In Vitro Profiling of Application Ready Human Surrogate Primary Progenitor Stromal Cell Fractions
Vasanthi Dasari, Paparao Bolimera, Leela Krishna Gorthi, Subhadra Dravida*
Transcell Oncologics Pvt Ltd, Technology Business Incubator – ASPIRE, University of Hyderabad, School of Life Sciences, Gachibowli, Hyderabad, Telangana, India
*Corresponding author: Subhadra Dravida, Transcell Oncologics Pvt Ltd, Technology Business Incubator – ASPIRE, University of Hyderabad, School of Life Sciences, Gachibowli, Hyderabad, Telangana, India
Received: 30 May 2022; Accepted: 08 June 2022; Published: 16 June 2022
Article Information
Citation: Vasanthi Dasari, Paparao Bolimera, Leela Krishna Gorthi, Subhadra Dravida. In Vitro Profiling of Application Ready Human Surrogate Primary Progenitor Stromal Cell Fractions. Archives of Clinical and Biomedical Research 6 (2022): 536-552.
View / Download Pdf Share at FacebookAbstract
Background: Human progenitor, primary cell-based models are emerging as alternatives to animal systems in discovery and development stages of pharma and biopharmaceutical candidates meant for clinical application. In vitro platforms available for such use including for assessments of human neuro related toxicities and virulence have been rarely reported. We characterized biological discard sourced primary progenitor cells configured as microphysiological systems in vitro to establish phenotype and genotype signatures with reference to pluripotency and neuronal lineage.
Methods and results: Tissue based method of stromal cell population isolation method, MACS to generate induced pluripotent cell fractions, conditioning medium to coax neuronal morphologies were employed to generate human microphysiological in vitro primary cellular platforms. The stromal cells, sorted cells and coaxed neuronal like cells (CNC) manifest distinct phenotypes and genotypes. The sorted cells show enrichment of pluripotent gene markers compared to heterogeneous stromal precursor cell population while CNCs show characteristic complementation with Nestin, EN, Tra1, Musashi, NFL genes.
Conclusion: We report here an in vitro human biological discard derived primary progenitor cell-based system with innate pluripotent genotype inclined towards neuronal lineage upon induction. Our study offers a possibility that human derived microphysiological systems generated in scale owing to the source advantage may bring novel insights into the design and choice of next generation assessment methodologies in testing neurotoxicity, neurovirulence like non-clinical safety parameters surrounding vaccines and drugs for safe delivery in line with FDA’s advice to the global industry.
Keywords
<p>Stromal cells; SSEA4; Pluripotent; Stem cells; Neuronal lineage</p>
Article Details
1. Introduction
Clinical in vivo studies have traditionally followed the establishment of a proof-of-concept in vitro [1, 2]. The in vitro model is therefore of primary importance, and much thought should go into the choice of the same. As with all model systems, the availability of the model, cost, and most importantly, the degree of resemblance to the human physiology, the degree to which the model reproduces the “real thing”, are all considerations in the choice of the model system. This is especially true in human neurological microphysiology modeling where tissue biopsies are difficult to acquire and often difficult to obtain large and viable cell culture systems to utilize in assay techniques. Immortalized cells can be used to certain extent but have significant genetic, transcriptional and protein expression changes in the assay’s baseline data set, affecting the readouts [3, 4]. Stem cell-based systems and especially human sourced are increasingly being used as surrogates for whole organisms, viz mice, rats, as part of a community-wide shift away from animal models [5-7].
Primary progenitor cells are preferable to established cell-lines for neurobiological studies, for more than one reason [8, 9]. The latter acquire polymorphisms with time and passage and a population often comprises a diversity of genetic backgrounds. Primary progenitor cells don’t suffer from this problem, and have the other qualities desirable in a cellular model system built on. Progenitor cells can be sourced from plentifully available biological discards like umbilical cord tissue collected at human birth [10]. Such tissues house stromal (mesenchymal) progenitor cells, which can give rise to both multipotent and human induced pluripotent cell populations (HiPSCs) in appropriate conditions, and thence to human neurons [11, 12]. If this transformation can be achieved at scale/ throughput, it will enable, and expand, the use of such “manufactured” neuronal cells in the biotech and pharma industries, especially in the testing of neurotoxicity and neurovirulence like non-clinical safety related end points. In the context of the biotech and pharma industry, such bespoke human surrogate cell based micro physiological systems can be used as real time platforms for the assay of neurovirulence and neurotoxicity in drugs, vaccines, and cosmetics meant for human consumption [13, 14].
In this article, we present three of our laboratory derived cellular systems produced from donated biological discard, i.e human umbilical cord. We detail their phenotype and genotype to establish the signature cellular features that can be leveraged for various applications and use cases.
