have been studied. Insect gut microbial communities, including bacterial species, have been investigated in various insects, including blood-sucking bugs [18], tsetse flies [19], mosquitoes [20-23], American cockroaches [24], sand flies [25-28], honeybees [29-31], and desert locusts [32]. The potential of the paratransgenesis approach against crop pests has been demonstrated in the Glassy-winged sharpshooter [33-34] and in honeybees [15].
Although C. suppressalis is the most important rice pest in the world, the microbial diversity associated with this insect pest has been little studied, with only two reports on the microbial diversity associated with the level of insecticide resistance among various populations of stem borer in China [6, 35]. In response to the lack of knowledge about these bacterial communities, and in the hope of finding bacterial candidates for paratransgenesis and RNAi in C. suppressalis, this study was conducted to assess the composition of aerobic gut bacterial communities of the insect at different life stages, from two different geographical regions in northern Iran. The gut bacteria were identified by culture-dependent isolation, followed by PCR-direct-sequencing of the 16S gene. The commensal bacterial communities of rice and sorghum in these geographic regions was also investigated, to examine any possible relationship between host plants and the microbiotal diversity of C. suppressalis. This information is important for the better understanding of symbiotic or commensal relationships between the bacteria and stem borer, the mechanisms that determine gut microbiota composition, and the potential for introduction of candidates for paratransgenesis or RNAi approaches against the pest in the study areas.
2. Materials and Methods
2.1 Sample collection
To cover the different life stages (larvae, pupae, and adult), the C. suppressalis populations were sampled in May-October 2018 (i.e. during the rice-growing season) from 18 rice fields in different regions of Mazandaran and Gillan provinces (Table 1). Sampling from rice and weeds was performed based on the distribution of rice fields in both provinces. Adults were captured using light traps, whereas the larvae and pupae were collected from the stems of the infested plants.
Each province was divided into three: eastern, central, and western parts, and in each part, where possible, three collection sites were selected based on their distances from the Caspian Sea. The specimens collected in the field were taken immediately to an experimental laboratory, where each sample was identified and used for isolation of bacteria from their alimentary canals.
2.2 Isolation of bacteria
The C. suppressalis specimens were immediately microdissected individually on sterile glass slides in a microbiological safety cabinet. First,the specimens were surface sterilized with bleach (10% sodium hypochlorite) for 5 min, and then rinsed with distilled water and air-dried. The gut of each specimen was then micro-dissected and homogenized in 1ml of sterile PBS by shaking with glass beads in a sterile tube.
To reduce the risk of laboratory-derived conta-mination, we used sterile workstations with sterile gloves, pipette tips with filters, and PCR grade RNAse-free water, and the experiments were performed under laminar flow hoods. The homogenized gut was transferred to screw-topped test tubes containing 5 ml of brain heart infusion (BHI) broth and incubated at 37°C for 24–48 h in aerobic conditions. Cloudy test tubes were considered as positive specimens. The grown bacteria were serially diluted or streaked on BHI agar plates and subcultured several times on the agar plates in the same conditions to achieve individual purified colonies. Test tubes containing BHI broth were opened near the dissection area under sterile conditions during the micro-dissection processes. Individual colonies were selected and used for further molecular identification.
The richness of each bacterial family was calculated, based on the total number of 16S sequences. The water used for the final rinsing of the cuticles was used as negative controls and plated in parallel. Gloves were changed frequently to avoid RNAse/DNAse contamination. Surface sterilization of the workstation was performed with bleach (10% sodium hypochlorite) followed by alcohol (70%) before and after each experiment. Also, instruments were autoclaved before and after handling each sample, and we avoided talking, sneezing, and coughing, and touching areas where DNA might exist.
2.3 16S rRNA gene amplification
DNA extraction from individual colonies was carried out using the phenol/chloroform DNA extraction method, as previously described [26]. Nearly 1500 bp of the bacterial 16S rRNA gene, including the less variable V1 and V2 regions and the highly variable V3-V5 regions were amplified using the universal primers 16suF (5'-GAG TTT GAT CCT GGC TCA G-3') and 16suR (5'-GTT ACC TTG TTA CGA CTT-3') [36]. Polymerase chain reaction (PCR) amplification was carried out using a Maxime PCR PreMix Kit (i-Taq) in 20μl reaction mixtures containing 1μl of each primer at 10μM concentration and 1–2μl (~0.1μg) of extracted genomic DNA.
BHI agar media and ddH2O were used as negative controls. The thermal cycler conditions were set as follows: an initial denaturation at 94°C for 10 min, followed by 35 cycles of denaturation at 95°C for 30 s, annealing at 57.5°C for 40 s and extension at 72°C for 30 s. Final extension was at 72°C for 8 min. The PCR products were separated on a 1% agarose gel containing ethidium bromide and were visualized using a UV transilluminator. The QIAquick PCR Purification Kit (Qiagen, Hilden, Germany) was used to purify the PCR products before sequencing.
To assess the environmental contamination, the cuticles of each specimen were used as an environmental control and were removed from the carcass and subjected to DNA extraction by phenol/chloroform and PCR amplification of 16S rRNA gene, as before. Where the negative control was positive, the specimen was eliminated from further analysis.
