The Human Microbiome: An Emerging Key Player in Health and Disease
Hubert E. Blum
Department of Medicine II, University Hospital Freiburg, Freiburg, Germany
*Corresponding Author: Hubert E. Blum, Department of Medicine II, University Hospital Freiburg, Freiburg, Hugstetter Strasse 55, Germany, Tel: 0049-761-27018116;
Received: 22 March 2017; Accepted: 06 April 2017; Published: 11 April 2017
Article Information
View / Download Pdf Share at FacebookAbstract
Until recently, human microbiology was based on the identification of single microbes, such as bacteria, fungi and viruses, frequently isolated from patients with acute or chronic infections. Novel culture-independent molecular biochemical analyses (genomics, transcriptomics, proteomics, metabolomics) allow today to detect and classify the diverse microorganisms in a given ecosystem (microbiota), such as the gastrointestinal tract, the skin, the airway system, the urogenital tract and others, and to assess all genomes in these ecosystems (microbiome) as well as their gene products. These analyses revealed that each individual has its own microbiota that plays a role in health and disease. In addition, they greatly contributed to the recent advances in the understanding of the pathogenesis of a wide range of human diseases. It is to be expected that these new insights will translate into diagnostic, therapeutic and preventive measures in the context of personalized/ precision medicine.
Keywords
Microbiota; Fecal Transplantation; Inflammatory Bowel Disease; Obesity; Atherosclerosis; Neurodegenerative Diseases
Article Details
1. Introduction
The basic aspects of molecular and cell biology are not only an integral part of biomedical research, but are also translated into patient care. Several global consortia have been launched and in part completed during the last decades. All of them continuously transform basic biomedical research and translate into medical applications and, after evaluation in randomized clinical trials, enter clinical practice with a tremendous potential to advance the diagnosis, treatment and prevention of human diseases.
More than 15 years ago, the international human genome organization (HUGO) project established the complete sequence of the ca. 3 billion base pairs that make up the human genome. In order to utilize these data from the HUGO project for research as well as for clinical applications and to define the functions of newly identified genes, collectively termed ‘functional genomics’, strategies were developed to globally analyze genomic DNA sequences as well as their cell-, tissue- or organ-specific expression profile. Using chips, so-called 'microarrays', thousands or ten thousands of single-stranded DNA species, reverse transcribed RNA (cDNA) or oligonucleotides of known sequence can provide a global gene (genomics), gene expression (transcriptomics, proteomics) or metabolite (metabolomics) profile (‘signature’) that is characteristic for the disease of individual patients, including its natural course/ prognosis and response to therapy.
In 2005 the international haplotype map (HapMap) project was initiated to identify, based on genome-wide association studies (GWAS) in ethnically different populations, single nucleotide polymorphisms (SNPs) and their association with specific human diseases and individual phenotypic characteristics, respectively. Through GWAS an increasing number of gene loci have been identified that are associated with individual (future) phenotypic traits, such as hair or eye color, height, body mass index and others as well as with the disposition to develop a specific disease [1]. Further, genetic variants are associated with the individuals’ response to drug treatment, e.g., to lithium [2]. Overall, GWAS allow an increasingly better understanding of disease pathogenesis and a more accurate assessment of the individual risk to develop a specific disease. Clinically, this may eventually translate into clinical advances in disease prevention, early diagnosis and therapy. It should be cautioned, however, that the contribution of a defined SNP to the risk assessment for a given disease must be weighed against established clinical parameters and needs to be carefully evaluated before entering clinical practice.
2. Review
2.1. The Human Microbiome Project
The human microbiome project (HMP) was established in 2007 as another global consortium [3]. The HMP and the ‘Metagenomics of the Human Intestinal Tract (Meta-HiT) Consortium Europe’ aim at the sequencing of all microbes (eukaryotes, archaea, bacteria, viruses) that inhabit specific body sites, such as the mouth, throat and airways, stomach and intestine, the urogenital system and the skin, respectively (Figure 1A). Recent data demonstrate that specific compositions of the microbial community are associated with health and disease (Figure 1B) and suggest that the detailed characterization, function and variation of the microbial community will reveal an important commensal host-microbe as well as microbe-microbe interactions with diagnostic, therapeutic and preventive implications [4,5].