2. Materials and Methods
2.1 Primary cells harvest and In vitro Configuration
Stromal cell isolation: Umbilical cord (UC) samples received from donors (3) after obtaining informed consent were cleaned in sterile PBS containing antibiotic (two times) in the biosafety cabinet. The tissue was cut into small pieces (1-2 cm) with a sterile blade. These pieces were covered in knockout DMEM supplemented with 10% FBS, 20 ng/ml EGF, 10 ng/ml b-FGF, 2 mM L-glutamine, 100 IU/ml penicillin/streptomycin and incubated at 37°C in an atmosphere of 5% CO2 for 10-15 days (while changing the media every 2-3 days). The tissue pieces were discarded after the cells outgrowth reached a confluency of 50-80% and labeled as P0 (passage) stage. Cells were digested with 0.25% trypsin, centrifuged and sub-cultured in fresh flasks for subsequent passages.
2.1.1 Enrichment and fractionation: Stromal cells were collected at P3 stage processed for the isolation of SSEA4 positive population using Miltenyi Biotech kit in accordance with the protocol mentioned in the user’s manual. The enrichment procedure fractionates the positive cell population that is antibody bound (sorted cells) and unbound cells (flow through). The sorted cells were cultured in medium containing DMEM supplemented with 10% FBS, 20 ng/ml EGF, 10 ng/ml b-FGF, 2 mM L-glutamine and 100 IU/ml penicillin/streptomycin.
2.1.2 Generation of Coaxed Neuronal like Cells (CNCs): P3 stage sorted cells were conditioned for 48hrs in Neurobasal media containing N2 (1%), B27 (2%), b-FGF (10 ng/ml), EGF (20 ng/ml), GDNF (20 ng/ml), retinoic acid (20 ng/ml), db-cAMP (50 µM) and interleukin-1b (200 pg/ml).
Phenotyping
Microscopy:
Phase Contrast: Live cells were profiled under inverted phase contrast microscope at 20X magnification at >70% confluency unless otherwise specified.
2.1.3 Immunostaining & Confocal imaging: Cells were grown on precleaned 75 mm coverslips for 24 hrs. Cells were rinsed and fixed in freshly prepared 4% paraformaldehyde for 15 min. The cells were permeabilized with a blocking solution containing 0.1% Triton X-100, 1% BSA in PBS for 1 hr. Cells were incubated with a primary antibody (FITC conjugated SSEA4 antigen-Miltenyi) diluted in PBS (1:50) for 30 min at room temperature. Cells were rinsed with PBS once for 5min before proceeding for PI staining (1:5) for 15 min in PBS at room temperature. The cells were gently rinsed three times each in PBS and mounted in a mounting medium containing DAPI (1µg/ml; Molecular probes/ Invitrogen).
Leica TCS SP8 microscope inverted-based microscope equipped with 20, 40 and 63X objectives was used to obtain confocal images. FITC was excited at 488 nm; the emission filters 498–551 were used to collect the signal. PI was excited at 568 nm; emission filters 648– 701 were used to collect the signal. Images were then treated with Fiji to obtain fluorescent images using maximum Z-projections [15].
2.1.4 Karyotyping: Cells were sub-cultured at 1:4 dilution before harvesting and were collected with Trypsin EDTA prior to colchicine treatment. After centrifugation, the cells were resuspended and spread on a slide, fixed with aceto-methanol. Fixed cells were dropped on glass slides and allowed to air dry [16]. Chromosomes were G-banded after staining with Giemsa dye. A total of 5 metaphases were analyzed using a Leica microscope.
2.1.5 Population Doubling Time (PDT) scoring: Cells were seeded in a 12 well plate at a density of 0.02m and counted using Tali (Invitrogen) or by manual method using Hemocytometer for every 12hr until 100 hrs. PDT was calculated using the formula: t X Log2/(LogC2/LogC1) [ t= duration in hours, Log = 10 based Log, C1 = 1st cell count, C2 = 2nd cell count] (ATCC.org).
2.2 Flow cytometric analysis
Cells were detached and stained with FITC conjugated monoclonal antibody against SSEA4 antigen (Miltenyi) for 10 min at 4 C in a buffer containing antibody according to the manufacturer's protocol.
Prior to the analysis, cells were washed by adding 1ml of buffer and were resuspended in PBS and subjected to FACS analysis. At least 10,000 cells were analyzed per sample using FACS Calibur flow cytometer (BD biosciences).
