Huaier Induces Cancer Recovery by Rescuing Impaired Function of Transcription Control Based On the Individual Genomic Potential
Manami Tanaka1*, Tomoo Tanaka1, Fei Teng2, Hong Lin2, Ning Li3, Zhu Luo3, Ding Wei4, and Zhengxin Lu5
1Bradeion Institute of Medical Sciences, Co., Ltd., Itado 433-1, Isehara, Kanagawa 259-1145, Japan.
2BGI-Shenzhen, Building NO.7, BGI Park, No.21 Hongan 3rd Street, Yantian District, Shenzhen 518083, China 3BGI-Japan, Kobe KIMEC Center BLDG. 8F 1-5-2 Minatojima-minamimachi, Chuo-ku, Kobe 650-0047 Japan. 4Japan Kampo New Medicine, Co., Ltd., 2-8-10 Kayaba-Cho, Chuo-Ku, Tokyo 103−0025, Japan.
5QiDong Gaitianli Medicines Co., Ltd., No.88, Heping South Road, Qidong, Jiangsu Province, China.
*Corresponding Author: Manami Tanaka, Bradeion Institute of Medical Sciences, Co. Ltd., Itado 433-1, Isehara, Kanagawa 259-1145, Japan
Received: 18 November 2020; Accepted: 09 December 2020; Published: 28 December 2020
Article Information
Citation: Manami Tanaka, Tomoo Tanaka, Fei Teng, Hong Lin, Ning Li, Zhu Luo, Ding Wei, and Zhengxin Lu. Huaier Induces Cancer Recovery by Rescuing Impaired Function of Transcription Control Based On the Individual Genomic Potential. Archives of Clinical and Biomedical Research 4 (2020): 817-855.
View / Download Pdf Share at FacebookAbstract
Background: Clinical significance of anti-cancer effects of Huaier has been emphasized recently. We have proved that a broad spectrum of Huaier effects was based on the rescue of the disrupted Hippo signalling pathway in Drosophila mutants, especially through the rescue of transcriptional dysregulation.
Objective: We initiated clinical research for thorough understanding of molecular basis of Huaier effects.
Methods: The obtained peripheral blood samples were analyzed by total RNA and non-coding small RNA sequencing on BGISEQ-500 Platform.
Results: In the present study, we sequenced 92 samples sequencing from 31 patients, average generating over 7.4 Giga bases per sample. The results showed significant changes in transcriptional factors and corresponding genes, which resulted in quantitative and qualitative alterations of signaling networks responsible for rescuing all the molecular biological function, cellular component and biological processes. Those changes were identified through genome wide range, and that from RNA editing events, to the resulting transcriptomes with discovery of 24,344 novel transcripts. Drastic variance of up/down regulated genes were associated with siRNA- and miRNA-mediated post-transcription control. Transcriptional factor changes, mean 1,115 per person, integrated those altered transcripts and their functional networks for the rescue of damaged or disrupted control. We provided functional lineage map of these results to show the process of cancer recovery by the time course of Huaier administration.
Conclusion: The present study identified that the cancer recovery by Huaier therapy was based on the rescue of transcription control not only in cancer lesion, but also in almost all biological systems. These changes depended on individual capability and flexibility of g
Keywords
<p>Huaier (Trametes robiniophila murr); Cancer therapy; Total RNA- and small RNA-sequencing; Transcription Factor (TF); Differentially Expressed Genes (DEG); lineage map; Transcription control rescue</p>
Article Details
1. Introduction
Huaier has been introduced its significant anti-cancer efficacy, and more importantly, without any side effects and/or toxicity. Among many materials used in traditional medicine, Huaier demonstrated prominent potential especially on the recovery from hepatocellular carcinoma and breast cancer [1, 2]. We have reported with a simple and successful evidence that the rescue of Hippo signaling pathway was a molecular basis of anti-cancer effects of Huaier [3]. Thereafter many researches followed for the confirmation, to provide the evidence in vitro that dysregulation in Hippo signaling pathway, disruption of transcription control is the major course of carcinogenesis [4-11]. However, there have not been appeared the additional successful controlling materials which contributes for cancer recovery so far.
It is not enough, either, to explain all functions associated to anti- cancer effects of Huaier. The questions remains to be solved such as 1) how Huaier operates immunological influences both to enhance and also inhibit the excessive reaction, 2) how Huaier causes cancer specific cell death and simultaneous normal tissue reproduction, and 3) how Huaier improves immunodeficiency in cancer patients, 4) how Huaier effectively reduces (oxidative) stress materials in liver and central nervous system, and 5) how Huaier makes swift recovery from the opportunistic infections during cancer therapy, and etc. More importantly, the reason why Huaier has no toxicity or side effects is still unclear. Therefore, we initiated clinical research to answer these questions by identifying Huaier effects on all the biophysiological system. We hired the next generation sequencing methods by BGI for this purpose, by total RNA sequencing and also small non-coding RNA sequencing. At the same time, the samples used in this approach should be free from the effects of chemotherapeutic agents, especially molecular-targeting anti-cancer drugs, which disrupted largely molecular mechanisms.
Beyond our expectations, the results obtained in the present study brought us to overlook the whole process of genome dynamics in each individual just as tracing the evolutional process of human nature. Consequently, we identified striking genomic plasticity and flexibility involving every element required to maintain homeostasis, and in the process of transcription and translation. The patients who had enough genomic potential to tolerate total alterations to restore normal homeostasis resulted in successful cancer recovery, and vice versa. As predicted before, we confirmed the rescue of transcription control in Hippo signalling pathway in the present study, but the similar rescue was observed among almost all the transcription control systems with the massive alterations in transcriptional factors (mean 1,115/person). This observation contributed to rescue dysregulated functions not only in current cancer, but also in every physiological function damaged by ageing, stress accumulations, and any possible disorders inside. Thus, the data obtained here could provide the answers not only on anti-cancer effects of Huaier, but also corresponding questions about the broad efficacy on all the biophysiological system disorders caused mainly by ageing and environmental stresses. These effects are observed in a dose and duration dependent manner, and that without any toxicity, side effects, and resistance in a long period of use, since all the effects are originated as spontaneous reaction based on the individual genomic potential.
2. Materials and Methods
2.1 Project Design
The present study initiated systematic understanding of the genomic and genetic requirement for the cancer recovery. Huaier compounds were provided by the manufacturer for this purpose with a strict control on transfer to Japan, good condition for maintenance, and provision to the patient volunteers. The present study was strictly conducted according to the guidelines of the Declaration of Helsinki and the principles of good clinical practice. Written informed consent was obtained from the patients. This clinical research was applied according to the Consolidated Standards of Clinical Research Trials guidelines and was applied to the Japanese Medical Association on 9th February 2018, and approved on 5th March, 2018 (ID: JMA-IIA00335). The project has been strictly conducted with a monthly review by the ethics committee consisted by the experts on Medicine, Nursing, Laws,
Pharmaceutics and Business Community (first committee held on 9th February, 2018).
We used Huaier compounds as complementary therapy, without any chemotherapy and radiotherapy which disrupt the molecular systems. Only surgical operation was performed when applicable. We thus planned and initiated an open-style, before-after controlled study, using peripheral blood as sampling materials to understand the almost all molecular events in each Huaier taking patient. The sampling materials were total blood, the same as reported previously [12, 13], by the consideration of as follows; 1) easy to obtain from any candidate patients, 2) non- or less-hazardous procedure, and 3) applicable to any patients of any stages, even after the surgical operation. To compare with the other sampling, RNA extraction using nuclear cell components in peripheral blood rapidly reflects the biophysiological changes, and that more sensitive to monitor the course of any treatment than any other samples such as dissected organs. To avoid the other molecular influence than Huaier, conventional chemotherapy was not applied to the volunteer patients during a research period. Age-matched healthy controls with or without Huaier administration were used for comparison.
2.2 Patient characteristics and sample collection
Basically, the volunteers were chosen on the demand of each individual, with a condition as written below;
- diagnosed as cancer in the clinics,
- exclusive for anti-cancer chemotherapy, radiotherapy, or any measures to influence genomic or genetic events in physiological condition for 90 days research period,
- between ages of 35 to 85
The normal healthy volunteers were chosen from the group of 1) without any diagnosed diseases, symptoms, and disorders, and 2) no drug administration, with an age distribution from ca. 35 to 85 years old. The blood samples were obtained from the volunteer three times; 1: before, 2: 30 days after, and 3: 90 days after 20g per day Huaier administration [14, 15]. In case, more sampling was applied to some patients for follow- up study in continuously Huaier taking patients. The samples (4ml blood) in PAXgene Blood RNA Tube (QIAGEN, Valencia, CA) was handled and kept at –80°C until shipping to BGI Shenzhen by dry-ice cargo. The RNA extraction, quality check for sequencing were performed in BGI Shenzhen according to the BGI protocols.
