Information Fusion for Colorectal Polyps Medical Image Segmentation
Zhuo Zhou1, Yinghua Duan2, Weilan Huang1, Fengyun Pei3, Bo Liu4, Jun Huang3,5,6*
1Health Management Center, Third Affiliated Hospital of Sun Yat-sen University, Guangzhou, 510620, Guangdong, China
2Department of Traditional Chinese Medicine, First Affiliated Hospital of Sun Yat-sen University, Guangzhou, 510060, Guangdong, China
3Department of Colorectal Surgery, Sixth Affiliated Hospital of Sun Yat-sen University, Guangzhou, 510655, Guangdong, China
4Academy of Mathematics and Systems Science, Chinese Academy of Sciences, Beijing, China
5Guangdong Provincial Key Laboratory of Colorectal and Pelvic Floor Diseases, Sixth Affiliated Hospital of Sun Yat-sen University, Guangzhou, 510655, Guangdong, China
6Guangdong Institute of Gastroenterology, Guangzhou, 510655, Guangdong, China
*Corresponding author: Zhuo Zhou, Health Management Center, Third Affiliated Hospital of Sun Yat-sen University, Guangzhou, 510620, Guangdong, China.
Jun Huang, Department of Colorectal Surgery, Sixth Affiliated Hospital of Sun Yat-sen University, Guangzhou, 510655, Guangdong, China.
Received: 27 July 2022; Accepted: 04 August 2022; Published: 19 August 2022
Article Information
Citation: Zhuo Zhou, Yinghua Duan, Weilan Huang, Fengyun Pei, Bo Liu, Jun Huang. Information Fusion for Colorectal Polyps Medical Image Segmentation. Archives of Clinical and Biomedical Research 6 (2022): 744-748.
View / Download Pdf Share at FacebookAbstract
Training a deep neural network often requires a large amount of annotated data, which is scarce in the medical image analysis domain. In this work, we present a simple yet effective technique for enhancing medical image segmentation neural network through information fusion. The proposed approach utilizes information from different spatial scales and combines them in a learnable way. Experimental results on two benchmark datasets demonstrate that the proposed fusion module improves the segmentation performance of state-of-the-art neural networks.
Keywords
<p>Information Fusion; Medical Image Analysis; Polyps Segmentation; Neural Network; Semantic Segmentation</p>
Article Details
1. Background
Semantic segmentation is a common computer vision task in which each pixel in an image is assigned to a pre-defined category [1]. It has been widely used in applications such as landscape mapping, autonomous driving, and medical image analysis [2,3]. In the last decade, training a deep convolutional neural network (CNN) to segment the image has replaced the handcrafted features to become a common practice due to its superior performance. It is well understood that deep neural networks are notoriously data hungry insofar as effectively training the model is concerned. As op- posed to the application of semantic segmentation in the industry, deploying a segmentation system for medical image analysis has been found challenging. Lack of annotated data has always been a major obstacle in the medical image analysis community [4]. This is attribute to the manner on how medical images are labeled. Unlike natural images that can be easily obtained, medical images are hard to acquire due to the data privacy concern. To make matters worse the annotator is required to have domain expertise for labeling medical images. Colorectal cancer is the third most commonly diagnosed cancer and the fourth cause of cancer death around the world [5]. As a precursor to the colorectal cancer, polyps are abnormal tissue growths on the inner lining of the colon or rectum. In clinical practice, colonoscopy is the preferred technique for detection and assessment of polyps. Therefore, developing a method to accurately segment polyps in colonoscopy images is of great interest as it helps the doctor to identify polyps at an early stage. However, segmenting polyps is a challenging task as the same type of polyps appears in a high variety of color, size and texture. Despite the advancement in colonoscopy, the boundary between polyps and surrounding mucous membrane is usually not very sharp as shown in the Figure 1. To address these challenges, in this paper, we propose a novel information fusion module to improve the performance of existing segmentation networks. The key idea is to combine features from different layers in a data adaptive way. We evaluate the proposed method on two publicly available datasets, Kvasir-SEG [6] and CVC-612 [7], for polyp segmentation in colonoscopy images. The results demonstrate that the proposed fusion module improves on multiple network architectures for medical image segmentation. The remainder of this paper is organized as follows. We introduce the related work in Section 2 and describe the proposed approach in Section 3. Section 4 describes the two datasets along with the segmentation results. Concluding remarks are provided in Section 5.