2.3 RNA isolation and RT PCR analysis
Total RNA was extracted using the RNeasy (Qiagen) method. First strand cDNA was generated using Prime Script 1st Strand cDNA kit (Takara) according to manufacturer’s instructions. Real time PCR was performed using CFX96 PCR system with SYBR Premix Ex Taq II kit (Takara). The primers used for qRT-PCR analysis are listed in Table 2. Each PCR was carried out in at least three independent replicates. The p-values were estimated after comparing among the cell types using bilateral student test. In all analyses, ‘ns’ stands for not significant, for p- value> 0.05; ‘*' for p-value<0.05; ‘**' for p-value<0.01; ‘***' for p-value<0.001.
3. Results
3.1 Distinct phenotypes and normal karyotype as in vitro cellular systems’ signature
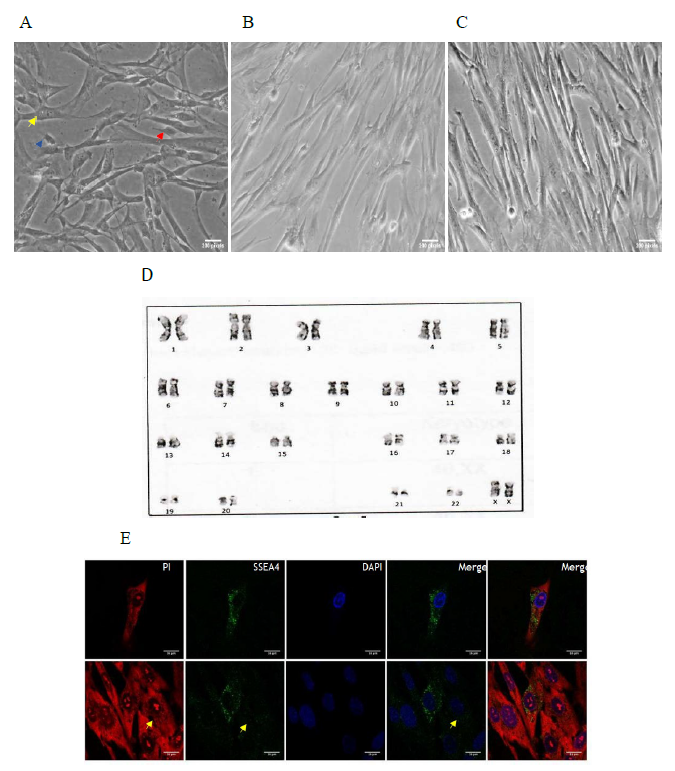
Stromal cells were derived from cord tissue explants by harvesting the outgrowth on coated plastic surface for 12 to 20 days as described above. Inverted phase contrast microscopy revealed that P3 stromal cells and configured sorted cells were documented to be spindle shaped, fibroblast-like (Figure 1A). We did not notice any major morphological differences between stromal cell type and sorted cell type although the later one was the fractionated and configured category. Sorted cells at P2 showed normal karyotype containing 46 chromosomes with no acquired numerical or structural aberrations or chromosome/chromatid breaks (Figure 1B).
Table 2: Primer sequences.
- Stromal cells with heterogeneous morphology. Yellow arrow depicts small cells; blue arrow shows fibroblast-like cells; and red arrow depicts small and round cells; B. Sorted cells in P1 seeded at 0.3 m in 35mm dish; C. Sorted cells in P5, with most of the cells seen as elongated and appear fibroblast like in morphology; D. Cytogenetic analysis of sorted cells at P3. Chromosomes maintained a normal karyotype without major alteration in morphology; E. immunofluorescence images of stromal cells. Expression of SSEA4+ (green) is shown. Cells counterstained with PI (red) to visualize morphology and DAPI (blue) to visualize nuclei. Yellow arrow mark depicts a cell with negative SSEA4 expression.
3.2 Pluripotent markers were expressed
In order to validate the ability of the sorted cells that are induced through SSEA4 fractionation and configured to the lineage differentiating potential, we assessed the expression status of pluripotent markers. We analyzed RT PCR data on three cell types: stromal cells, sorted cells and flow through cells.
The expression of OCT4, NANOG, SOX2, SSEA4 and REX was significantly enriched in sorted cells compared with that in stromal cells or with the flow through. Though, AFP and Desmin were enriched, this finding was not significant in our analysis.