2.3 RNA extraction & miRNA library construction
The obtained peripheral blood samples (92 samples from 31 cancer patients) were designated to be analyzed by RNA extraction in BGI Shenzhen, construction of whole transcriptome library for sequencing on BGISEQ-500 Platform, followed by whole transcriptome mapping [16-18]. Total RNA was extracted using Trizol (Invitrogen, Carlsbad, CA, USA), according to the manual instructions. The mix was centrifuge at 12,000×g for 5min at 4°C. The supernatant was transferred to a new 2.0 ml tube which was added 0.3ml of Chloroform/isoamyl alcohol (24:1) per 1.5ml of Trizol reagent. After the mix was centrifuged at 12,000×g for 10min at 4°C, the aqueous phase was transferred to a new 1.5ml tube which was add equal volume of supernatant of isopropyl alcohol. The mix was centrifuged at 12,000×g for 20min at 4°C and then removed the supernatant. After washed with 1ml 75% ethanol, the RNA pellet was air-dried in the biosafety cabinet and then dissolved by add 25μl~100μl of DEPC-treated water. Subsequently, total RNA was qualified and quantified using a Nano Drop and Agilent 2100 bioanalyzer (Thermo Fisher Scientific, MA, USA).
2.4 mRNA Library Construction
Oligo(dT)-attached magnetic beads were used to purified mRNA. Purified mRNA was fragmented into small pieces with fragment buffer at appropriate temperature. Then First-strand cDNA was generated using random hexamer-primed reverse transcription, followed by a second-strand cDNA synthesis. afterwards, A- Tailing Mix and RNA Index Adapters were added by incubating to end repair. The cDNA fragments obtained from previous step were amplified by PCR, and products were purified by Ampure XP Beads, then dissolved in EB solution. The product was validated on the Agilent Technologies 2100 bioanalyzer for quality control. The double stranded PCR products from previous step were heated denatured and circularized by the splint oligo sequence to get the final library. The single strand circle DNA (ssCir DNA) was formatted as the final library. The final library was amplified with phi29 to make DNA nanoball (DNB) which had more than 300 copies of one molecular, DNBs were loaded into the patterned nanoarray and single end 50 bases reads were generated on BGIseq500 platform (BGI-Shenzhen, China).
Library was prepared with 1 μg total RNA for each sample. Total RNA was purified by electrophoretic separation on a 15% urea denaturing polyacrylamide gel electrophoresis (PAGE) gel and small RNA regions corresponding to the 18–30 nt bands in the marker lane (14-30 ssRNA Ladder Marker, TAKARA, Japan) were excised and recovered. Then the 18–30 nt small RNAs were ligated to a 5′-adaptor and a 3′-adaptor. The adapter-ligated small RNAs were subsequently transcribed into cDNA by SuperScript II Reverse Transcriptase (Invitrogen, USA) and then several rounds of PCR amplification with PCR Primer Cocktail and PCR Mix were performed to enrich the cDNA fragments. The PCR products were selected by agarose gel electrophoresis with target fragments 100~120 bp, and then purified by QIAquick Gel Extraction Kit (QIAGEN, Valencia, CA). The library was quality and quantitated in two methods: check the distribution of the fragments size using the Agilent 2100 bioanalyzer, and quantify the library using real- time quantitative PCR (QPCR) (TaqMan Probe). The final ligation PCR products were sequenced using the BGISEQ-500 platform (BGI-Shenzhen, China).
2.5 Bioinformatics Workflow
Bioinformatics Work was processed in BGI, Shenzhen. The detailed protocols were provided and demonstrated at BGI website: http://www.bgitechsolutions.com/). We first removed the reads mapped to rRNAs and get raw data, then we filtered the low quality reads (More than 20% of the bases qualities are lower than 10), reads with adaptors and reads with unknown bases (N bases more than 5%) to get the clean reads. After reads filtering, we mapped those clean reads only to reference genome using HISAT1 [19], and confirmed the sample quality. the bam files for genome mapping results were prepared using Integrative Genome Viewer (IGV) [20] provided by BGI. After genome mapping, STRINGTIE
[21] was used to reconstruct transcripts, and with genome annotation information, novel transcripts were identified by using Cuffcompare (a tool of Cufflinks) [22], and the coding ability of those new transcripts were predicted by CPC (Coding Potential Calculator) [23]. Then we map those clean reads onto reference genome(hg38_UCSC_20180118), followed with novel gene prediction, SNP & INDEL calling and gene splicing detection. We used GATK (Genome Analysis Toolkit) [24] to identify single nucleotide polymorphisms (SNP) and insertions and deletions (INDL) variants for each sample. This process supported importing multiple samples for comparison and show the distribution of reads in the exon, intron, UTR, intergenic areas based on the annotation results.
Finally, we identify DEGs (differentially expressed genes) between samples and do clustering analysis and functional annotations. In
the transcriptome analysis, all samples had library construction and sequenced on the MGISEQ-2000 and BGISEQ-500 platform. Paired-end reads were aligned to UCSC hg38 using Hisat2 and Bowtie2. We calculated the expression of genes by RSEM, call DEGs by PossionDis method, predicted novel transcripts by StringTie and call SNP by GATK. The obtained novel genes have been deposited to The NCBI GEO (GSE157086). With quantitative analysis of DEGs, we performed Gene Ontology (GO) classification and functional enrichment. Basically, GO has three ontologies: molecular biological function, cellular component and biological process, and the results were compared by each patient with a consideration of time course of Huaier administration. We also performed KEGG pathway classification [25] according to each cancer analyzed in the present study (https://www.genome.jp/kegg/) and functional enrichment. With the KEGG annotation result, we classify DEGs according to official classification, and were also perform pathway functional enrichment (only significant one). Furthermore, we applied the enrichment analysis of DEG in KEGG database.
2.6 Small RNA analysis
We have also performed total non-coding small RNA sequencing on the same BGISEQ-500 Platform, and the length of the small RNAs were found to be between 18 to 30 nucleotides. After filtering, clean tags were mapped to sRNA database such as miRBase [26], Rfam [27]. To make unique small RNA mapped to only one annotation, we follow priority rule: miRNA>piRNA>Rfam>other sRNA. After sRNA annotation, those unknown tags will be used to predict novel sRNA based on their architectural features. Thus, novel miRNA, piRNA, and siRNA were predicted.
Singled-end reads were aligned to UCSC hg38 MirBase22 and Rfam by Bowtie2. We calculated the expression of miRNAs by in- house program, predicted miRNA by mirdeep2, and predicted target genes by miRanda and TargetScan. The following processes were applied to the subsequent data analysis.
- Eliminate the low-quality reads, small tags, adaptors and other contaminants to get clean
- Summarize the length distribution of the clean tags, common and specific sequences between
- Annotate the clean tags into different
- Predict the novel miRNA by exploring the characteristic hairpin structure of miRNA
- Functionannotation of known miRNAs, including
Gene Ontology and Pathway.
The obtained novel small nuclear non-coding RNA have been deposited to The NCBI GEO (GSE157086).
3. Results
- Characterization of volunteer patients and normal controls The medical characteristics of the 31 volunteer patients were summarized in Table 1. The mean age is 67 years old, since this research excluded the congenital cancer or The present study summarizes the results from 92 sample sequencing from 31 patients, including 2 normal controls (patient No. 21 and 26), and 4 with benign tumors (No. 13, 14, 18, and 20). Normal control No. 21 was the same as No. 25, the opportunistic infection case after 8 months of the end of first research period as No. 21. In order to compare the results with the other volunteer patients with opportunistic virus infections, the sampling was continued over one year from the patients No.5, 8, and 10. The t volunteer No. 20 was designated to be normal healthy control without Huaier administration, was later identified adenoma progression in colon, and successfully removed by colon endoscopic dissection one month later.
The cancer origin and the stages (benign tumor to cancer TNM stages 0 to IV) varies among patients, and their symptom changes and anti-cancer effects after Huaier administration were confirmed by many ordinary clinical tests in clinics, and by the evaluation of CT, MRI, and X-ray images. The volunteers summarized in Table
1 were free from any conventional anti-cancer chemotherapy during research period (3 months). TNM Stage IV patients were only treated by Huaier. Interestingly, there were 5 volunteers who had virus infection during the observation period (No. 5, 8, 10, 12, 21/25). The methodology hired in the present study was also valid to identify virus infection, and that could analyse in quantity (Figure 4b). These results confirmed that Huaier could not prevent virus infection, however, the infection remained silent and totally cured within reasonable period of time (one month to half a year).