While the HMP has meanwhile developed into a major field of biomedical research, the intestinal microbial community in particular has turned out to play a major role in human health and disease pathogenesis as will be discussed in more detail below [6].
2.2. The Intestinal Microbial Community
In recent years the intestinal microbial community has been studied in great detail. It represents a microbial ecosystem consisting of trillion microbial cells with an aggregate 9.9 million microbial genes across the fecal microbiome [7].
While until recently, the environment in utero has been considered sterile, DNA-based analyses identified bacterial species in maternal placenta, amniotic fluid and meconium. The colonization of the human gut begins at birth with a rapid expansion of bacterial diversity and is characterized by a successively changing composition that eventually becomes relatively stable in adulthood. While the specific microbial species and subspecies and their proportions vary greatly from person to person the individual microbiome is unique and becomes more diverse in the elderly.
Important factors for the composition of the intestinal microbial community are endogenous and exogenous factors. Examples are mode of delivery of the neonates, diet (dietary supplements, breast-feeding, formula-feeding), xenobiotics, including antibiotics and other drugs. Further, infections and exposure to environmental microbial agents are established risk factors for childhood diseases, such as obesity and allergy [8]. Recent evidence further suggests that human genetic variation also influences the abundance of specific members of the intestinal microbial community.
Taken together, the emerging data suggest that the detailed characterization of the human intestinal microbiome composition, function and variation across different body sites will reveal an important commensal host-microbe as well as microbe, microbe interactions that may play a role in human health and disease (Figure 1A and B).
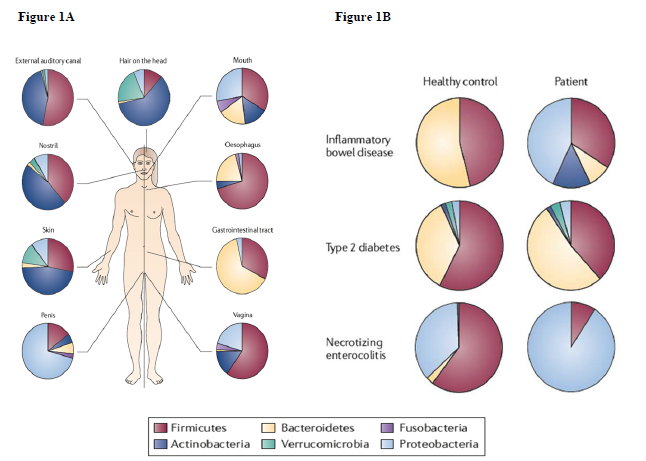
Figure 1: (A) Different microbiomes in humans; (B) The intestinal microbiome in healthy individuals and patients [27].
In view of the numerous and diverse physiological functions of the intestinal microbiota in human health (Table 1) it is not surprising that it is also involved in gastrointestinal as well as non-gastrointestinal diseases, such as obesity/ metabolic syndrome, and atherosclerosis/ cardiovascular as well as neurologic, psychiatric and neurodegenerative diseases, making it one of the most dynamic current topics in biomedical research (Table 2). In the following, a few examples will be discussed in more detail.
| Host Physiology | References |
|---|---|
| Adaptive immunity | 20 |
| Autoimmunity | 21 |
| Innate immunity | 22 |
| Cell proliferation | 23 |
| Bone density | 24 |
| Vascularization | 25 |
| Neurological signalling | 26 |
| Biosynthesis | |
| Neurotransmitters | |
| Steroid hormones | |
| Vitamins | |
| Metabolism | |
| Dietary components | |
| Bile salts | |
| Drugs | |
| Xenobiotics |
Table 1: Functions of the intestinal microbial community in human health (examples).