3.3 Proliferation capacities determined
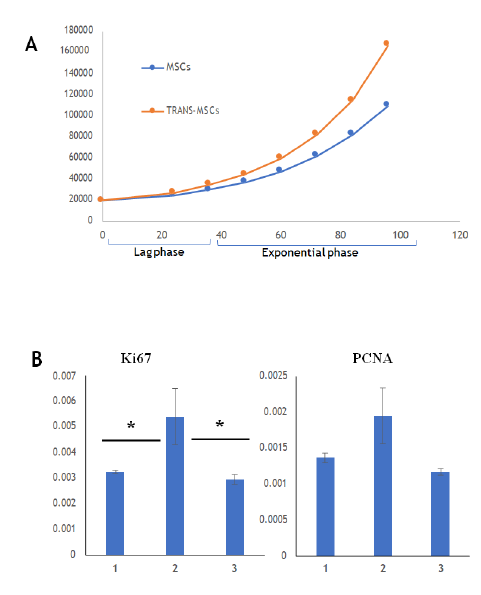
The proliferation capacity of sorted cells was assessed using two strategies. As shown in Figure 3A, Ki 67 expression was significantly increased in sorted cells compared to that in the stromal cells and flow through. Further, PCNA expression was found to be increased in sorted cells hinting at their higher proliferation capacity than stromal cells. We confirmed the same by cell growth curve assay as shown in Figure 3B that reads sorted cells undergoing strong proliferation in in vitro culture conditions created. The average doubling time, calculated in the exponential phase, for sorted cells was 45.9 ± 1.8hrs which is higher than stromal cells (54.9 ± 1.5hrs).
A. Relative gene expression of proliferation markers – Ki67 and PCNA for proliferation at P3 stage of stromal cells, sorted cells and flow through; B. Growth curves of stromal cells and sorted cells at P3 stage.
3.4 Sorted cells coaxed to neuronal lineage
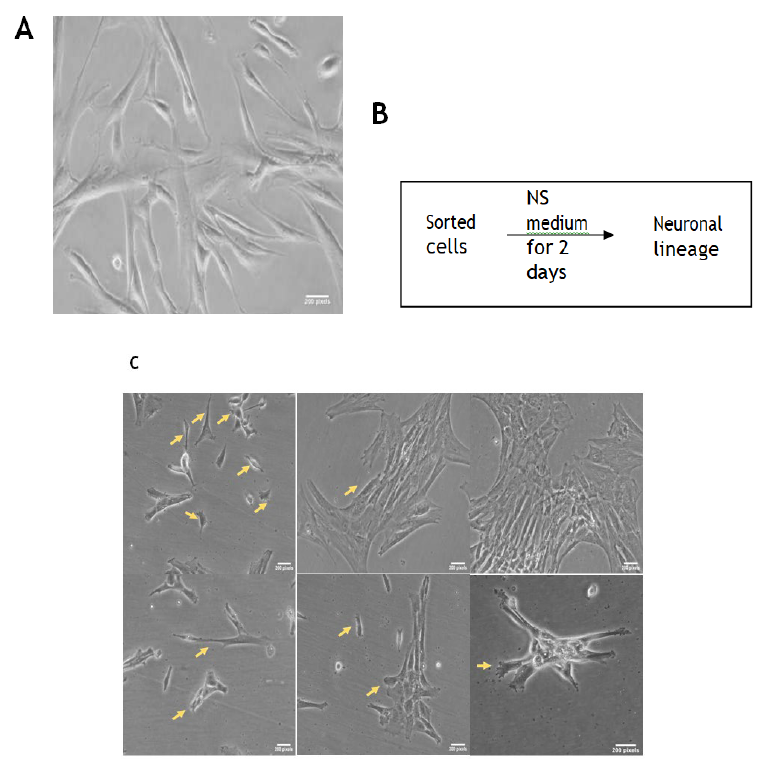
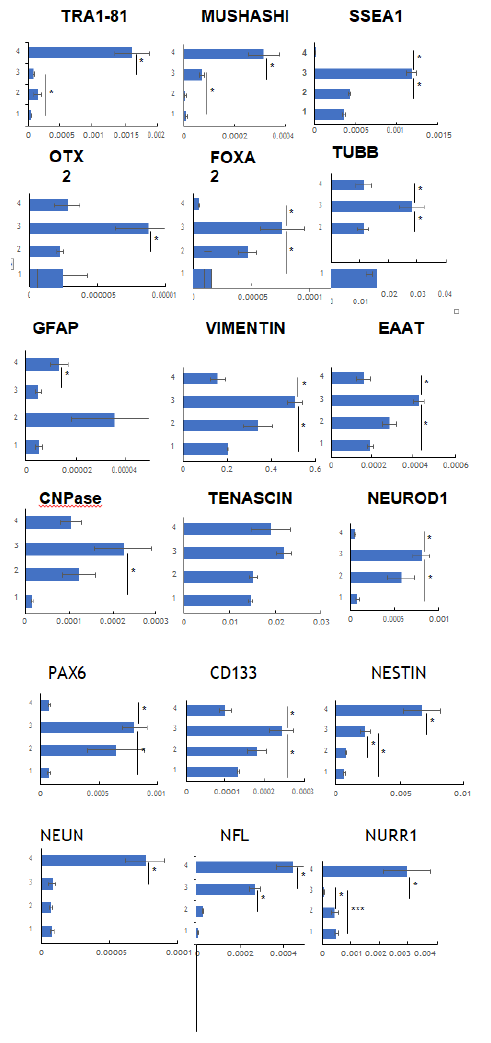
It has been noticed previously by our group as well as others in the field that it usually takes 5-7 days to differentiate iPSC into neuronal cells upon culturing in presence of various growth factors. We conditioned sorted cells that are more pluripotent than the stromal cells upon exposing to the neuronal induction medium (NS media) for 48hrs that yielded neuron-like morphologies with identities noticed as shown in Figure 4B. Initially cells remained adhered to the surface and maintained their spindle shaped morphology for the first 12hrs. Upon 24hrs of the neuronal induction medium conditioning, the cells started to show different morphologies as well as some losing the potential to adhere and form aggregates on the surface. On day 2, the cells showed various morphologies along with spindle shape as depicted in Figure 4. Few cells with dendrites had process resembling an axon, oligodendrocytes, radial glial cells as well as intermediate progenitors. Further, finer characterization involving gene expression studies was performed using quantitative real time PCR technique (Figure 5 and Table 1) [17]. GAPDH was used as a reference housekeeping gene and the expression of other genes was measured as relative expression as shown in the graphs (Figure 5). Tra1, Musashi and SSEA1, which are markers for neural stem cells as well as pluripotent stem cells, were shown to be enriched in CNC compared to the stromal cells [18]. Additionally, SSEA1, a germline marker was not enriched in SH5SY cells (Neuronal cell line derived from SKNSH cells) compared to the stromal cells or to the CNC, while Musahshi and Tra1 were found to be enriched in the cell lines. Pro-neurogenic factors like Otx2 and FoxA2 but not EN, that are critical for visceral endoderm development in turn neural plate during embryonic stages were enriched significantly in CNC compared to either stromal cells or to SH5SY cells [19, 20]. Also, CNPase, an oligodendrocyte marker was enhanced in CNCs [21]. Further, among all the tubulins, Tubulin Beta 3 Class III usually expressed primarily in neurons and responsible for cell migration was shown to be fortified in CNC [22, 23].