3.2 Summary of sequence events
The results were compared between the samples obtained before and after 1month, and 3 months of Huaier administration. The quantitative analysis of > 7.0 GB RNA sequencing of each sample provided estimated transcribed genes of >27,000; mean number of 27, 447 per sample (Table 2). The average mapping ratio with reference genome is 89.64 % (varies from 71.49% to 94.51%); 43,296 genes were identified per sample (in which 18,952 of them are already known genes). and 24,344 of them are novel genes. Total number of the novel transcripts identified was 172,487, composed of 1) 108,991 for previously unknown splicing event for known genes, 2) 24,344 novel coding transcript without any known features, and 3) 39,152 long non- coding RNA. The obtained novel genes and small nuclear non- coding RNA obtained in the present study have been deposited to the NCBI GEO (GSE157086).
3.3 Summary of RNA editing events
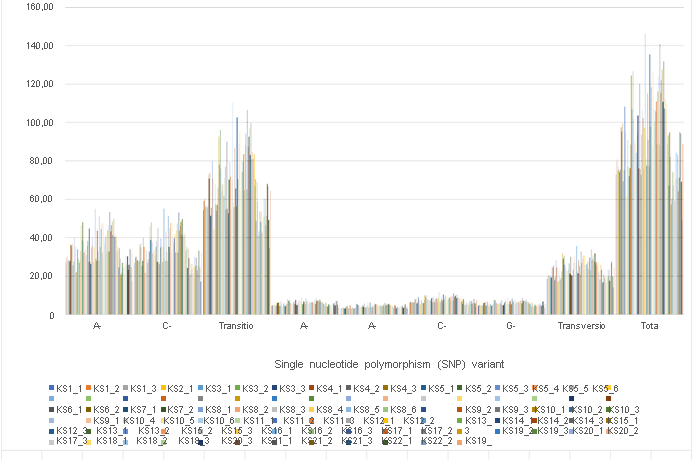
Final results are shown in Figure 1a. We identified total 8,503,297 SNP variant types in 92 sample analysis, and 92,427 SNP per sample in mean number (ranging from 48,994 to 146,102), whereas 22,688 in total among normal healthy individuals [16]. As for the SNP variations, A-G > C-T transitions were the most common mutations (3,152,487 and 3,142,470 respectively, 74.0%), followed by A-C > A-T (552,611 and 406,169, respectively, 11.3%), C-G > G-T (700,403 and 549,157 respectively, 14.7%) transversions, which is consistent with the previous reports on oesophageal squamous carcinoma cells [13]. Mean number of transitions per sample is 68,423 (ranging from 34,672 to 110,293), whereas
24,004 transversions (ranging from 14,322 to 35,809) (Table 2). These ratios found no significant differences among samples, but such a gross increase of SNP variants has not been reported before using the samples from the same person. The incidence ratio was three times higher in transition, which is much higher than that identified in oesophageal squamous carcinoma cells [13], and any other cancer cells analyzed before [28]. There were no significant differences identified by cancer origin, gender, and the stages of cancer. In contrast, the significant increases were observed at the time of opportunistic virus infections in patient No. 8, 10, 12, and 21.
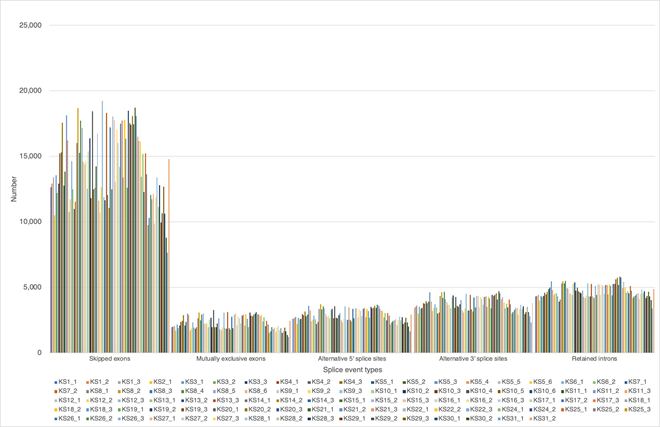
Distribution of SNP location is about 40% each in both exons and introns, and the rest located in intergenic region (12%) and within 2,000 bp up- and down-stream region (2% each) of the expressed genes. The distribution of numerous INDL (>2,000,000 per sample) location showed the similar results. Both SNP and INDL location was identified 40 to 45 % in exons, and up to 60% including the adjacent area. Thus, These RNA editing events did not randomly scatter among the whole genome, nor any correlation to cancer origin or stages. We also detect the level of differentially splicing genes (DSG) compared before and after Huaier administration, using rMATS [29]. The summary of splicing variation comparison is shown in Figure 1b. DSGs are regulated by alternative splicing (AS), which allows the production of a variety of different isoforms from one gene only. Changes in relative abundance of isoforms, regardless to the expression change, indicates a splicing-related mechanism. We detected five types of AS events, including skipped exon (SE), alternative 5’ splicing site (A5SS), alternative 3’ splicing site (A3SS), mutually exclusive exons (MXE), and retained intron (RI). Approximately total 2,557,753 splicing events in 92 samples were observed, in which were 1,303,783 SE (51%); 211,336 MXE (8.2%); 264,368 A5SS (10.3%); 343,124 A3SS (13.4%); and 435,142 RI (17.1%), respectively. Mean numbers of each event per sample (= per person) were 14,172 (ranging from 7,567 to19,240) SE; 2,297 (ranging from 1,229 to 3,258) MXE; 2,874 (ranging from 1,655 to 3,704) A5SS; 3,730 (ranging from 2,341 to 4,699) A3SS; and 27,802 (ranging from 16,294 to 36,079) RI, respectively. Just like SNP variants, the ratio of splicing events showed no significant differences among patients, and irrespectively to cancer origin, cancer stage, age, or gender (Table 2), only exception at the time of virus infections.
3.4 Comparison of up- and down-regulated transcriptomes before and after Huaier administration
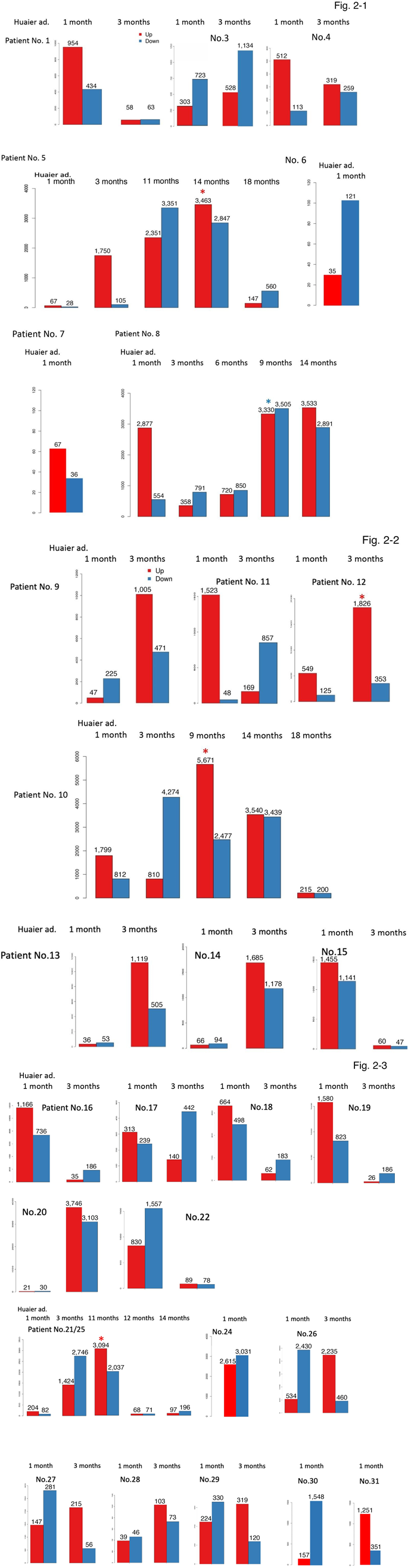
After the genome mapping and Gene Ontology classification, we defined 24,344 novel transcripts. Then we merge novel coding transcripts with reference ones to get complete reference, and map clean reads using Bowtie [30], followed by calculate gene expression level for each sample with RSEM [31]. The results provided the total numbers of predicted transcripts were 27,447, and the gene mean mapping ratio was 89.64 % (ranging from 71.49% to 94.51%). Based on the expression information, box plot analysis was completed o show the distribution of the gene expression level of each sample, followed by construction of the density map which revealed the change of gene abundance by the time course of Huaier administration. As a result of those analysis, gene expression level analysis enabled us to compare the levels of Differentially Expression Genes (DEG) among samples of the same patients according to the time course of Huaier administration (Figure 2). The numbers of up- and down- regulated DEGs were summarized in Table 3. We also used MAplot, Volcano plot, Scatter plot, and Heatmap plot to show the distributions of DEGs (details demonstrated at BGI website: http://www.bgitechsolutions.com/). As shown in Table 3, the average number of up-regulated genes were 2,589 out of the total 27,447 transcripts (ranging from 21 to 5,671), whereas down-regulated genes were 2,273 (ranging from 28 to 4,227).