| Allergies/ Allergy protection |
|---|
| Atherosclerosis/ Thrombosis/ Cardiovascular Disease |
| Cancer |
| Diabetes mellitus |
| Immune-Mediated Inflammatory Diseases |
| Inflammatory bowel diseases Multiple sclerosis Rheumatoid arthritis Psoriasis |
| Kwashiorkor |
| Liver Diseases |
| Metabolic Syndrome/ Obesity |
| Neurodevelopmental, Psychiatric and Neurodegenerative Diseases |
| Autism Depression Alzheimer’s Disease, Parkinson’s Disease |
Table 2: Disease associations with the intestinal microbial community (examples)
2.2.1. Inflammatory Bowel Diseases and Colon Cancer
The inflammatory bowel diseases (IBD) in humans include ulcerative colitis (UC) and Crohn disease (CD). These are characterized by inflammation limited to the mucosal layer of the colon in UC and the transmural involvement of the gastrointestinal tract, including extraintestinal sites in CD. While the pathogenesis of IBD is not fully understood, it is clear that its pathology depends among others on the intestinal microbial community. Further, a case-control study identified ‘IBD-specific’ alterations of the intestinal microbiota that may serve as biomarkers for the prediction of disease predisposition, activity/ severity and responsiveness to therapy.
The three components - environment, host genetics and the microbial community - interact to maintain homeostasis in the intestine. The disruption of the stability of this interaction may be a trigger for disease development. Two recent publications shed a new light on the pathogenesis of IBD through the change of the intestinal microbial composition involving two different pathways: helminth infection [9] and lipocalin-2 expression [10], respectively.
Helminth infection, microbial community and IBD. Epidemiologic studies demonstrated a major increase of the incidence of IBD in the developed world, suggesting a change in the environment, including an alteration of the intestinal microbiome and a decreased exposure to intestinal parasites, such as helminths. In mice deficient for the CD susceptibility gene Nod2 (Nod2-/- knockout) [11] it could be demonstrated that small intestinal abnormalities develop in the face of a sustained colonization with the inflammatory bacterium Bacteroides vulgatus, an ubiquitous member of the intestinal microbial community.
Chronic infection of Nod2-/- mice with the parasitic worm Trichuris muris, however, inhibited colonization with inflammatory bacteroides species and promoted the establishment of a protective microbial environment enriched in Clostridiales [9]. Further, the authors demonstrated that individuals from helminth-endemic regions harbour a similarprotective microbial community and deworming treatment reduced Clostridiales and increased Bacteriodales, resulting in an increased IBD incidence. These data support the ‘hygiene hypothesis’ whereby certain individuals are genetically susceptible to the consequences of a changing intestinal microbial community that favors IBD development.
Lipocalin-2 protection from IBD and colon cancer. Lipocalin-2 (Lcn2) is an antimicrobial peptide with high mucosal and fecal concentrations in patients with IBD. It is produced by various cell types, including epithelial cells, and acts as an antimicrobial defense mediator by binding to a subset of bacterial siderophores, thereby preventing bacterial iron acquisition and growth of siderophore-dependent strains. While it has been implicated in several biologic processes, such as acute phase response, erythropoiesis and iron metabolism, its functional role in contributing to IBD development remained unclear.
To decipher the role of Lcn2 in colon inflammation, mice double deficient in Lcn2 and IL-10 (Lcn2-/-/ IL10-/- double knockout) were generated and compared to single knockouts and wild-type animals. The experimental data indicate that Lcn2 expression protects from early onset colitis and the spontaneous emergence of right-sided colonic tumors that result from IL-10 deficiency. The inflammation is driven by IL-6 which also controls tumorigenesis. The Lcn2-/-/ IL10-/- double knockout mice showed major alterations of their intestinal microbial community, especially with respect to the facultative pathogenic Alistipes spp. These contribute to inflammation and tumorigenesis as shown by the transmissibility of the phenotype and the protection by antibiotic therapy. Taken together, the authors demonstrate that Lcn2 protects against intestinal inflammation and tumorigenesis in the face of an altered intestinal microbial composition [10].
Recently, it was discovered that the probiotic Lactobacillus casei strain ATCC 334 produces ferrichrome that inhibits colon cancer progression by apoptosis mediated through the c-Jun N-terminal kinase pathway [12], possibly representing a novel tumor-suppressing strategy.