Among all the Radial glial cell markers, Vimentin and EAAT were significantly measured to be more in CNC compared to stromal cells or with cell line while Tenascin was unaffected [24]. NeuroD1, which is a known marker for immature neurons, was found to be enriched significantly in CNC compared to SH5SY cell line or with stromal cells [25]. Pax6 and CD133 were documented to be substantially expressed in CNC compared to the stromal cells or with SH5SY cell line which are markers for Dopaminergic neurons. Nestin expression was shown to be enhanced both in CNC and SH5SY cell line compared to stromal cells alone. Further, NEUN, Nurr1, TH, Tenascin, MAP2, GFAP markers for mature neurons were not detected to be enriched in CNC.
A. Morphology of sorted cells; B. Schematic diagram for the treatment used for inducing the differentiation towards neuronal lineage from sorted cells; C. Morphology of CNCs. Arrows depict altered cell morphologies that are close to the neuronal lineage features.
Relative expression of gene markers corresponding to various cell types in neuronal lineage. 1: stromal cells 2: Sorted cells 3: CNC and 4: SH5SY cell line. All the experiments were performed in triplicates and error bars represent s.e.m.
Table 1: Summary table comparing the gene expression status of CNCs with other cells.
4. Discussion
Use of cell lines in testing the safety or efficacy of drugs and vaccines is reported to offer numerous benefits over animal models. These immortalized commercially available cell lines derived from miscellaneous sources have been used to study mechanisms involved in discovery and development of neurological diseases, pathology [26]. Human neuronal framework is a complex connection of cell-to-cell interactions along with the external stimuli. There have been commendable advances in developing neuronal models built on mono immortalized cultures with genetic and transcriptional divergence, but the research translatability was achieved when in vitro models had heterogeneous cell types [27]. A comprehensive analysis of embryonic stem cell line and bone marrow-derived MSCs did demonstrate great similarities in their cellular genetic and proteomic profiles [28]. Our study was inspired in part by a recently conducted analysis of cultured in vitro cell based system while the aim of our study was to characterize biological discard derived primary progenitor stromal cells, sorted cells and CNC, to leverage their signature features in developing application ready in vitro human surrogate micro physiological systems [29].