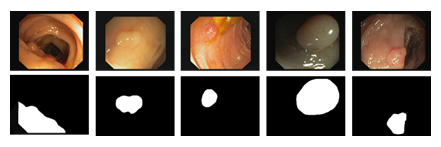
Figure 1: Example colonoscopy images of polyps (top row) and associated labels (bottom row) from Kvasir-SEG (6) dataset.
2. Related work
To address the label scarcity problem, plenty of machine learning techniques have proposed in the last few years. Existing techniques can be roughly grouped into two broad categories: efficient training and data feature enhancement. Active learning seeks to train the network with as little labeled data as possible by picking the most informative samples to increase network training efficiency [8]. By pre-training the network on a large-scale dataset like ImageNet [9], where labels are relatively easy to collect, transfer learning allows a model to be fine-tuned with a considerably less quantity of data. Semi-supervised learning differs from supervised learning in that it extracts informative characteristics from unlabeled data [10]. Data feature enhancement, on the other hand, focuses on creating more discriminative features and exploiting the relationship between data samples. Synthetic data generated with generative adversarial network (GAN) [11] has proven to be very effective for improving the model performance [12,13]. Statistical shape and appearance models have also been used to generate training samples with higher data variance that helps improve the robust- ness of the model [14]. The hierarchical features in CNN provide a natural way to address the object scale variance issue in medical image segmentation [15]. U-Net [16] and its variants [17-20] capture the multi-scale information by connecting features from shallow and deep layers. Attention- based methods [21] guide the training to focus on important image regions such as object edges such that the segmentation accuracy can be improved.
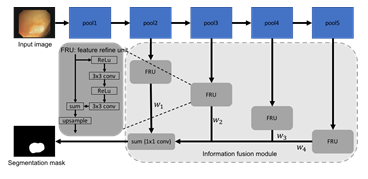
Figure 2: Network architecture of a standard FCN with the proposed information fusion module.
3. Methods
In this section, we introduce the proposed information fusion module. We begin by describing the base network, fully convolutional network (FCN) (1), and the feature refinement unit [22]. We then explain how the multi-scale features are combined through the information fusion module. Figure 2 de- picts our network architecture. Fully convolutional network (FCN) has been extensively used for semantic segmentation since it was introduced in 2015 [1]. To simplify the flowchart, we omit the convolutional layers and the output (softmax) layer in FCN in Figure 2. The convolutional features after four pooling layers, which are commonly known as FCN-32s, FCN-16s, FCN-8s, and FCN-4s, are sent through the information fusion module before making the prediction. The information fusion module not only improves the discrimination ability of features, but also aggregates features at different level in a learnable fashion. In particular, the convolution features at each stage are first connected to a feature refinement unit (FRU). The detailed architecture of a FRU is shown in Figure 2. Inspired by the RefineNet [22], we add two additional convolutional layers and associated ReLU activation layers [23] in combine with a residual connection. This refinement process has shown to be effective in dealing with fine-scale features [22]. When used in colonoscopy image segmentation, the feature refinement is helpful for segmenting polyps that are relatively small as compared to the image size. It is well known that early layers in a CNN are responsible for extracting shallow, fine, and appearance information from the data, while layers in later stages tend to produce features at a coarse scale and are related to the semantic information [24,25]. The success of U-Net [16] has shown that combining these two types of features is critical for producing accurate segmentation map for medical images. Instead of directly concatenating all features, the outputs of FRU are added together and followed by a 1 × 1 convolution. This essentially learns a weight for features from each layer. Compared to a feature concatenation or a feature summation, weights for each component can be learned through the error back propagation. Therefore, features that contain different information can be combined in a data-driven way. It is worth noting that we choose FCN as the base network for illustration purpose. The proposed information fusion module is flexible and generic. We can attach different number of FRUs based on the feature groups. For example, when the proposed information fusion module is used with U-Net, we can attach one FRU after each skip connection, which results in a 4-stage fusion. The information fusion module can also be used with other network architectures as long as the convolutional features can be extracted from different depths. The standard pixel-wise cross entropy loss is used to train the network.