Figure 2 demonstrates those changes by individuals. It is obvious that there were high-responders as well as low-responders to Huaier, indicating the individual genomic potential. Generally, TNM stage IV cancer patients were typical low-responder (such as Patient No.7; colorectal cancer with type I diabetes, with a long- term history of chemotherapy combinations before Huaier taking). A sudden increase of the up-regulated genes was a typical first observation, followed by the inversion to the increase of down- regulated genes. The endoscopic dissection of cancer lesion often caused massive increase of up-regulated genes (patients Nos. 9-11, 13, 14, 19), more or less 10 to 25 % of total transcripts, which might reflect the repair function and tissue reproduction by Huaier. The history of chemotherapy and radiotherapy delayed any responses (5, 6, 9, 24, 27, and 30), typically observed in patient No.
- Interestingly, just as observed in RNA editing events, the opportunistic virus infection caused the utmost increase of up- regulated genes, highlighted by asterisks in Figure 2. Huaier administration successfully down-regulated those changes to normal level, within 30 days in normal control, 3 to 6 months in cancer patients (Patient No. 5, 8, 10), with an exception in patient No.12, most severe case of stage IV lung cancer patients with multiple metastasis without any additional medical care or hospitalization.
Detailed alterations identified by KEGG pathway classification and GO functional enrichment
With quantitative analysis of DEGs, we performed Gene Ontology (GO) classification (Figure 3a) and functional enrichment. Basically, GO has three ontologies: molecular biological function, cellular component and biological process, and the results were compared by each patient with a consideration of time course of Huaier administration. We also performed KEGG pathway classification [25] according to each cancer analysed in the present study (Figure 3b) and functional enrichment (Figure 3c). Since there were no basic figure on oesophageal cancer, we did not demonstrate the molecular linkages in this cancer. There were a broad range of enriched pathway and up- and down-regulations among cases, and the alterations in transcripts were observed among all the molecular function categories, i.e., every process involved in molecular biological function, cellular component and biological process classified by KEGG pathways (the numbers of DEGs were shown in Extended Table 1).
In Figure 3, the typical GO changes were shown by three types of methods. Since abundant numbers and kinds of alterations identified so far, these panels were focused on the influence of Huaier on; a. the opportunistic infection, b. the responsible molecules altered in each cancer and signaling pathway, and, c. the responsible alterations observed in colorectal cancer by progression of the stage (from adenoma to stage IV) by the pathway functional enrichment analysis.
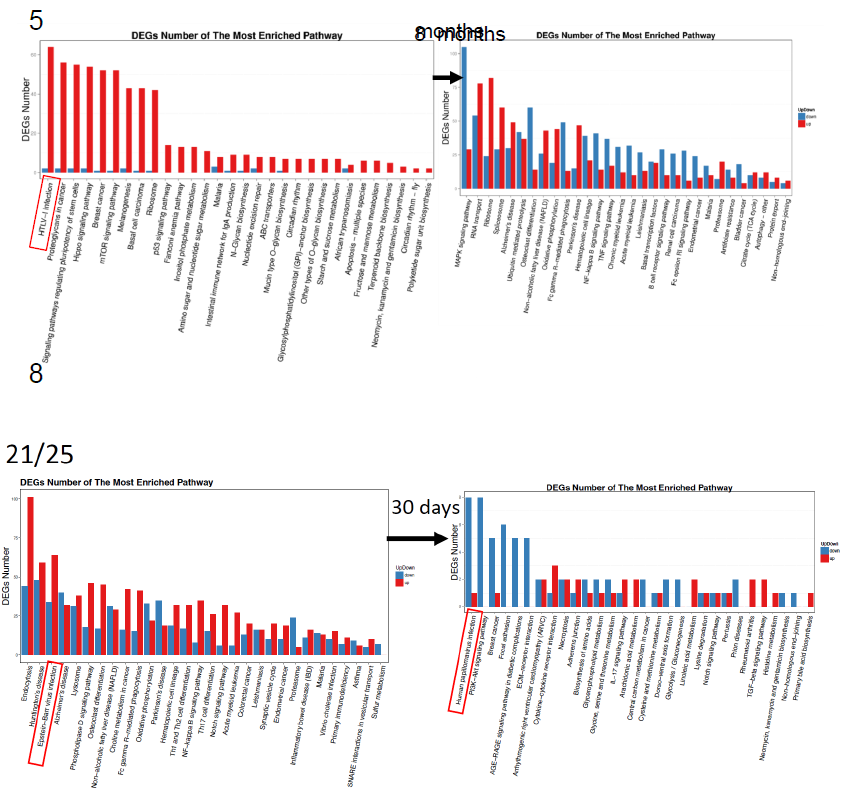
In Figure 3a, the utmost DEG changes were shown. The largest changes were highlighted by red box, the level was 3,478 in mean number (ranging from 1,826 to 5,671). These changes were indicated by asterisks in Figure 2 and Table 3, too. The virus origins in 5 patients, were; HTLV-1 or related RNA viruses (patient No. 5, 10, 12), Papilloma virus (patient No.8), and Epstein-Barr virus (patient No. 21/25) by KEGG classification. Those patients were significantly improved with administration of Huaier, however, the patient No.12 (deceased at 150 days after Huaier administration). Patient No.8 had surgical dissection of facial basal cell cancer just before the last sampling.
As shown in Figure 3a, continuous Huaier administration (20g per day) could completely rescue the molecular functions to the normal level (to recover homeostasis). The required time for the recovery seemed to vary among patients and physiological situation (from 30
days to several months), and severe case such as patient No.12 could not recover from the immunodeficiency treated by Huaier only. However, even in this case, the prognosis was improved up to 150 days, significantly improved with better QOL than predicted. Figure 3b demonstrated detailed factors and molecules with up/down regulation according to the KEGG pathway map [25] (https://www.genome.jp/kegg/). Immunological improvements seemed to be closely linked to the regulation of NFkB (nuclear factor kappa-light-chain-enhancer of activated B cells) signaling pathways, which also closely related to improve prognosis (comparison among patients 8, 21, and 12). The panels represented Huaier effects on each cancer (noted by upper left) after 90 days administration, with alterations of DEGs indicated by red colored box (up-regulation) and blue-colored box (down-regulation). In each panel, the specified DEG names were highlighted in grey box, especially related to oncogenes and tumor suppressor genes. The numbers of DEGs classified by KEGG pathways in all the patients by the time course were shown in extended Table 1.
We searched for the possible molecular changes by the comparison between the patients recovered and deceased (patients number noted in Table 1 were written on the left side of each panel). The representative changes were highlighted in grey box such as: Oncogenes; c-Myc, TERT, NRF2, c-Met, PIK3CA, k-Ra, HER2/neu, EGFR, MDM2, PDGF, PDGFR, CDK4; Tumor
suppressors; Rb, p53, PTEN, p16, Smad4, BRAC2, IKN4a/ARF, AXIN, KEAP1, ARIDIA, and DNA repair genes; hMLH1, hMSH2, hMSH3, hMSH6.
This could be typically observed in the results obtained in the patient No.12, by up-regulation of DEGs responsible for signal transfer in NFkB and TGF-b (transforming growth factor beta) signaling pathway [28], with further influences to related regulatory systems.
Immunological systems and pathways are multi-functional biosystems, and both deficiency as well as excessive functional enhancement cause various disorders and diseases. The results obtained in patient No.12 provided a hint for the major molecules and pathways. The continuous up-regulation of DEGs was observed which was responsible for signal transfer in NFkB and TGF-b signaling pathway [17], whereas the other patients successfully recovered showed the down-regulation of these systems at a certain point of Huaier complementally therapy
(Figure 3b). We provided the copy number changes which might link to NFkB by NFkB subunits 1 and 2 (Extended Table 2); molecules related to the production of iPS cells (KIT, Myc, OCT3/4, SOX2, LIN28A and NANOG) (Extended Table 3). However, there observed no significant increase and/or decrease among patients. Not only the genes correlating with signal transduction and carcinogenesis, many other genes of physiological importance should be also affected as a link to main pathways for maintenance of homeostasis. More importantly, almost all the transcriptional misregulation were rescued chiefly by the up- regulated transcription factors, with the down-regulation of Nuclear receptor co-repressor (N-CoR) in TRANSCRIPTIONAL MISREGULATION IN CANCER panel of Figure 3b. As reported before [14], Figure 3b confirmed the transcription control rescue in Hippo signaling pathway [3] and corresponding Wnt and MAPK signaling pathways.