Stromal cells are multipotent stem cells that can be isolated and cryopreserved at scale from various adult or fetal tissues like fat bodies, dental pulp, placenta, muscle, and umbilical cord blood and tissue [21, 30]. Cultured cells accrue genetic variations with successive passages and are also variably heterogeneous in this respect as well as in their plasticity. Cultured primary cells harvested from biological tissues will have variable phenotypes and genetic background, as these attributes are functions of source and passages. The quality of their preparation is important if they are to be reliably and reproducibly used for a multitude of in vitro applications and use cases. For this reason, the process and methods to generate homogeneous configurations and some threshold quality standards as signatures for the cell preparations are the key elements in developing in vitro micro physiological systems.
In this study, we employed the MACS process to isolate positive SSEA4 cells from the stromal cell population (with 5-20% of SSEA4 positivity), isolated from the biological discard: human umbilical cord tissue. In other words, we configured it by enriching the cellular population with the SSEA4 cells (30-72%) and further culture expanding them. Cells bearing this marker are known to have the capacity for triploblastic differentiation into three germ layers and self- renewal [29]. The identification of SSEA4 positive cells from human tissue has precedence; SSEA4-positive cells have been previously identified from several human tissues like human fetal forebrain, forebrain-derived neurosphere cells, and human adult ovaries [31]. MSCs derived from umbilical cords appear to have greater neurogenic potential than those from other sources. Limbal SSEA4 positive cells that resemble fibroblasts have been shown to be pluripotent, and to have the ability to differentiate in-vitro [32]. Multipotent MSCs from umbilical cord matrix and lining have been isolated and characterized, and the latter was shown to differentiate into all three germ layers [33, 34]. The SSEA4 protein marker is targeted to the cell surface, and is present at very low levels in the cytoplasm in DU145 cells [35], and staining analyses of the sorted cells in our system reveal the same. Not all studies on SSEA4 cells are in accord though. SSEA4’s value as a marker for proliferation and multipotency has been contested in the past [36], but here we demonstrated the role of SSEA4 influencing pluripotency in the context of neuronal lineage as part of the in vitro culture systems handled.
It has been previously reported, that iPSCs typically differentiate into neuronal cells in 5-7 days upon treatment with various growth factors [37]. We treated sorted cell configured system in a conditioning medium and induced neuron-like morphologies (Figure 4b). The transition and progression from sorted cells to neuron-like phenotype was prominent by the second day with spindle shape changing to a variety of morphologies resembling: dendrite- and axon-like cellular structures, cells resembling oligodendrocytes, radial glial cells [38].
The progression and response in gross morphology was reflected in genetic analysis/ cellular gene expression (Table 1). Stromal cells in general, and sorted stromal cells in particular, also express the Sox2 and the Pax6 protein marker, the latter is the characteristic of neuronal cells and one that enables differentiation into the neuronal lineage [39]. Previously, it has been observed that UC derived Mesenchymal Stem Cells have better neurogenic potential than other sources [40] .The Neuronal like cells derived in this study, CNCs, show characteristic enrichment of Pax6, CD133, SSEA1, FOXA2, EAAT, Vimentin, TUBB and NeuroD1 while showed downregulated expression compared to SH5SY cell line for Musashi, Tra1-81, Nestin, Nurr1, EN, GFAP, NEUN, NF (Table 1) [41]. In vitro differentiation of configured SSEA4 cells (precursors) isolated from stromal cells by our method of conditioning is therefore of value for large scale studies as the human microphysiological systems can be created in vitro in reasonable time period. The systematic sorting of large scale SSE4A cells in the labs owing to the biological discards availability and handling them to obtain barcoded neuronal phenotype with genotype can be considered as a next generation strategy in generating application specific cellular signatures for their use as models for testing like assays retaining scalability and flexibility like advantages. Importantly, these are attractive alternative human surrogate in vitro systems to animal models for the routine testing of cosmetics, vaccine, and pharmaceutical candidates. Such cellular platforms can be engineered further with ease of obtaining large batches applied to read neurotoxicity and neurovirulence like non-clinical end points with speed and throughput. Our data profiling stromal cells, sorted cells and CNCs do suggest various possibilities and way forward applications of these in vitro especially human micro physiological systems in developing non-animal, cruelty free workstation solutions integrated with data analytics in discovery, development and manufacturing stages.
TRANS-MSC is the trade name given to configured sorted cells by Transcell Oncologics that has a Science Exchange registered vertical www.transtoxbio.com with all new league of human surrogate in vitro micro physiological platforms used as tools in preclinical research and testings.
Acknowledgements
We thank Karthik B from CCMB, Hyderabad for help in karyotyping. We thank Prof Krishnaveni Mishra and Prof Bindu Madhav Reddy from School of Life Sciences, HCU, Hyderabad for assistance in confocal microscopy and SH5SY cell line. We thank Dr Lakshman Varanasi, Krea University, India for critical reading of our manuscript.