4. Experiments
In this section, we evaluate the proposed method by applying them to two publicly available polyp segmentation datasets. We compare the performance of multiple networks with and without adding the information fusion module.
4.1 Datasets and Baselines
Kvasir-SEG (6) is a recently released dataset for polyp segmentation. It consists of 1000 images and their corresponding ground truth. These images were manually labeled by a medical doctor and reviewed by an experienced gastroenterologist from Vestre Viken Health Trust in Norway. CVC-612 (7) is a benchmark dataset that has been used in many polyps segmentation studies. It contains 612 images that were extracted from 31 colonoscopy videos. For both datasets, each pixel in the image is labeled as either polyp or non-polyp. To evaluate the efficacy of the proposed method, we compare the performance before and after adding the information fusion module with multiple networks, including FCN [1], U-Net [16], U-Net++ [17], ResUNet++ [19], PraNet [26]. Since the PraNet has a similar multi-level feature aggregation design, we compare it with the proposed method by replacing the parallel reverse attention module in PraNet with the proposed information fusion module.
4.2 Results
For both datasets, we follow the common experimental setup used in [19,26] by randomly split the data into three sets, training (80%), validation (10%), and testing (10%). The performance of network is measured with standard semantic segmentation metrics, Dice and mean Intersection over Union (mIoU). All networks are implemented using PyTroch framework [27]. The experiments are conducted on an AWS EC2 p3.2xlarge instance. For baseline networks, i.e., without information fusion module, we use numbers reported by the original authors. To have a fair comparison, we use the same hyperparameters as the authors specified in the original papers when training the network with information fusion module. For networks that the training details are not provided, we train the network 100 epochs with a batch size 16. The initial learning rate is set to 0.01. For optimization, we use the SGD optimizer with cosine decay for learning rate schedule [28]. We also use the standard data augmentation, such as random flip, crop, rotation, and color jittering, etc., to improve the model performance.
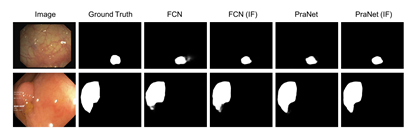
Figure 3: Segmentation results of different methods on CVC-612 (top row) and Kvasir- SEG (bottom row) datatsets.
|
Method |
Kvasir-SEG |
CVC-612 |
|||
|
Dice |
mIoU |
Dice |
mIoU |
||
|
FCN [1] |
0.794 |
0.763 |
0.768 |
0.728 |
|
|
FCN (IF) |
0.822 |
0.792 |
0.791 |
0.762 |
|
|
U-Net [16] |
0.818 |
0.746 |
0.823 |
0.755 |
|
|
U-Net (IF) |
0.833 |
0.77 |
0.845 |
0.773 |
|
|
U-Net++ [17] |
0.821 |
0.743 |
0.794 |
0.729 |
|
|
U-Net++ (IF) |
0.831 |
0.767 |
0.819 |
0.742 |
|
|
ResUNet++ [19] |
0.813 |
0.793 |
0.796 |
0.796 |
|
|
ResUNet++(IF) |
0.827 |
0.812 |
0.815 |
0.816 |
|
|
PraNet [29] |
0.898 |
0.84 |
0.899 |
0.849 |
|
|
PraNet (IF) |
0.899 |
0.849 |
0.903 |
0.857 |
|
Table 2: Segmentation results of networks with (indicated by (IF)) and without the information fusion module on Kvasir-SEG and CVC-612 dataset.