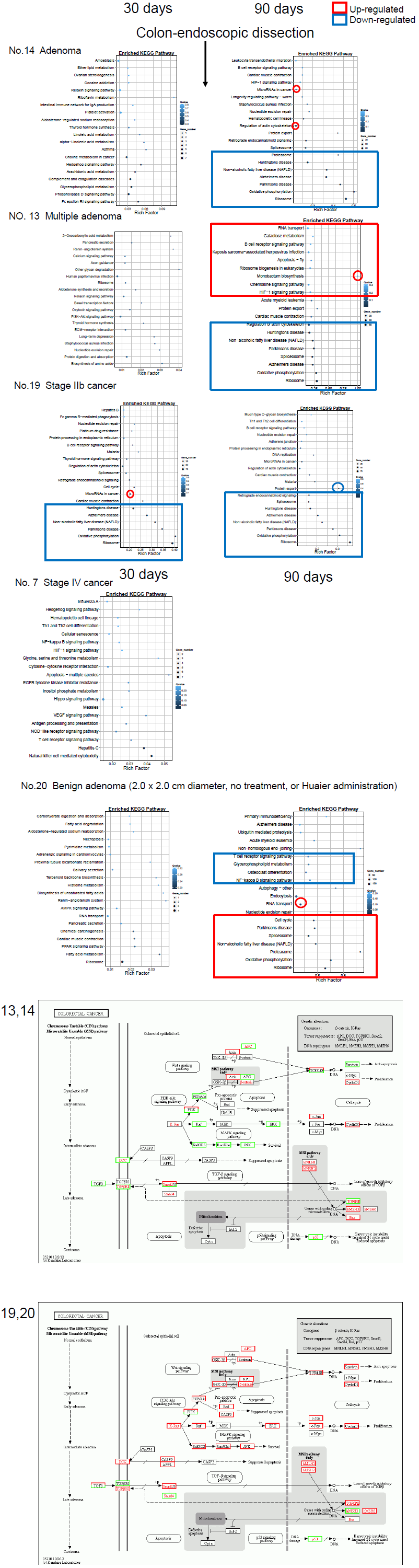
The functional enrichment classified by KEGG pathway [25] specifically using the results of every TNM stages of colorectal cancer were shown in Figure 3c. Blue box represents down- regulated genes and pathways, and red box indicates up-regulated ones. The panels were placed according to the stage of cancer progression, and the last panels showed the comparison of Huaier effects on benign adenoma vs. adenocarcinoma. The Patient No. 20 was a control without Huaier administration, with a latent progression of benign adenoma (dissected endoscopically one month after the final sampling). There were significantly more suppressive effects in the responsible oncogenes and tumor suppressors in benign adenoma cases than progressive adenocarcinoma cases. Details of relating signaling pathways were shown in Figure 3b.
The results obtained from the Patient No.20 indicated that, on the early process of cancer progression, the immunological pathways were down-regulated, whereas cell cycle signaling and oxidative stress accumulation were up-regulated. Interestingly, the alterations in the oxidative stress reduction systems by Huaier were identical in different cancers, especially strong similarity found in hepatoma and meningioma (Patient No.8 and No.10 in Fig. 3b), which might suggest the required rescue system and/or functional damaging factors by stress accumulation were identical in those two organs, liver and central nervous system. The typical low responder to Huaier, the patient No.7, did not demonstrate any significant DEGs alterations in any pathways.
3.6 Huaier effects on transcriptional factors and their quantitative rearrangements in vivo
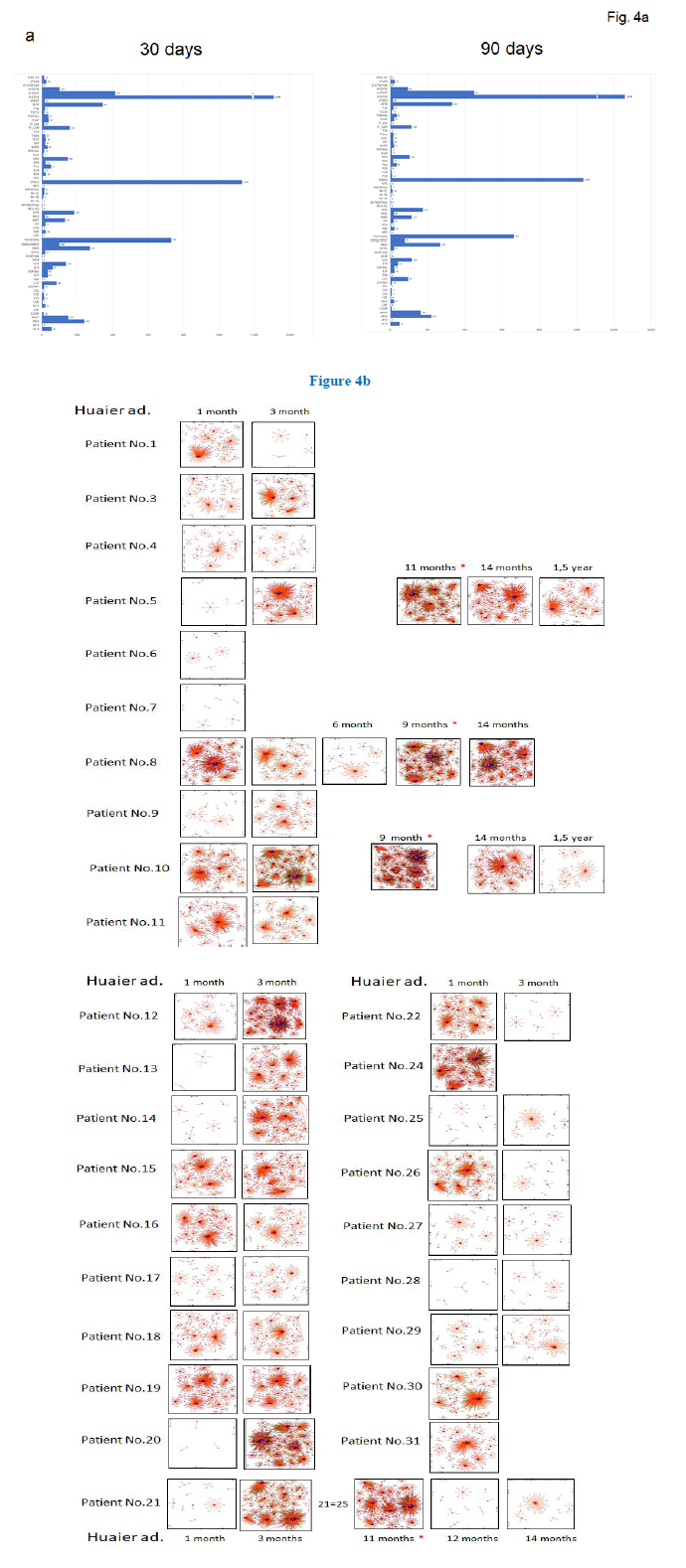
We analyzed the relationship of those quantitative and qualitative DEG alterations to encoding transcription factors (TF). Figure 4a showed transcription factor families and their quantitative changes classified by the DEGs which belonged to. Total 7,664 DEGs coding transcriptional factors were altered to rescue and/or modify transcriptional misregulation noted in Figure 3b, within 30 days after Huaier administration. We have already indicated Huaier effects on the rescue of transcription control using Drosophila mutants [3], and the present study provides the systematic and quantitative background. These dynamic changes in transcription factors might explain the enormous range of transcription control rescue by Huaier. As shown in Table 4, 66 kinds of transcription families were altered and modified by up/down regulation of encoding DEGs, maximum differences identified are 2,535 DEGs within 30 days, and 2.293 DEGS at 90 days after Huaier administration. Again, we have to emphasise that these effects were observed with no side effects, toxicity, which resulted in the improvement of QOL of the patients.
As a consequence, the process of cancer recovery was clearly demonstrated by the TF-DEG network changes (Figure 4b). These functional linkage map of TF-DEG network were simplified form to indicate that the Huaier effects depended on the time course of Huaier administration. The red and green dots represent the up- regulated and down-regulated DEGs, respectively. Purple ball represents each TF as a core. The greater the core size, the more DEGs regulated by the core transcription factor. The results indicated the extreme lineage of gene expression was evoked soon after the Huaier administration in cancer patients as shown in the former tables and figures, suggesting the genetic changes would be based on transcription control via small nuclear RNAs.
3.7 Analysis of differentially expressed small RNAs (DSGs); siRNA (30 nucleotides), piRNA (24 nucleotides), and miRNA (21-22 nucleotides)
The numbers of detected small non-coding RNA for each sample were summarized in Table 5. Table 6 showed the results of different kinds of sRNA in all the samples (the ratio of each components was shown in Extended Table 4). The structure of miRNAs and the sequences of the novel small RNAs were deposited to the NCBI GEO (GSE157086), and shown and stored in precursor.tar.gz.