2.4 16S rRNA gene sequencing and analysis
First, 16S rRNA amplicons were amplified using each forward or reverse primer, and then sequenced using the Sanger method (Macrogen, Seoul, S. Korea). To compare these sequences with those available in ribosomal databases, eight databases of prokaryotic 16S rRNA gene were used: NCBI (nucleotide collection; http:// blast.ncbi.nlm.ni-h.gov/Blast.cgi), EMBL (http://www.ebi.ac.uk/ena), RDP (http://rdp.cme.msu.edu/seqmatch/ seqma-tch_intro.jsp), EzTaxon-e (http://eztaxon-e.ezbiocloud.net), Greengenes
(http://greengenes.lbl.gov/cgibin/nph-index.cgi), DDBJ (http://blast.ddbj.nig.ac.jp/ ?lang=en), leBIBI (http://umr5558-sud-str1.univ-lyon1.fr /lebibi/lebi-bi.cgi), and Blast2Tree (http://bioinfo.unice.fr/ blast) [37, 38]. Sequence homology with available data was assessed, based on the number and quality of nucleotides of the sequence reads using appropriate features of the data, such as cultivable and/or non-cultivable phenotype and type and/or non-type specimens.
The sequences were assigned at the species level, based on either the most common nomenclature within the results of the eight databases or the highest similarity rate.
Nucleotide homology >95% and >98% were considered as lower thresholds at genus and species levels, respectively (https://rdp.cme.msu.edu).
The partial 16S rDNA consensus sequences obtained in this study were annotated in the GenBank database using the 16S ribosomal RNA database (https://submit.ncbi.nlm. nih.gov/subs/genbank). Species assignation of the bacteria was verified by phylogenetic analysis using 16S rRNA gene sequences, based on the neighbor-joining algorithm of MEGA7 Software. Cytoscape Software (http://www.cytoscape.org) was used as a tool for visualizing complex networks between data, to visualize bacterial richness and shared bacteria in the three life stages through the network analysis. Data, as CYS files containing vertices or nodes (representing bacteria and life stages, or hosts, or location) and edges (representing links), were submitted to Cytoscape software v.3.5.1. GraphPad Prism software v.5.00 for Windows (GraphPad, San Diego, USA) and Student’s t-test embedded in the software was used for graphical representation and statistical analysis, respectively.
3. Results
In total, 1150 C. suppressalis specimens, comprising 671 (58.35%) larvae, 199 (17.3%) pupae, and 207 (24.35%) adults were collected from the study areas. Almost identical numbers of specimens were collected from each of the provinces of Mazandaran and Gillan (573 versus 577) (Table 1). All the specimens were processed individually for their midgut bacterial composition.
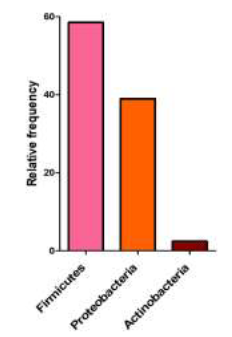
In total, 250 bacterial colonies, comprising 40 species, were isolated and identified from the midgut of the infected C. suppressalis specimens (Tables 2-3). The bacteria in this study were assigned to three phyla, nine families, and thirteen genera. Amongst the detected phyla, the Firmicutes were predominant, with a mean relative frequency of 58.5%, followed by Proteobacteria with a mean relative frequency of 39.0%, and Actinobacteria with a mean relative frequency of 2.5% (Figure 1). The family Bacillaceae was the most predominant group, with mean relative frequencies of 45 % (Tables 2-3).
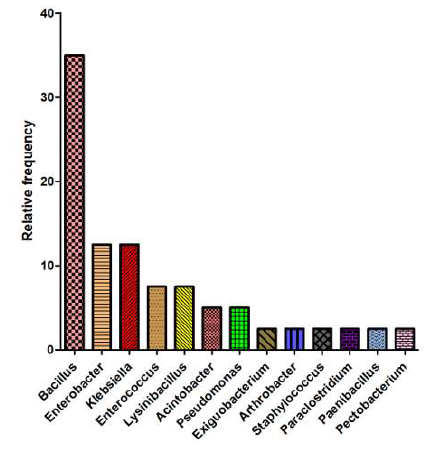
The most abundant of the genera in the C. suppressalis specimens was Bacillus, with a mean relative frequency of 35% (Figure 2). The second most abundant genera were Enterobacter and Klebsiella, each with a mean relative frequency of 12.5% (Figure 2). Among the gut bacteria, B. subtilis and Enterobacter spp. were found often in the C. suppressalis samples. These bacteria are potential candidates for paratransgenesis or RNAi approaches against this important pest.
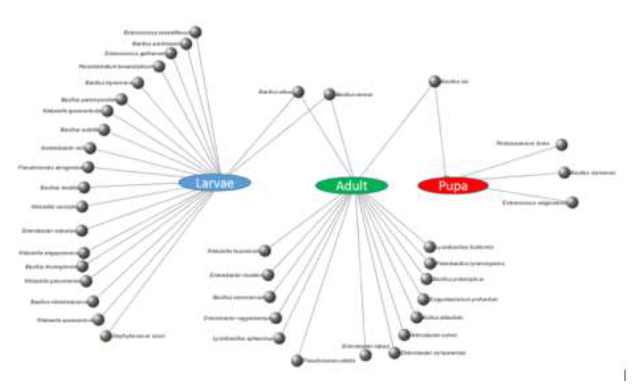
The frequency and diversity of bacteria in adults and larvae of C. suppressalis was very similar (n=17 versus n=21); however, there was a huge reduction in the diversity and abundance of bacteria in the pupal stage (n=4) (Figure 3). The most abundant bacteria in the larvae were of the genus Bacillus (42.9%), followed by Klebsiella (19%), whereas the most abundant bacteria in the adults were Bacillus (35.3%) and Enterobacter (23.5%).
Bacillaceae were present at high relative abundance in both C. suppressalis adults and larvae, with mean relative frequencies of 64.7% and 47.6%, respectively. However, there were significant differences between the composition of the bacterial communities in larvae and adults (F=4.35839, p=0.001; Tables 2-3), with only Bacillus cereus and B. albus being common to both (Figure 3). Although there was a slight increase in the abundance of Enterobacteriaceae in adults, the Enterobacteriaceae bacteria showed similar relative abundances in adults and larvae. Of the four bacteria species observed in pupae, only B. luti was common to pupae and adults (Figure 3).