Author Contributions
SD and LG conceived the study. VD executed the in vitro experimentation, design, analysis and drafting of the manuscript. PB was involved in generation of primary cells data and experimental work. VD and SD analyzed the data, contributing to manuscript preparation.
Funding
The financial support for the research, authorship, publication of this article was from Transcell Oncologics investors and Indian Angel Network.
Conflict of Interest
The authors have no relevant financial or non-financial interests to disclose.
Ethical Approval
The study was performed in line with Ethics committee approval handling biological discards for in vitro experimentation.
Consent to Participate
Informed consent was obtained from the donors to collect their biological discards utilized in the study.
References
- Nagamura-Inoue T, He H. Umbilical cord-derived mesenchymal stem cells: Their advantages and potential clinical utility. World J Stem Cells 6 (2014):195-202.
- Hughes JP, Rees S, Kalindjian SB, Philpott KL. Principles of early drug discovery. Br J Pharmacol 162 (2011): 1239-1249.
- Edwards MA, Loxley RA, Williams AJ, Connor M, Phillips JK. Lack of functional expression of NMDA receptors in PC12 cells. Neurotoxicology 28 (2007): 876-885.
- LePage KT, Dickey RW, Gerwick WH, Jester EL, Murray TF. On the use of neuro-2a neuroblastoma cells versus intact neurons in primary culture for neurotoxicity studies. Crit Rev Neurobiol 17 (2005): 27-50.
- Gamo NJ, Sawa A. Human stem cells and surrogate tissues for basic and translational study of mental disorders. Biol Psychiatry 75 (2014): 918-919.
- Handa K, Matsubara K, Fukumitsu K, Guzman-Lepe J, Watson A, et al. Assembly of human organs from stem cells to study liver disease. Am J Pathol 184 (2014): 348-357.
- Mahla RS. Stem Cells Applications in Regenerative Medicine and Disease Therapeutics. Int J Cell Biol 2016 (2016): 6940283.
- Azari H, Reynolds BA. In Vitro Models for Neurogenesis. Cold Spring Harb Perspect Biol 8 (2016).
- Gordon J, Amini S, White MK. General overview of neuronal cell culture. Methods Mol Biol 1078 (2013): 1-8.
- Weiss ML, Troyer DL. Stem cells in the umbilical cord. Stem Cell Rev 2 (2006): 155-162.
- Pastor DM, Poritz LS, Olson TL, Kline CL, Harris LR, et al. Primary cell lines: false representation or model system? a comparison of four human colorectal tumors and their coordinately established cell lines. Int J Clin Exp Med 3 (2010): 69-83.
- Richter M, Piwocka O, Musielak M, Piotrowski I, Suchorska WM, et al. From Donor to the Lab: A Fascinating Journey of Primary Cell Lines. Front Cell Dev Biol 9 (2021): 711381.
- May Fulton C, Bailey WJ. Live Viral Vaccine Neurovirulence Screening: Current and Future Models. Vaccines (Basel) 9 (2021).
- Schultz L, Zurich MG, Culot M, da Costa A, Landry C, et al. Evaluation of drug- induced neurotoxicity based on metabolomics, proteomics and electrical activity measurements in complementary CNS in vitro models. Toxicol In Vitro 30 (2015):138-165.
- Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, et al. Fiji: an open-source platform for biological-image analysis. Nat Methods 9 (2012): 676-682.
- Campos PB, Sartore RC, Abdalla SN, Rehen SK. Chromosomal spread preparation of human embryonic stem cells for karyotyping. J Vis Exp (2009).
- Yang N, Ng YH, Pang ZP, Sudhof TC, Wernig M. Induced neuronal cells: how to make and define a neuron. Cell Stem Cell 9 (2011): 517-525.
- Zhao W, Ji X, Zhang F, Li L, Ma L. Embryonic stem cell markers. Molecules 17 (2012): 6196-6236.
- Costello I, Nowotschin S, Sun X, Mould AW, Hadjantonakis AK, et al. Lhx1 functions together with Otx2, Foxa2, and Ldb1 to govern anterior mesendoderm, node, and midline development. Genes Dev 29 (2015): 2108-2122.
- Bielinska M, Narita N, Wilson DB. Distinct roles for visceral endoderm during embryonic mouse development. Int J Dev Biol 43 (1999):183-205.
- Michalski D, Keck AL, Grosche J, Martens H, Hartig W. Immunosignals of Oligodendrocyte Markers and Myelin-Associated Proteins Are Critically Affected after Experimental Stroke in Wild-Type and Alzheimer Modeling Mice of Different Ages. Front Cell Neurosci 12 (2018): 23.
- Sferra A, Petrini S, Bellacchio E, Nicita F, Scibelli F, et al. TUBB Variants Underlying Different Phenotypes Result in Altered Vesicle Trafficking and Microtubule Dynamics. Int J Mol Sci 21 (2020).