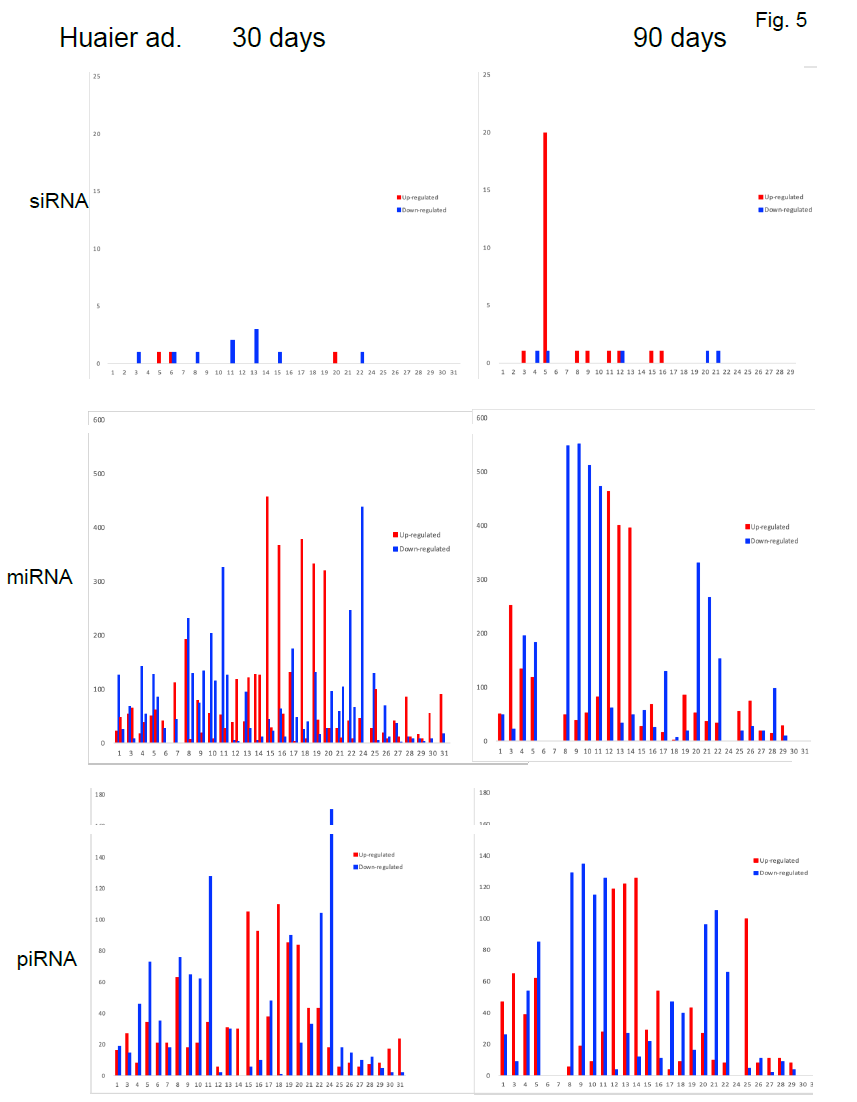
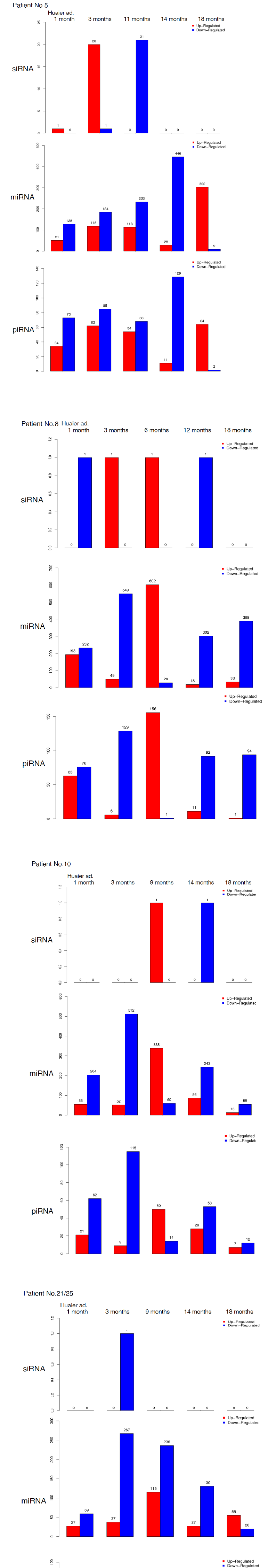
The small RNA expression level was calculated by using TPM [32] (Transcripts Per Kilobase Million). The analyzed differentially expressed small RNAs (DESs) were designated to compare between samples. DESs in each pairwise were shown in Figure 5. The changes in each patient by time course were demonstrated in Figure 6. In the patient No.5 (oesophageal squamous cell cancer with lung metastasis, we identified 20 up-regulation at 90 days after Huaier administration, and with one down-regulated element, total
21 novel elements were all down-regulated at 11 months after administration (Figure 6). The slow onset of novel siRNA function was also noteworthy in this patient. Compared with the changes in transcriptomes (Figure 2) and TF-DEG network (Figure 4b), these siRNAs seemed to contribute to the drastic alterations of downregulation of the transcriptomes, followed by up-regulation of over 3,000 transcripts later. DEG changes were also associated with the drastic up-regulation of transcription factor families.
SiRNA has been reported to interfere with the expression of specific genes with complementary nucleotide sequences by degrading mRNA after transcription, preventing translation [33]. In addition, siRNAs typically work by cleaving the mRNA before translation, and have 100% complementarity, thus very tight target specificity [34]. These sequential genetic events found in patient No.5 in total would rescue the misregulation of translational and transcriptional systems in vivo, required for cancer recovery. In the other patients, no significant siRNA function was identified.
On the other hand, as shown in Table 5, average 1,003 miRNAs (ranging from 685 to 1,378) per sample were identified within 90 days after Huaier administration. DESs in each pair wise was shown in Table 5, Figure 5 and Figure 6, revealed several hundreds of miRNA alterations in each patient, and over a half of total identified miRNAs were differentially expressed. Figure 6 well demonstrated miRNA-mediating gene silencing effects with a comparison to the results in Figure 2. For example, the significant changes in up- and down-regulated DEGs were in inverse proportion to those observed in quantitative DES changes, such as significant increase of up-regulated miRNAs in the patient No. 8 (Figure 6, middle column) resulted in the significant decrease of both up- and down-regulated DEGs in Figure 2. In contrast, the significant decrease of up-regulated DESs in Figure 6, related to human papillomavirus infection suggested by Figure 3a, drastically increased both up- and down-regulation of DEGs up to 3,330 and 3,505, respectively (12.2% of the total DEGs). The total numbers of DEG changes in the Patient No.8 became 85% of total transcripts (23,210/27,447).
Three months after Huaier administration of the patient 10, the increase of down-regulated miRNA was observed when massive increase of down-regulated DEGs (4,274; 15.6% of the total). In both patients No. 8 and 10, the opportunistic virus infection might influence the drastic decrease in number of miRNAs.
In the present study, haemoglobin counts in TNM stage IV colorectal cancer improved from 6.0 to 12.0 g/dl after 4 months’ treatment with 20g Huaier per day. MiRNAs also play crucial roles in the regulation of complex enzymatic cascades including the haemostatic blood coagulation system [35]. Large scale studies of functional miRNA targeting have recently uncovered rationale therapeutic targets in the haemostatic system [36]. Piwi-interacting RNA (piRNA) is the largest class of small non-coding RNA molecules (containing about 24 nucleotides in the present study) expressed in animal cells [37]. PiRNAs were reported distinct from microRNA (miRNA) in size (24 nucleotides as opposed to 21–22 nt), lack of sequence conservation, increased complexity. However, as shown in Figure 6, the dynamic statistics of piRNA revealed similar movement as miRNAs, although the total numbers of the known and the novel piRNAs were much smaller compared with miRNAs. The present study, together with other small RNA information, shed lights into the function of piRNA function related to RNA-mediated adaptive immunity against transposon expansions and invasions, especially in the case with the opportunistic infection.
Figure 3: Pathway classification and functional enrichment of DEGs. a. Pathway functional enrichment results for up/down regulation genes. X axis represents the terms of pathway. Y axis represents the number of up/down regulation genes. b. The detailed analysis of the Huaier effects on each cancer by KEGG biological pathways [25]. The pathways responsible for cancer progression, stress reduction, and major immunological responses related to NFkB and TGFb signaling pathway were specifically compared among patients with recovery and poor prognosis. c. Functional enrichment of the factors classified by KEGG pathways. The color indicates the q-value (high: white, low: blue), the lower q-value indicates the more significant enrichment. Point size indicates DEG number (the bigger dots refer to larger amount). Rich Factor refers to the value of enrichment factor, which is the quotient of foreground value (the number of DEGs) and background value (total gene amount). The larger the value, the more significant enrichment.