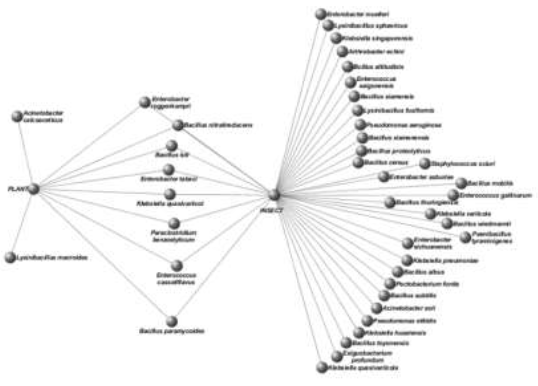
The host of the C. suppressalis specimens was, almost exclusively, the Rice plant. Although six host weeds (Johnson/Aleppo millet grass, cockspur grass, Bidens, Willow weed/Curlytop knotweed/Ladys thumb, sorghum, and one unknown species) in the study area were investigated, only a few weed plants were infested with C. suppressalis. Moreover, the majority of the C. suppressalis specimens of these hosts did not harbor bacteria in their midgut. To find the possible origins of the gut microbiome of C. suppressalis, we tested the bacteria on the surface of leaves and stems of the plant hosts (rice and weeds) and matched them with those isolated from the C. suppressalis guts. Rice plants were found to harbor more bacteria species than the weed plants (8 versus 2). Out of the ten bacteria species observed on the plants, eight were common to both the guts and the host plants (Figure 4). Of these eight shared bacteria, five were observed in larvae, two in adults, and one in a pupa (Tables 2,3). The other two bacteria were not observed in the insect guts.
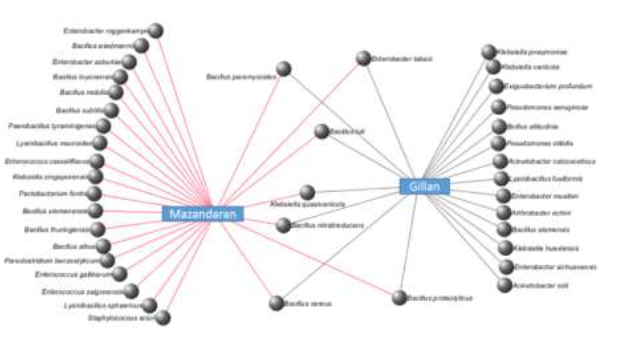
Although Mazandaran province showed more abundant bacterial communities than Gillan, statistical analysis confirmed that there were no significant differences between the microbial abundance of C. suppressalis guts in the two provinces (26 versus 21) (Figure 5). However, there were significant differences in the microbial composition of the samples from the two provinces (F=1.93071, p=0.041), where only 7 bacterial species were shared between the two locations (Figure 5).
|
Province |
Location |
Latitude & Longitude |
No of specimens on |
Total |
|||
|
Rice |
Weeds |
||||||
|
L |
P |
A |
L |
||||
|
Gillan |
Astara |
N:37°-16'-82'' |
11 |
10 |
24 |
21 |
66 |
|
E:49°-53'-40.9'' |
|||||||
|
Talesh |
N:37°-42'-59.8'' |
15 |
12 |
23 |
24 |
74 |
|
|
E:48°-57'-10.6'' |
|||||||
|
Asalem |
N:37°-33'-55.8'' |
11 |
10 |
19 |
12 |
52 |
|
|
E:49°-6'-55.3'' |
|||||||
|
Anzali |
N:37°-27'-21.5'' |
12 |
12 |
17 |
30 |
71 |
|
|
E:49°-36'-11.6'' |
|||||||
|
KHomam |
N:37°-22'-56.6'' |
12 |
10 |
21 |
21 |
64 |
|
|
HW |
E:49°-39'-33.5'' |
||||||
|
Rasht |
N:37°-12'-17.8'' |
11 |
10 |
11 |
21 |
53 |
|
|
Res Ins |
E:49°-38'-29.5'' |
||||||
|
Lahijan |
N:37°-13'-7'' |
13 |
10 |
22 |
25 |
70 |
|
|
E:49°-58'-26.5'' |
|||||||
|
Navideh |
N:37°- 16'- 35.3'' |
12 |
10 |
11 |
27 |
60 |
|
|
E:49°-44'-54.2'' |
|||||||
|
Klachai |
N:37°-40'-32.6'' |
13 |
19 |
11 |
24 |
67 |
|
|
E:50°-23'-13.3'' |
|||||||
|
Subtotal |
110 |
103 |
159 |
205 |
577 |
||
|
Mazandaran |
Shirood |
N:36°-51'-53.3'' |
7 |
11 |
10 |
6 |
34 |
|
E:50°-46-?54.7'' |
|||||||
|
Tonekabon |
N:36 °- 47'-43.7'' |
10 |
10 |
20 |
32 |
72 |
|
|
RW |
E:50°-54'-49.2'' |
||||||
|
Chalous |
N:36°-37' -55.615'' |
19 |
11 |
10 |
34 |
74 |
|
|
RW(Kosksara) |
E:51° -27'-24.777'' |
||||||
|
Amol, MohamadAbad |
N:36°-28' -30.685'' |
6 |
9 |
10 |
30 |
55 |
|
|
E:52°-27'-48.617'' |
|||||||
|
Amol Rice Sta. (Galekash) |
N: 36°-21'-48.911" |
10 |
10 |
10 |
32 |
62 |
|
|
E: 52°-21'-17.892" |
|||||||
|
Amol, |
N: 36° -35'30.615'' |
10 |
12 |
9 |
34 |
65 |
|
|
Marzango |
E: 52° 28'-38.671'' |
||||||
|
Ghaemshahr, CheftKola |
N: 36°- 25' 34.281'' |
10 |
10 |
11 |
30 |
61 |
|
|
E: 52°-48' -44.993'' |
|||||||
|
Ghaemshahr, Telar Hotel |
N: 36°-28'-56.524'' |
12 |
10 |
30 |
33 |
85 |
|
|
E: 52° -48'-54.242'' |
|||||||
|
Jouybar (Sarvkola) |
N: 36°-35'-26.104'' |
10 |
13 |
11 |
31 |
65 |
|
|
E: 52°-54'-22.013'' |
|||||||
|
Subtotal |
94 |
96 |
121 |
262 |
573 |
||
|
Total |
204 |
199 |
280 |
467 |
1150 |
||
L: larva; P: pupa; A: adult.