- Breuss MW, Leca I, Gstrein T, Hansen AH, Keays DA. Tubulins and brain development - The origins of functional specification. Mol Cell Neurosci 84 (2017): 58-67.
- Kriegstein A, Alvarez-Buylla A. The glial nature of embryonic and adult neural stem cells. Annu Rev Neurosci 32 (2009):149-184.
- Boutin C, Hardt O, de Chevigny A, Core N, Goebbels S, et al. NeuroD1 induces terminal neuronal differentiation in olfactory neurogenesis. Proc Natl Acad Sci USA 107 (2010): 1201-1206.
- Slanzi A, Iannoto G, Rossi B, Zenaro E, Constantin G. In vitro Models of Neurodegenerative Diseases. Front Cell Dev Biol 8 (2020): 328.
- Altschuler SJ, Wu LF. Cellular heterogeneity: do differences make a difference? Cell 141 (2010): 559-563.
- Balasubramanian S, Thej C, Venugopal P, Priya N, Zakaria Z, et al. Higher propensity of Wharton's jelly derived mesenchymal stromal cells towards neuronal lineage in comparison to those derived from adipose and bone marrow. Cell Biol Int 37 (2013): 507-515.
- Guenther R, Dreschers S, Maassen J, Reibert D, Skazik-Voogt C, et al. The Treasury of Wharton's Jelly. Stem Cell Rev Rep (2021).
- Abdulrazzak H, Moschidou D, Jones G, Guillot PV. Biological characteristics of stem cells from foetal, cord blood and extraembryonic tissues. J R Soc Interface 7 (2010): S689-S706.
- Vasicek J, Balazi A, Bauer M, Chrenek P. SSEA-4 Antigen Is Expressed on Rabbit Lymphocyte Subsets. Magnetochemistry 7 (2021).
- Dravida S, Pal R, Khanna A, Tipnis SP, Ravindran G, et al. The transdifferentiation potential of limbal fibroblast-like cells. Brain Res Dev Brain Res 160 (2005): 239-251.
- Lee KS, Kang HW, Lee HT, Kim HJ, Kim CL, et al. Sequential sub-passage decreases the differentiation potential of canine adipose-derived mesenchymal stem cells. Res Vet Sci 96 (2014): 267-275.
- Gonzalez R, Griparic L, Umana M, Burgee K, Vargas V, R et al. An efficient approach to isolation and characterization of pre- and postnatal umbilical cord lining stem cells for clinical applications. Cell Transplant 19 (2010):1439-1449.
- Sivasubramaniyan K, Harichandan A, Schilbach K, Mack AF, Bedke J, et al. Expression of stage-specific embryonic antigen-4 (SSEA-4) defines spontaneous loss of epithelial phenotype in human solid tumor cells. Glycobiology 25 (2015): 902-917.
- Petrenko Y, Vackova I, Kekulova K, Chudickova M, Koci Z, et al. A Comparative Analysis of Multipotent Mesenchymal Stromal Cells derived from Different Sources, with a Focus on Neuroregenerative Potential. Sci Rep 10 (2020): 4290.
- Hu BY, Weick JP, Yu J, Ma LX, Zhang XQ, et al. Neural differentiation of human induced pluripotent stem cells follows developmental principles but with variable potency. Proc Natl Acad Sci USA 107 (2010): 4335-4340.
- Leyh J, Paeschke S, Mages B, Michalski D, Nowicki M, et al. Classification of Microglial Morphological Phenotypes Using Machine Learning. Front Cell Neurosci 15 (2021): 701673.
- Gomez-Lopez S, Wiskow O, Favaro R, Nicolis SK, Price DJ, et al. Sox2 and Pax6 maintain the proliferative and developmental potential of gliogenic neural stem cells In vitro. Glia 59 (2011):1588-1599.
- Urrutia DN, Caviedes P, Mardones R, Minguell JPJ, Vega-Letter AM, et al. Comparative study of the neural differentiation capacity of mesenchymal stromal cells from different tissue sources: An approach for their use in neural regeneration therapies. Plos One 14 (2019).
- Ashraf R, Sofi HS, Sheikh FA. Experimental Protocol of MSC Differentiation into Neural Lineage for Nerve Tissue Regeneration Using Polymeric Scaffolds. Methods Mol Biol. 2125 (2020):109-117.


 Impact Factor: * 5.8
Impact Factor: * 5.8 Acceptance Rate: 71.20%
Acceptance Rate: 71.20%  Time to first decision: 10.4 days
Time to first decision: 10.4 days  Time from article received to acceptance: 2-3 weeks
Time from article received to acceptance: 2-3 weeks 