Figure 3a
Figure 3: Pathway classification and functional enrichment of DEGs. a. Pathway functional enrichment results for up/down regulation genes. X axis represents the terms of pathway. Y axis represents the number of up/down regulation genes. b. The detailed analysis of the Huaier effects on each cancer by KEGG biological pathways [25]. The pathways responsible for cancer progression, stress reduction, and major immunological responses related to NFkB and TGFb signaling pathway were specifically compared among patients with recovery and poor prognosis. c. Functional enrichment of the factors classified by KEGG pathways. The color indicates the q-value (high: white, low: blue), the lower q-value indicates the more significant enrichment. Point size indicates DEG number (the bigger dots refer to larger amount). Rich Factor refers to the value of enrichment factor, which is the quotient of foreground value (the number of DEGs) and background value (total gene amount). The larger the value, the more significant enrichment.
Figure 3b
Figure 3: Pathway classification and functional enrichment of DEGs. a. Pathway functional enrichment results for up/down regulation genes. X axis represents the terms of pathway. Y axis represents the number of up/down regulation genes. b. The detailed analysis of the Huaier effects on each cancer by KEGG biological pathways [25]. The pathways responsible for cancer progression, stress reduction, and major immunological responses related to NFkB and TGFb signaling pathway were specifically compared among patients with recovery and poor prognosis. c. Functional enrichment of the factors classified by KEGG pathways. The color indicates the q-value (high: white, low: blue), the lower q-value indicates the more significant enrichment. Point size indicates DEG number (the bigger dots refer to larger amount). Rich Factor refers to the value of enrichment factor, which is the quotient of foreground value (the number of DEGs) and background value (total gene amount). The larger the value, the more significant enrichment.
Figure 3c
Figure 4: TF-DEG network in each patient by the time course of Huaier administration. a. TF families classified by the encoding DEGs, with each number of up/down regulation. b. TF-DEG predicted network. The red and green dots represent the up-regulated and down-regulated DEGs, respectively. Purple ball represents transcription factor, the greater the node the more DEGs the transcription factor regulate.
4. Discussion
The results obtained in the present study clearly indicated the anti- cancer effects of Huaier were based on the rescue of transcription control which contributes to maintain and restore homeostasis. It was based on the individual genomic potential to undergo the drastic changes among genome wide range, and not influenced by cancer origin, stage, and personal differences, except the extrinsic factors like virus infections. It is surprising to see the genomic flexibility and capability, i.e., individual genome potential which can tolerate extensive genomic and genetic changes. At the beginning, the average number of RNA editing events such as SNP and INDL showed that Huaier administration induced approximately 4 times higher movability of the genome, ranging 2 times to 7 times higher than those detected as identified among normal population [16]. The statistics of splicing events resulted in expanded the plasticity and flexibility, and consequently enhanced the genomic capability to manage or cope with the following processed in transcription and translation. These results brought over 24,344 novel transcripts, and also caused the drastic rearrangements in number of up/down-regulated transcripts. The ratio of the quantitative changes in translation at maximum was 85% (23,210/27,447). These changes in transcripts were successfully demonstrated as Transcription Factor (TF)- Differentially Expressed Genes (DEG) lineage map (see Figure 4). The process and time course toward cancer recovery was clearly shown in this figure sets, and more importantly, the recovery from the influence of environmental stress typically indicated by the opportunistic virus infections could be observed even in normal healthy controls, too. The duration period required for the recovery was dependent on the physiological status in each individual, one month for healthy control and about 5-6 months for cancer patients. This observation was identical to the clinical observation reported previously.
The comparison between the patients who successfully recovered and failed revealed that the major difference existed in immunological factors, especially observed in the control in NFkB and TGF-b signalling pathways (Fig. 3b). The cancer recovery seemed to require practical inhibition and down-regulation of NFkB and TGF-b and functionally linked molecules after significant up-regulation. These sequential changes occurred by the time course of Huaier administration, and the same was true to the other signalling pathways. The sequential control of immunological reactions, both activation as well as modulation of excessive
enhancement, were required for the recovery of cancer, by NFkB and TGF-b signaling pathways. The significant effects on the recovery from oesophageal cancer (squamous cell cancer and adenocarcinoma) was observed, and that even with multiple metastasis (patient No. 5, 9, 28, and 29). Interestingly, the patient No.29 with lung metastasis was successfully recovered judged by CT images and endoscopic examination, without any significant increase of both up- and down-regulated transcripts. The clinical examinations showed a good coincidence to the results obtained here, with a significant improvement/elimination of metastatic lesion, and that without any more progression of the disease.
In addition, it is emphasized here that genomic plasticity has been continuously observed even in normal healthy individuals. It seems that, even without any medical problems and disorders, every person should have influences from environmental stress, infective agents and ageing processes which disrupted physiological functions. Huaier revealed its efficacy even on the healthy individual, according to the extent of functional disruption. The observation provided of the potential of Huaier to rescue the silent disruptions in molecular level to recover the normal homeostasis. The consequent restoration of transcription control seemed to be the major molecular mechanisms induced by Huaier administration. The recovery of cancer was easily identified to see the quantitative level of regulation in transcriptomes, with a confirmation of CT, MRI, and X-ray images of the patients (data not shown). However, there were many patients without significant gene expression changes. It is surprising to find no significant quantitative changes found specially in the patient with oesophageal cancer even with multiple metastasis (patient No.29).
Huaier administration induced as such many clusters of oncogenes and tumor suppressor genes, and that in both up- and down- regulation. There have not been reported yet about the possible compounds or natural herbs to show the similar efficacy in vivo. Many trials were performed, but failed from the strong toxicity even to in vitro cultured cells. In contrast, Huaier influenced almost all the main factors correlating to the carcinogenesis and tumor suppressor functions, and that contained a whole molecular system linked by those molecules. Huaier seemed to compensate for the limitation of targeting system by conventional chemotherapy, by the rescue of all the required physiological functions. Clinically, the rescued functions seemed to be responsible for the specific cell death in occult metastasis in the peripheral blood (prevent the recurrence and relapse of cancer), and for the recovery from anemia and hypoleukocytemia.
The present study also increased the information of the many novel miRNAs significantly contributed to cancer recovery, and further investigation of these molecules in vitro and in vivo are expected. MiRNAs have been reported to typically silence genes by repression of translation, and with broader specificity than siRNA, to functions in RNA silencing and post-transcriptional regulation of gene expression [34, 38, 39]. The small nuclear RNAs were also affected, and the consequent gene-silencing were also detected by miRNA-mediated post-transcription control, and numerous novel molecules were identified (deposited to NCBI GEO: GSE157086). While many unusual expressions of long ncRNAs in disease states have been reported [40], it is still remained unknown how those ncRNAs function in causing diseases. Transcriptome analyses here identified numerous differentially expressed novel ncRNAs in various forms of cancer such as colorectal carcinoma and hepatocellular carcinoma. Many SNPs have been found to be associated with certain disease conditions, which function in association with targeting ncRNAs in the transcriptional regulation [40]. All the novel small RNA (DSG) sequences identified were also deposited to the NCBI GEO (GSE157086).
In addition, we identified the similar drastic changes derived from opportunistic infections, RNA virus infection as COVID-19. The severity of symptoms and the recovery period dependent on the genomic potential, too, and in fact, the level of severity to the lethal situation in case were drastically decreased by clinical trial of Huaier against COVID-19 (WHO ID:NCT04291053). Huaier also prevented reinfection after 3-6 months.
4. Conclusion
The present study provided almost all the genomic and genetic events in cancer patients, especially identified from the recovered patients. The recovery from cancer was identical to the recovery of homeostasis, through the modulation and rescue of transcriptional dysregulation using the multi-elements involved in the series of each process in transcription and translation. SncRNAs (small non- coding RNAs) together with dynamic RNA editing in SNP variation directly influenced these processes, which resulted in large-scale alterations in DEGs. Huaier effects could be quantitatively and widely observed in the biophysiological systems classified by Gene Ontology, and in total were derived from the
physiological capability and flexibility of each patient. The individual genomic potential to undergo those changes and modulations is the key for cancer recovery. The effect was spontaneous reaction indirectly induced by Huaier administration.
Although there have been many reports regarding specific cancer cell death as anti-cancer effects of Huaier [1-3, 14, 28], it should be emphasized the efficacy of Huaier on the significant potential for the recovery from opportunistic virus infections, and also for tissue reproduction to regenerate the damaged tissues and organs after the removal and/or cancer cells in the lesion. The elimination of occult metastasis (after surgical treatment) should be emphasized, too. These effects are spontaneous, and dose- and duration-dependent. It is the major reason why Huaier does not cause any side effects and/or toxicity.
5. Acknowledgements
The authors wish to thank cancer patient volunteers and many healthy volunteers kindly collaborated with the present study. We also wish to thank Prof. Dr. Tongbiao Zhao, Professor of Stem Cell and Immunology, Institute of Zoology, Chinese Academy of Sciences, China, for critical revision of the manuscript. The present study was grant-in-aid from QiDong Gaitianli Medicines Co., Ltd. And Japan Kampo NewMedicine, Co., Ltd.