Table 1: Details of C. suppressalis specimens collected in rice fields during the growing season, northern Iran, 2018.
|
Assigned bacterial spp. |
Origin |
Plant host |
In Dev. Stage |
Family |
Phylum |
Genbank ID |
|
Bacillus xiamenensis |
In |
NA |
A |
Bacillacae |
Firmicutes |
MT176557 |
|
Bacillus wiedmannii |
In |
Rice |
L |
Bacillacae |
Firmicutes |
MT355806 |
|
Bacillus toyonensis |
In |
Rice |
L |
Bacillacae |
Firmicutes |
MT176558 MT176559 |
|
Bacillus thurengiensis |
In |
Rice |
L |
Bacillacae |
Firmicutes |
MT176560 |
|
Bacillus proteolyticus |
In |
NA |
A |
Bacillacae |
Firmicutes |
MT176561 |
|
Bacillus paramycoides |
Pl |
Rice |
NA |
Bacillacae |
Firmicutes |
MT176562 |
|
In |
Rice |
L |
MT176563 |
|||
|
Bacillus nitratireducens |
Pl |
Weed |
NA |
Bacillacae |
Firmicutes |
MT176564 MT176565 |
|
In |
L |
|||||
|
Bacillus subtilis |
In |
Rice |
L |
Bacillacae |
Firmicutes |
MT176566 |
|
Bacillus mobilis |
In |
Rice |
L |
Bacillacae |
Firmicutes |
MT176567 |
|
Bacillus luti
|
Pl |
Rice |
NA |
Bacillacae |
Firmicutes |
MT176568 |
|
In |
Rice |
P |
MT176569 |
|||
|
In |
Rice |
P |
MT176570 |
|||
|
Bacillus cereus
|
In |
Rice |
L |
Bacillacae |
Firmicutes |
MT176571 |
|
In |
NA |
A |
MT176574 |
|||
|
In |
NA |
A |
MT176572 |
|||
|
In |
NA |
A |
MT176573 |
|||
|
In |
NA |
A |
MT176575 |
|||
|
Bacillus albus |
In |
Rice |
L |
Bacillacae |
Firmicutes |
MT176576 MT176577 |
|
NA |
NA |
A |
||||
|
Lysinibacillus sphaericus |
In |
NA |
A |
Bacillacae |
Firmicutes |
MT176578 |
|
Lysinibacillus macroides |
Pl |
Rice |
NA |
Bacillacae |
Firmicutes |
MT176579 |
|
Paraclostridium benzoelyticum
|
Pl |
Rice |
NA |
Peptostreptococcaceae |
Firmicutes |
MT176580 MT176582 MT176581 |
|
In |
Rice |
L |
||||
|
In |
Rice |
L |
||||
|
Paenibacillus tyraminigenes |
In |
NA |
A |
Flexibacteraceae |
Firmicutes |
MT355807 |
|
Enterococcus saigonensis |
In |
Rice |
P |
Enterococcaceae |
Firmicutes |
MT176583 |
|
Enterococcus gallinarum |
In |
Rice |
L |
Enterococcaceae |
Firmicutes |
MT176584 |
|
Enterococcus casseliflavus |
Pl In |
Rice Weed |
NA L |
Enterococcaceae |
Firmicutes |
MT176585 MT176586 |
|
Staphylococcus sciuri |
In |
Rice |
L |
Staphylococcaceae |
Firmicutes |
MT176587 |
|
Enterobacter Tabaco |
Pl In |
Rice NA |
NA A |
Enterobacteriaceae |
Proteobacteria |
MT176588 MT366203 |
|
Enterobacter roggenkampii |
Pl In |
Rice NA |
NA A |
Enterobacteriaceae |
Proteobacteria |
MT176589 MT366202 |
|
Enterobacter asburiae |
In |
Rice |
L |
Enterococcaceae |
Proteobacteria |
MT176601 |
|
Pectobacterium fontis |
In |
Rice |
P |
Pectobacteriaceae |
Proteobacteria |
MT355808 |
|
Klebsiella singaporensis |
In |
Rice |
L |
Enterobacteriaceae |
Proteobacteria |
MT355805 |
|
Klebsiella quasivariicola |
Pl In |
Rice Rice |
NA L |
Enterobacteriaceae |
Proteobacteria |
MT176590 MT366204 |
In: insect; Pl: plant; A: adult; L: larva; P: pupa; NA: not applicable; Dev: developmental.