Table 1 shows the segmentation results of all networks. As can be seen that the information fusion module (indicated by (IF) in the table) consistently improved the model performance on both datasets. When applied to a basic network like FCN [1], the information fusion module improved Dice from 0.794 to 0.822 by 3.5%, and mIoU from 0.763 to 0.792 by 3.8% on the Kvasir-SEG dataset. A similar trend can be observed on the CVC-612 dataset as well as for U-Net. With advanced variants of U-Net, the additional connections in U-Net++ [17] and ResUNet++ [19] are expected to enhance the feature combination from different layers. The model with in- formation fusion module still achieved higher Dice and mIoU. We attribute this to the weighted summation with learned weights compared to simple feature concatenation and feature summation. The improvement on the recently published PraNet [26] is relatively small as opposed to other networks. This is because the PraNet also adopts multi-scale feature fusion using an attention module, which shares the same motivation as the proposed information fusion module. It is worth mentioning that the performance achieved by the PraNet is the state-of-the-art on both datasets. Considering that the PraNet results are close to the empirical upper bound of these two datasets, the improvement achieved by the information fusion module is considerably significant. Fig. 3 shows the qualitative results for models with and without the in- formation fusion module on CVC-612 and Kvasir-SEG datasets. We observe similar trends from all the compared networks and choose FCN and PraNet as examples. As can be seen that networks with the proposed information fusion module produced segmentation masks with sharper edges compared to the ones produced by networks without the proposed module. The ba- sic network (FCN and PraNet) tended to make mistake when the boundary between polyps and surrounding mucous membrane is blurry. With the in- formation fusion module, the networks (FCN (IF) and PraNet (IF)) were able to separate these polyps from background. We attribute this improvement to the ability to combine fine-scale features with coarse-scale features.
5. Conclusion
In this paper, we have proposed a novel approach to improve semantic segmentation networks for medical image segmentation. The proposed information fusion can be easily incorporated as a module for all common segmentation networks. The feature refinement and weighted feature summation that provided by the information fusion module have been demonstrated to be effective in improving the segmentation performance on real polyp segmentation datasets when compared with other basic networks.
Acknowledgments
This work was supported by the National Natural Science Foundation of China [Grant No. 81972885], the 1010 project of the 6th Affiliated Hospital of Sun Yat-sen University [1010CG (2020)-20].
References
- Long J, Shelhamer E, Darrell T. Fully convolutional networks for se- mantic segmentation, in: Proceedings of the IEEE conference on com- puter vision and pattern recognition (2015): 3431-3440.
- Krizhevsky A, Sutskever I, Hinton GE. Imagenet classification with deep convolutional neural networks, in: Advances in neural information processing systems (2012): 1097-1105.
- Girshick R, Donahue J, Darrell T, et al. Rich feature hierarchies for accurate object detection and semantic segmentation, in: Proceedings of the IEEE conference on computer vision and pattern recognition (2014): 580-587.
- Tajbakhsh N, Shin JY, Gurudu SR, et al. Convolutional neural networks for medical im- age analysis: Full training or fine tuning?, IEEE transactions on medical imaging 35 (2016): 1299-1312.
- Favoriti P, Carbone G, Greco M, et al. Cor- cione, Worldwide burden of colorectal cancer: a review, Updates in surgery 68 (2016): 7-11.
- Jha D, Smedsrud PH, Riegler MA, et al. A segmented polyp dataset, in: International Conference on Multimedia Modeling, Springer (2020): 451-462.
- Bernal J, S´anchez FJ, Fern´andez-Esparrach G, et al. Vilarin˜o, Wm-dova maps for accurate polyp highlighting in colonoscopy: Validation vs. saliency maps from physicians, Comput- erized Medical Imaging and Graphics 43 (2015): 99-111.
- Yang L, Zhang Y, Chen J, et al. Suggestive annota- ti.on: A deep active learning framework for biomedical image segmen- tation, in: International conference on medical image computing and computer-assisted intervention, Springer (2017): 399-407.
- Deng J, Dong W, Socher R, et al. Imagenet: A large-scale hierarchical image database, in: 2009 IEEE conference on computer vision and pattern recognition, IEEE (2009): 248-255.