6. Author contributions
T.T., M.T., designed the study from the clinical observation of the cancer patients with Huaier treatment (as a complementally therapy), and managed the sampling and clinical assessment of the patient volunteers, statistically analyzed the data, and drafted the manuscript. F.T., H.L., managed total RNA and small nuclear RNA sequencing and conducted systematic analysis of the data. Z.L., D.W., contributed to the provision of Huaier granules and clinical evaluation of the data, especially focused on Immunological evaluation.
Conflict of Interest
The authors have no competing interest to declare.
Author information
The authors have no competing interest to declare. Readers are welcome to comment on the paper. Correspondence should be addressed to T.T. (ttanaka@bradeion.com) and M.T. (manami- tanaka@bradeion.com), and those researchers contributed equally to this work. Requests for Huaier extract and commercially- available granules should be addressed to D.W. (teii@newkampo.co.jp) and M.T. (manami-tanaka@bradeion.com)
References
- Chen Q, Shu, C, Laurence, AD, Chen Y, Peng BG, et al. Effect of Huaier granule on recurrence after curative resection of HCC: A multicentre, randomised clinical trial. Gut 67 (2018): 2006–2016.
- Zhang Y, Wang X, Chen T. Efficacy of Huaier granule in patients with breast cancer. Clin Transl Oncol 21 (2019): 588-595.
- Tanaka T, Suzuki T, Nakamura J, Kawamura Y, Sadahiro S, et al. Huaier Regulates Cell Fate by the Rescue of Disrupted Transcription Control in the Hippo Signaling Pathway. Arch. Clin Biomed Res 1 (2017): 179-199.
- Mo JS, Park JW, Guan KL. The Hippo signaling pathway in stem cell biology and cancer. EMBO Rep 25 (2014): 642-656.
- Zhang H, Lang TY, Zou DL, Zhou L, Lou M, et al. miR- 520b Promotes Breast Cancer Stemness Through Hippo/YAP Signaling Pathway. Onco Targets Ther 12 (2019):11691-11700.
- Wu L, Yang Targeting the Hippo Pathway for Breast Cancer Therapy. Cancers (Basel) 10 (2018): 422-437.
- Wei C, Wang Y, Li X. The role of Hippo signal pathway in breast cancer metastasis. Onco Targets Ther 11 (2018): 2185-2193.
- Feng X, Zhang M, Wang B, Zhou C, Mu Y, et al. CRABP2 regulates invasion and metastasis of breast cancer through hippo pathway dependent on ER status. J Exp Clin Cancer Res 38 (2019): 361-368.
- Shen J, Cao B, Wang Y, Ma C, Zeng Z, et Hippo component YAP promotes focal adhesion and tumour aggressiveness via transcriptionally activating THBS1/FAK signalling in breast cancer. J Exp Clin Cancer Res 37(2018): 175-191.
- Qiao K, Ning S, Wan L, Wu H, Wang Q, et al, Pang D. LINC00673 is activated by YY1 and promotes the proliferation of breast cancer cells via the miR-515- 5p/MARK4Hippo signaling pathway. J Exp Clin Cancer Res 38 (2019): 418-432.
- Wang S, Su X, Xu M, Xiao X, Li X, et al. Exosomes secreted by mesenchymal stromal/stem cell-derived adipocytes promote breast cancer cell growth via activation of Hippo signaling pathway. Stem Cell Res Ther 10 (2019): 117-128.
- Jiang Y, Sun A, ZhaoY, Ying W, Sun H, et al. Proteomics identifies new therapeutic targets of early-stage hepatocellular carcinoma. Nature 567 (2019): 257-261.
- Song Y, Li L, Ou Y, Gao Z, Li E, et al. Identification of genomic alterations in oesophageal squamous cell cancer. Nature 509 (2014): 91-95.
- Song X, Li Y, Zhang H, Yang, Q. The anticancer effect of Huaier (Review). Oncol. Rep. 34 (2015): 12-21.
- Wang X, Wang N, Cheung F, Lao L, Li C, et al. Chinese medicines for prevention and treatment of human hepatocellular carcinoma: current progress on pharmacological actions and mechanisms. J Integr Med 13 (2015): 142-164.
- Peng Z, Cheng Y, Tan BC, Kang L, Tian Z, et al.Comprehensive analysis of RNA-Seq data reveals extensive RNA editing in a human transcriptome. Nat Biotechnol 30 (2012): 253-260.
- Ramsköld D,Luo S,Wang Y, Li R, Deng Q, et al. Full- Length mRNA-Seq from single cell levels of RNA and individual circulating tumor cells. Nat Biotechnol 30 (2013): 777-782
- Cock PJA, Fields CJ, Goto N, Heuer ML, Rice PM. The Sanger FASTQ file format for sequences with quality scores, and the Solexa/Illumina FASTQ variants. Nucleic Acids Res 38 (2009): 1767-1771.
- Kim D, Langmead B, Salzberg SL. HISAT: a fast spliced aligner with low memory requirements Daehwan HHS Public Access. Nat Methods 12 (2015): 357-360.
- Robinson JT, Thorvaldsdóttir H, Winckler W, Guttman M, Lander ES, et al. Integrative Genome Viewer. Nat Biotechnol 29 (2011): 24-26.
- Pertea M, Pertea GM, Antonescu CM, Chang T, Mendell JT, et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat Biotechnol 33 (2015): 290-295.
- Trapnell C. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc 7 (2013): 562-578.
- Kong L, Zhang Y, Ye Z, Liu X, Zhao S, et al. CPC: assess the protein-coding potential of transcripts using sequence features and support vector machine. Nucleic Acids Res 35 (2007): 345-349.
- DePristo MA, Banks E, Poplin R, Garimella KV Maguire JR, et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Genet 43 (2011): 491-498.
- Kanehisa M, Araki M, Goto S, Hattori M, Hirakawa M, et al. KEGG for linking genomes to life and the Nucleic Acids Res 36 (2008): 480-484.
- Griffiths-Jones S. miRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res 34 (2006): D140-D144.
- Nawrocki EP, Burge SW, Bateman A, Daub J, Eberhardt RY, et al. Rfam 12.0: Updates to the RNA families database. Nucleic Acids Res 43 (2015): D130-D137.
- Li C, Wu X, Zhang H, Yang G, Hao M, et al. A Huaier polysaccharide inhibits hepatocellular carcinoma growth and metastasis. Tumour Biol 36 (2015): 1739-1745.
- Shen S, Park JW, Lu Z, Lin L, Henry MD, et al. rMATS: Robust and flexible detection of differential alternative splicing from replicate RNA-Seq data. Proc. Natl Acad Sci USA 111 (2014): E5593-E5601.
- Langmead B, Trapnell C, Pop M. Salzberg SL. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10 (2009).
- Li B. Dewey CN. RSEM: Accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12 (2011).
- Li B, Ruotti V, Stewart RM, Thomson JA, Dewey CN. RNA-Seq gene expression estimation with read mapping uncertainty. Bioinformatics 26 (2009): 493-500.
- Laganà A, Veneziano D, Russo F, Pulvirenti A, Giugno R, et al. Computational design of artificial RNA molecules for gene regulation. Methods Mol Biol 1269 (2015): 393- 412.
- Mack, G. S. MicroRNA gets down to business. Nat. Biotechnol 25 (2007): 631-638.
- Teruel-Montoya R, Rosendaal FR, Martínez C. MicroRNAs in hemostasis. J. Thrombosis and Haemostasis 13 (2015): 170-181.
- Nourse J, Braun J, Lackner K, Hüttelmaier S, Danckwardt
- Large-scale identification of functional microRNA targeting reveals cooperative regulation of the hemostatic system. J Thrombosis and Haemostasis 16 (2018): 2233- 2245.
- Seto AG, Kingston RE, & Lau, NC. The Coming of Age for Piwi Proteins. Mol Cell 26 (2007): 603-609.
- Fabian MR, Sonenberg N, Filipowicz W. Regulation of mRNA Translation and Stability by microRNAs. Annu Rev Biochem 79 (2010): 351-379.
- Bartel DP. MicroRNAs: Target Recognition and Regulatory Functions. Cell 136 (2009): 215-233.
- Ma L, Cao J, Liu L, Du Q, Li Z, et al. Lncbook: A curated knowledgebase of human long non-coding rnas. Nucleic Acids Res 47 (2019): D128-D134.




 Impact Factor:
* 5.8
Impact Factor:
* 5.8
 Acceptance Rate:
71.20%
Acceptance Rate:
71.20%
 Time to first decision: 10.4 days
Time to first decision: 10.4 days
 Time from article received to acceptance:
2-3 weeks
Time from article received to acceptance:
2-3 weeks