Table 2: Details of the 26 bacterial species (42 strains) isolated from the midgut of C. suppressalis or its host plants in Mazandaran province, east coast of Caspian Sea, north Iran.
|
Assigned bacterial spp. |
Origin |
Plant host |
In Dev. Stage |
Family |
Phylum |
Genbank ID |
|
Bcillus altitudinis |
In |
NA |
A |
Bacillacae |
Firmicutes |
MT176591 |
|
Bacillus cereus |
In |
Rice |
L |
Bacillacae |
Firmicutes |
MT176592 |
|
In |
NA |
A |
MT366205 |
|||
|
Bacillus luti |
In |
NA |
A |
Bacillacae |
Firmicutes |
MT355810 |
|
Bacillus nitratireducens |
In |
Rice |
L |
Bacillacae |
Firmicutes |
MT176593 |
|
Bacillus paramycoides |
Pl |
Rice |
NA |
Bacillacae |
Firmicutes |
MT176594 |
|
Bacillus proteolyticus |
In |
NA |
A |
Bacillacae |
Firmicutes |
MT176595 |
|
Bacillus siamensis |
In |
Rice |
P |
Bacillacae |
Firmicutes |
MT176596 |
|
Lysinibacillus fusiformis |
In |
NA |
A |
Bacillacae |
Firmicutes |
MT176597 |
|
Exiguobacterium profundum |
In |
NA |
A |
Bacillacae |
Firmicutes |
MT355804 |
|
Acinetobacter calcoaceticus |
Pl |
Rice |
NA |
Moraxellaceae |
Proteobacteria |
MT176598 |
|
Acinetobacter soli |
In |
Rice |
L |
Moraxellaceae |
Proteobacteria |
MT176599 |
|
Arthrobacter echini |
In |
NA |
A |
Micrococcaceae |
Actinobacteria |
MT176600 |
|
Enterobacter muelleri |
In |
NA |
A |
Enterobacteriaceae |
Proteobacteria |
MT176601 MT366201 |
|
Enterobacter sichuanensis |
In |
NA |
A |
Enterobacteriaceae |
Proteobacteria |
MT176602 |
|
Enterobacter tabaci/mori |
In |
NA |
A |
Enterobacteriaceae |
Proteobacteria |
MT176603 |
|
MT355803 |
||||||
|
Pseudomonas aeruginosa |
In |
Rice |
L |
Pseudomonadaceae |
Proteobacteria |
MT355802 |
|
Pseudomonas otitidis |
In |
NA |
A |
Pseudomonadaceae |
Proteobacteria |
MT176604 |
|
Klebsiella quasivariicola |
In |
Rice |
L |
Enterobacteriaceae |
Proteobacteria |
MT176605 MT176606 |
|
Klebsiella variicola |
In |
Rice |
L |
Enterobacteriaceae |
Proteobacteria |
MT355809 |
|
Klebsiella pneumoniae |
In |
Rice |
L |
Enterobacteriaceae |
Proteobacteria |
MT176607 |
|
Klebsiella huaxiensis |
In |
NA |
A |
Enterobacteriaceae |
Proteobacteria |
MT176608 |
In: Insect; Pl: Plant; A: Adult; L: Larva, P: Pupa; NA: Not applicable; Dev: Developmental
Table 3: Details of the 21 bacterial species (25 strains) isolated from the midgut of C. suppressalis and its host plants in Gillan province, west coast of Caspian Sea, north Iran.
4. Discussion
This study is the first significant survey of the bacterial communities in C. suppressalis in Iran. Understanding the microorganisms that accompany insects, especially those of international economic importance, is essential for developing microbial-based tools for insect pest management, such as paratransgenesis, gene silencing or RNA interference, as well as novel chemical and microbial biopesticides, sterile insect technique (SIT), and incompatible insect technique (IIT) [39]. Here, we report an association between the bacterial communities of C. suppressalis and the bacteria of the host plant on which the larvae or adults feed. These results suggest new opportunities to develop paratransgenesis approaches, or combinations of paratransgenesis and RNAi approaches, by manipulation of the C. suppressalis microbiome to produce effector molecules with high impacts on the insect fitness contributing to their pest status. The presence of B. subtilis and Enterobacter in the C. suppressalis samples suggests that these bacteria are potential candidates for a paratransgenesis plus RNAi approach against the pest. These bacteria can be cultured, transformed, and readily delivered by the host plant (rice) to the C. suppressalis gut. These bacteria have been already used for the production of paratransgenic sand flies [11] and mosquitoes [40].
During this study, we characterized the bacterial community of C. suppressalis larvae, pupae, and adults and their plant hosts using culture-dependent isolation, followed by sequencing of the 16S gene. For the first time, to our knowledge, the possible transstadial transmission of bacteria from immature stages to the adult was investigated, as well as the possible acquisition of bacteria from host plants in this insect pest. Delivery of the modified bacteria to the insect gut is one of the most challenging issues of paratransgenesis or RNAi approaches to insect control, and the results of this study show that C. suppressalis may acquire bacteria from the host plant on which they feed at the larval stage, and via polluted sugar meals from plant flowers at the adult stage. However, in order to use these bacteria for paratransgenic approaches, it will be important to examine their capacity to efficiently colonize the gut or reproductive organs of C. suppressalis, and to express enough effector molecules or dsRNA to inhibit the target gene.
In this study we used culture-dependent isolation, followed by PCR-direct sequencing of the 16S gene to detect bacterial species; this method might have some disadvantages in comparison with high-throughput sequencing techniques (e.g. next generation sequencing: NGS). However, the method we used can potentially exclude slower-growing species and allow the observation of diverse characteristics of the isolated organisms, including the physiological characteristics, such as antibiotic resistance. Moreover, this method facilitates bacterial genome sequencing, and eliminates those bacteria incapable of being propagated on the culture media. Thus, it offers the best way of assessing the validity of the candidate bacteria, as it allows testing of their capacity to accept foreign DNA (genes or plasmids) for paratransgenesis and RNAi. To date, two studies on microbial diversity have been conducted on the larvae of C. suppressalis in China, using degeneration gradient gel electrophoresis (DGGE) to clarify the changes of gut microbial diversity before and after treatment with Bacillus thuringiensis (Bt) insecticidal proteins [6, 35]. They suggested that the changes in the bacterial abundance in midguts of larval C. suppressalis are related to the difference in Bt insecticidal proteins, geographical sites and successive rearing times.