- Bai W, Oktay O, Sinclair M, et al. Semi-supervised learning for network-based cardiac mr image segmentation, in: Interna- tional Conference on Medical Image Computing and Computer-Assisted Intervention, Springer (2017): 253-260.
- Goodfellow I, Pouget-Abadie J, et al. Generative adversarial nets, in: Ad- vances in neural information processing systems (2014): 2672-2680.
- Frid-Adar M, Diamant I, Klang E, et al. Greenspan, Gan-based synthetic medical image augmentation for in- creased cnn performance in liver lesion classification, Neurocomputing 321 (2018): 321-331.
- Frid-Adar M, Klang E, Amitai M, et al. Syn- thetic data augmentation using gan for improved liver lesion classifica- tion, in: 2018 IEEE 15th international symposium on biomedical imag- ing (ISBI 2018), IEEE (2018): 289-293.
- Uzunova H, Wilms M, Handels H, et al. Training cnns for image registration from few samples with model-based data augmentation, in: International Conference on Medical Image Computing and Computer- Assisted Intervention, Springer (2017): 223-231.
- Shi C, Zhang J, Zhang X, et al. A recurrent skip deep learning network for accurate image segmentation, Biomedical Signal Processing and Control 74 (2022): 103533.
- Ronneberger O, Fischer P, Brox T. U-net: Convolutional networks for biomedical image segmentation, in: International Conference on Medical image computing and computer-assisted intervention, Springer (2015): 234-241.
- Zhou Z, Siddiquee MMR, Tajbakhsh N, et al. Unet++: A nested u-net architecture for medical image segmentation, in: Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support, Springer (2018): 3-11.
- Alom MZ, Yakopcic C, Hasan M, et al. Recur- rent residual u-net for medical image segmentation, Journal of Medical Imaging 6 (2019): 014006.
- Jha D, Smedsrud PH, Riegler MA, et al. Resunet++: An advanced architecture for medical image segmentation, in: 2019 IEEE International Symposium on Multimedia (ISM), IEEE (2019): 2225-2255.
- Jha D, Riegler MA, Johansen D, et al. Doubleu-net: A deep convolutional neural network for medical image segmentation (2020).
- Yu J, Yang D, Zhao H. Ffanet: Feature fusion attention network to medical image segmentation, Biomedical Signal Processing and Control 69 (2021): 102912.
- Lin G, Milan A, Shen C. Refinenet: Multi-path refinement networks for high-resolution semantic segmentation, in: Proceedings of the IEEE conference on computer vision and pattern recognition (2017): 1925-1934.
- Nair V, Hinton GE. Rectified linear units improve restricted boltz- mann machines, in: ICML (2010).
- Yosinski J, Clune J, Bengio Y, et al. How transferable are fea- tures in deep neural networks?, in: Advances in neural information pro- cessing systems (2014): 3320–3328.
- Huh M, Agrawal P, Efros AA. What makes imagenet good for trans- fer learning?, arXiv preprint arXiv:1608.08614 (2016).
- Fan DP, Ji GP, Zhou T, et al. Pranet: Parallel reverse attention network for polyp segmentation, in: Interna- tional Conference on Medical Image Computing and Computer-Assisted Intervention, Springer (2020): 263-273.
- Paszke A, Gross S, Massa F, et al. An imperative style, high- performance deep learning library, in: Wallach, H. Larochelle, A. Beygelzimer, F. d'Alch´e-Buc, E. Fox, R. Garnett (Eds.), Advances in Neural Information Processing Systems 32, Curran Associates, Inc (2019): 8024-8035.
- Loshchilov I, Hutter F. Sgdr: Stochastic gradient descent with warm restarts, arXiv preprint arXiv: 1608.03983 (2016).


 Impact Factor: * 5.8
Impact Factor: * 5.8 Acceptance Rate: 71.20%
Acceptance Rate: 71.20%  Time to first decision: 10.4 days
Time to first decision: 10.4 days  Time from article received to acceptance: 2-3 weeks
Time from article received to acceptance: 2-3 weeks 