The bacterial communities of the C. suppressalis specimens analyzed in our study predominantly consisted of the phylum Firmicutes (58.5%), and, to a lesser extent, the Proteobacteria (39.9%), and Actinobacteria (2.5%). A study of the C. suppressalis gut microbiota by Zhang et al (2013) had also found that it was dominated by the Firmicutes, followed by Proteobacteria, Chloroflexi, and Bacteroidetes. However, several studies on the composition of gut microbial communities across the Lepidoptera [41] have shown that most of the gut bacteria detected in butterflies and moths belong to the Proteobacteria phylum (42%), which differs from the C. suppressalis gut microbiome. However, those studies also found that the gut microbiome of Lepidoptera is highly variable between and within species; it was suggested that differences in the insect habitat and in the experimental methodologies used in the studies, such as variations in insect diet, the life stage, and the screening procedures (culture-based, culture-independent) may explain this variability. The different bacterial communities of C. suppressalis found in Iran and China may reflect the different methodologies, environmental conditions, life stages, and host plants used in the two studies. Therefore, caution is needed when recommending a specific bacterium for paratransgenesis or other symbiont-based control methods, because some bacteria are indigenous only in certain geographic regions and cannot be established in other areas. For example, the Bacillacae and Enterobacteriaceae were dominant in the microbial community of the Iranian C. suppressalis samples, differing from the microbial community found in this pest in China, where Enterococcaceae were the most common [6]. However, studies of the microbiome of 30 lepidopteran species indicated that the Bacillaceae and Enterobacteriaceae are two of the most common groups found in moths and butterflies [41]. The Pseudomonadaceae are only the third most widespread group across lepidopteran species and are also found less frequently in C. suppressalis specimens. The effect of geographical location on the C. suppressalis gut bacterial communities was shown in this study when comparing the provinces of Mazandaran and Gillan. The communities were relatively distinct, even though the host plant (rice) was the same at both sites, probably due to differences in phyllosphere bacteria inhabiting the host plants at each site. This finding is consistent with previous studies indicating that habitat may significantly affect the bacteria associated with lepidopteran species [42, 43].
This study has also shown that the C. suppressalis larvae (caterpillars) harbor a lower gut bacterial community than the adults, which agrees with a previous study indicating that microbial symbionts are generally absent or present only in low numbers in guts of lepidopteran larvae [44]. Caterpillars have a simple alimentary canal, lacking the intricate pouch-like structures (diverticula or caeca) that are known to carry bacterial symbionts in other insect taxa [45, 46]. Guts with many pouches might favor the establishment of a strong bacterial community, as occurs in non-lepidopteran insects with extremely rich bacterial gut communities (24, 32, 47-49]. Insects that feed on wood, decomposing matter, or garbage, such as termites, cockroaches, crickets, and some beetles have the most diverse gut bacterial communities [32, 49]. Several other factors, including short gut, highly alkaline conditions (pH values >10-12), host antimicrobial peptides, rapid food passage, and lining of the midgut epithelium with peritrophic matrix may also hinder microbial colonization in the gut of caterpillars [50-56]. In contrast, some adult butterflies and moths may have high gut microbial loads [57], although many Lepidopteran adults do not feed [44]. It is known that adults of C. suppressalis visit rice and other host plants for nectar and pollen [58] that may provide a source of bacteria.
Our results showed that three species of the plant bacteria were present in the guts of C. suppressalis adults, suggesting plant hosts as a source of bacterial acquisition in adults of this insect pest. These observations suggest that paratransgenesis or RNAi approaches are more applicable to the adult than to the larval stage of moths, because the bacteria can colonize the adult alimentary canal.
Acknowledgments
We are grateful to Mahmoud Hasanzadeh assistance and contributions to the project.
Declaration of Authorships
Funding
Partial support was received from Tehran University of Medical Sciences, Iran.
Conflicts of Interest/Competing Interests
The authors declare that they have no conflict of interest.
Ethics Approval
Not applicable
Consent to Participate
Not applicable
Consent for Publication
Not applicable.
Availability of Data and Material
The sequence data generated for this study can be found in GenBank under accession numbers MT176557-MT176608, MT355802-MT355810, and MT366201-MT366204.
Code Availability
Not applicable
Authors' Contributions
AH performed field work and species identification. MAO contributed to conceptualization and management of the project and writing and revising the manuscript. NC and MK performed the experiments, generated sequencing data and performed molecular analyses. FK drafted the manuscript and collaborated in imaging analysis. All authors read and approved the final version of the manuscript.
References
- Khan ZR. World bibliography of rice stems borers: 1794-1990. Int Rice Res Inst, Manila Philippines (1991).
- Zahiri R, Sarafrazi A, Salehi L, et al. A geometric morphometric study on populations of the Rice Stem Borer, Chilo suppressalis Walker (Lepidoptera: Crambidae) in northern Iran. Middle East 38 (2006): 73-84.
- Zibaee A, Sendi JJ, Alinia F, et al. A study on biochemical differences among five different groups of rice striped stem borer Chilo suppressalis Walker (Lepidoptera: Pyralidae). Invertebr Surviv J 5 (2008): 20-29.
- Chai HN, Du YZ, Zhai BP. Characterization of the complete mitochondrial genomes of Cnaphalocrocis medinalis and Chilo suppressalis (Lepidoptera: Pyralidae). Int J Biol Sci 8 (2012): 561-579.
- Heydari A, Oshaghi MA, Nazari A, et al. Genetic structure of rice striped stem borer, Chilo suppressalis (Lepidoptera:Crambidae) in North of Iran, Mitochondrial DNA Part A 31 (2020): 327-334.
- Li Z, Han L, Liu Y, et al. Change in the diversity of bacterial community in larval midguts of the striped stem borer, Chilo suppressalis (Lepidoptera: Crambidae), after treatment with Bacillus thuringiensis insecticidal proteins. Acta Entomol Sin 59 (2016): 292-300.
- Wang Y, Huang C, Hu B, et al. Gene flow across host?associated populations of the rice stem borer Chilo suppressalis Walker (Lepidoptera: Crambidae): implications for Bt resistance management in rice. Pest Manag Sci 76 (2020): 695-703.
- Yao R, Zhao DD, Zhang S, et al. Monitoring and mechanisms of insecticide resistance in Chilo suppressalis (Lepidoptera: Crambidae), with special reference to diamides. Pest Manag Sci 73 (2017): 1169-1178.
- Dillon R, Charnley K. Mutualism between the desert locust Schistocerca gregaria and its gut microbiota. Res Microbiol 153 (2002): 503-509.
- Coutinho-Abreu IV, Zhu KY, Ramalho-Ortigao M. Transgenesis and paratransgenesis to control insect-borne diseases: current status and future challenges. Parasitol Int 59 (2010): 1-8.
- Hurwitz I, Hillesland H, Fieck A, et al. The paratransgenic sand fly: a platform for control of Leishmania Parasit Vectors 4 (2011): 82.
- Taracena ML, Oliveira PL, Almendares O, et al. Genetically modifying the insect gut microbiota to control Chagas disease vectors through systemic RNAi. PLoS Negl Trop Dis 9 (2015): e0003358.
- Wang S, Dos-Santos AL, Huang W, et al. Driving mosquito refractoriness to Plasmodium falciparum with engineered symbiotic bacteria. Science 357 (2017): 1399-1402.
- Koosha M, Vatandoost H, Karimian F, et al. Delivery of a genetically marked Serratia AS1 to medically important arthropods for use in RNAi and paratransgenic control strategies. Microb Ecol 78 (2019 a): 185-194.
- Leonard SP, Powell JE, Perutka J, et al. Engineered symbionts activate honeybee immunity and limit pathogens. Science 367 (2020): 573-576.
- Favia G, Ricci I, Marzorati M, et al. Bacteria of the genus Asaia: a potential paratransgenic weapon against malaria. In: Aksoy S (ed.). Transgenesis and the management of vector-borne disease. New York, Springer (2008): pp 49-59.
- Koosha M, Vatandoost H, Karimian F, et al. Effect of Serratia AS1 (Enterobacteriaceae: Enterobacteriales) on the Fitness of Culex pipiens (Diptera: Culicidae) for Paratransgenic and RNAi Approaches. J Med Entomol 56 (2019 b): 553-559.
- Durvasula RV, Gumbs A, Panackal A, et al. Prevention of insect-borne disease: an approach using transgenic symbiotic bacteria. Proc Natl Acad Sci 94 (1997): 3274-3278.
- Medlock J, Atkins KE, Thomas DN, et al. Evaluating paratransgenesis as a potential control strategy for African trypanosomiasis. PLoS Negl Trop Dis 7 (2013): e2374.
- Favia G, Ricci I, Damiani C, et al. Bacteria of the genus Asaia stably associate with Anopheles stephensi, an Asian malarial mosquito vector. Proc Natl Acad Sci 104 (2007): 9047-9051.
- Chavshin AR, Oshaghi MA, Vatandoost H, et al. Identification of bacterial microflora in the midgut of the larvae and adult of wild caught Anopheles stephensi: a step toward finding suitable paratransgenesis candidates. Acta Trop 121 (2012): 129-134.
- Chavshin AR, Oshaghi MA, Vatandoost H, et al. Escherichia coli expressing a green fluorescent protein (GFP) in Anopheles stephensi: a preliminary model for paratransgenesis. Symbiosis 60 (2013): 17-24.
- Chavshin AR, Oshaghi MA, Vatandoost H, et al. Isolation and identification of culturable bacteria from wild Anopheles culicifacies, a first step in a paratransgenesis approach. Parasit Vectors 7 (2014): 419.
- Akbari S, Oshaghi MA, Hashemi-Aghdam SS, et al. Aerobic bacterial community of American cockroach Periplaneta americana, a step toward finding suitable paratransgenesis candidates. J Arthropod Borne Dis 9 (2015): 35-48.
- Hillesland H, Read A, Subhadra B, et al. Identification of aerobic gut bacteria from the kala azar vector, Phlebotomus argentipes: a platform for potential paratransgenic manipulation of sand flies. Am J Trop Med Hyg 79 (2008): 881-886.
- Maleki-Ravasan N, Oshaghi MA, Hajikhani S, et al. Aerobic microbial community of insectary population of Phlebotomus papatasi. J Arthropod Borne Dis 8 (2014): 69-81.
- Maleki-Ravasan N, Oshaghi MA, Afshar D, et al. Aerobic bacterial flora of biotic and abiotic compartments of a hyperendemic Zoonotic Cutaneous Leishmaniasis (ZCL) focus. Parasit Vectors 8 (2015): 63.
- Karimian F, Vatandoost H, Rassi Y, et al. Aerobic midgut microbiota of sand fly vectors of zoonotic visceral leishmaniasis from northern Iran, a step toward finding potential paratransgenic candidates. Parasit Vectors 12 (2019): 1-12.
- Kwong WK, Moran NA. Gut microbial communities of social bees. Nat Rev Microbiol 14 (2016): 374-384.
- Zheng H, Steele MI, Leonard SP, et al. Honeybees as models for gut microbiota research. Lab anim 47 (2018): 317-325.
- Romero S, Nastasa A, Chapman A, et al. The honeybee gut microbiota: strategies for study and characterization. Insect Mol Biol 28 (2019): 455-472.
- Dillon RJ, Dillon VM. The gut bacteria of insects: nonpathogenic interactions. Annu Rev Entomol 49 (2004): 71-92.
- Bextine BR, Harshman D, Johnson MC, et al. Impact of pymetrozine on glassy-winged sharpshooter feeding behavior and rate of Xylella fastidiosa J Insect Sci 4 (2004): 34.
- Arora AK, Forshaw A, Miller TA, et al. A delivery system for field application of paratransgenic control. BMC Biotechnol 15 (2015): 59.
- Zhang J, He Y, Chen J. Diversity analysis of bacterial community in midguts of larvae of the striped stem borer, Chilo suppressalis (Lepidoptera: Crambidae), with different levels of resistance to insecticides. Acta Entomol Sin 56 (2013): 1075-1082.
- Weisburg WG, Barns SM, Pelletier DA, et al. 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173 (1991): 0697-703.
- Devulder G, Perriere G, Baty F, et al. BIBI, a bioinformatics bacterial identification tool. J Clin Microbiol 41 (2003): 1785-1787.
- Chun J, Lee JH, Jung Y, et al. EzTaxon: a web-based tool for the identification of prokaryotes based on 16S ribosomal RNA gene sequences. Int J Syst Evol Microbiol 57 (2007): 2259-2261.
- Qadri M, Short S, Gast K, et al. Microbiome Innovation in Agriculture: Development of Microbial Based Tools for Insect Pest Management. Front Sustain Food Syst 4 (2020): 547751.
- Dehghan H, Oshaghi MA, Moosa-Kazemi SH, et al. Dynamics of Transgenic Enterobacter cloacae Expressing Green Fluorescent Protein Defensin (GFP-D) in Anopheles stephensi Under Laboratory Condition. J Arthropod Borne Dis 11 (2017): 515-532.
- Paniagua Voirol LR, Frago E, Kaltenpoth M, et al. Bacterial symbionts in Lepidoptera: Their diversity, transmission, and impact on the host. Front Microbiol 9 (2018): 556.
- Staudacher H, Kaltenpoth M, Breeuwer JA, et al. Variability of bacterial communities in the moth Heliothis virescens indicates transient association with the host. PLoS One 11 (2016): e0154514.
- Jones AG, Mason CJ, Felton GW, et al. Host plant and population source drive diversity of microbial gut communities in two polyphagous insects. Sci Rep 9 (2019): 1-11.
- Hammer TJ, Janzen DH, Hallwachs W, et al. Caterpillars lack a resident gut microbiome. Proc Natl Acad Sci. 114 (2017): 9641-9646.
- Alonso-Pernas P, Arias-Cordero E, Novoselov A, et al. Bacterial community, and PHB-accumulating bacteria associated with the wall and specialized niches of the hindgut of the forest cockchafer (Melolontha hippocastani). Front Microbiol 8 (2017): 291.
- Sudakaran S, Kost C, Kaltenpoth M. Symbiont acquisition and replacement as a source of ecological innovation. Trends Microbiol 25 (2017): 375-390.
- Prem Anand AA, Vennison SJ, Sankar SG, et al. 2010. Isolation and characterization of bacteria from the gut of Bombyx mori that degrade cellulose, xylan, pectin and starch and their impact on digestion. J Insect Sci 10 (2010): 107.
- Barbehenn RV, Constabel CP. Tannins in plant–herbivore interactions. Phytochemistry 72 (2011): 1551-1565.
- Engel P, Moran NA. The gut microbiota of insects–diversity in structure and function. FEMS Microbiol Rev 37 (2013): 699-735.
- Buchler ER. 1975. Food transit time in Myotis lucifugus Chiroptera: Vespertilionidae. J Mammal 56 (1975): 252-255.
- Dow JA. Extremely high pH in biological systems: a model for carbonate transport. Am J Physiol Regul Integr Comp Physiol 246 (1984): R633-R636.
- Buchsbaum R, Wilson J, Valiela I. Digestibility of plant constitutents by Canada Geese and Atlantic Brant. Ecology 67 (1986): 386-393.
- Johnson KS, Felton GW. Potential influence of midgut pH and redox potential on protein utilization in insect herbivores. Arch Insect Biochem Physiol 32 (1996): 85-105.
- Brinkmann N, Tebbe CC. Leaf-feeding larvae of Manduca sexta (Insecta, Lepidoptera) drastically reduce copy numbers of aadA antibiotic resistance genes from transplastomic tobacco but maintain intact aadA genes in their feces. Environ Biosaf Res 6 (2017): 121-133.
- Hegedus D, Erlandson M, Gillott C, et al. New insights into peritrophic matrix synthesis, architecture, and function. Annu Rev Entomol 54 (2009): 285-302.
- Jiang H, Vilcinskas A, Kanost MR. Immunity in lepidopteran insects. In Söderhäll K (ed.) Invertebrate immunity. Springer, US (2010): pp 181-204.
- Ravenscraft A, Berry M, Hammer T, et al. Structure and function of the bacterial and fungal gut microbiota of Neotropical butterflies. Ecol Monogr 89 (2019): e01346.
- Lu YH, Zheng XS, Lu ZX. Application of vetiver grass Vetiveria zizanioides: Poaceae (L.) as a trap plant for rice stem borer Chilo suppressalis: Crambidae (Walker) in the paddy fields. J Integr Agric 18 (2019): 797-804.


 Impact Factor: * 4.1
Impact Factor: * 4.1 Acceptance Rate: 75.32%
Acceptance Rate: 75.32%  Time to first decision: 10.4 days
Time to first decision: 10.4 days  Time from article received to acceptance: 2-3 weeks
Time from article received to acceptance: 2-3 weeks 