2.2.2. Obesity
Obesity [13], insulin resistance [14] and khwashiorkor [15] are examples for which a correlation between microbial dysbiosis and the clinical state has been demonstrated. Further, in genetically susceptible hosts, the transplantation of fecal microbiota from healthy donors to patients resulted in clinical improvement. The underlying concept is a ‘common ground hypothesis’ that involves a leaky mucosa caused by various endogenous or exogenous factors, the expansion of opportunistic microbes (dysbiotic pathobionts) with the generation of pathogenic microbial gene products which can be transferred to genetically susceptible individuals [6].
2.2.3. Atherosclerosis and Thrombosis Risk
Recent studies suggest that intestinal microbes are involved in atherosclerosis. In this context, foods rich in choline, phosphatidylcholine and carnitine such as meat, egg yolk and high-fat dairy products, serve as precursors of trimethylamine (TMA) and TMA N-oxide (TMAO) that accelerates atherosclerosis and elevated TMAO blood levels are associated with an increased risk for atherosclerotic heart disease and incident major adverse cardiac events. Further, TMAO enhances platelet hyperreactivity and thrombotic events in studies of animal model, employing dietary choline or TMAO, germ-free mice and microbial transplantation, collectively confirming the key role of intestinal microbiota [16]. Taken together, these results reveal a previously unknown link between specific dietary nutrients, intestinal microbes and atherosclerosis/ thrombosis risk (Figure 2).
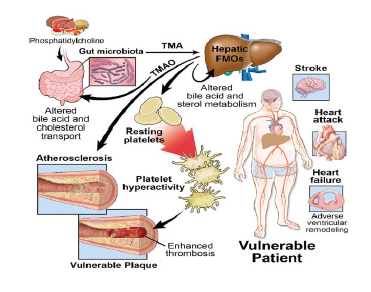
Figure 2: The intestinal microbial metabolite trimethylamine (TMA) and the atherosclerosis/ thrombosis risk [16]
The intestinal TMAO formation is a two-step process involving the generation of TMA by intestinal microbes after food ingestion and the hepatic conversion of TMA to TMAO by host flavin monooxygenases. In a recent, study Wang et al. [17] demonstrated that 3,3-dimethyl-1-butanol (DMB), a structural analog of choline, blocks the intestinal TMA formation through the inhibition of the microbial TMA lyase, and results in reduced TMAO levels. Thus, ‘drugging the microbiome’ with DMB may be a novel approach for the prevention/ treatment of atherosclerosis (Figure 3).
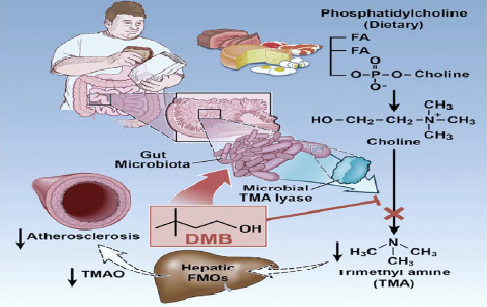
Figure 3: Inhibition of intestinal microbial trimethylamine (TMA) synthesis by 3,3-dimethyl-1-butanol (DMB) and attenuation of atherosclerosis [17].
2.2.4. Neurodevelopmental, Psychiatric and Neurodegenerative Diseases
Studies investigating the intestinal microbial-brain communication (gut-brain axis) demonstrate a critical role of the intestinal microbial community in modulating the maturation and function of tissue-resident immune cells in the central nervous system (CNS) as well as the activation of peripheral immune cells involved in neuroinflammation, brain injury, autoimmunity and neurogenesis [18]. Germ-free mice raised under sterile conditions or mice depleted of their intestinal microbiota by antibiotics show major alterations in behaviours or neuropathologies that characterize neurodevelopmental, psychiatric and neurodegenerative disorders. These include among others autism spectrum disorders, depression and Alzheimer’s or Parkinson’s disease (Table 2).
An impressive example for a pathogenic role of the intestinal microbial community is Parkinson’s disease (PD). In patients with PD, plaques in brain cells as well as in the intestine containing the neurotoxic protein alpha-synuclein (AS) are a hallmark of the disease. For example, in PD patients gastric motility is frequently impaired and AS levels in the intestine are elevated.
In an animal model, mice overexpressing AS indeed develop neurologic deficits resembling those of patients with PD. Recently, three lines of evidence demonstrated a central role of the intestinal microbial community in the pathogenesis of PD (Figure 4): (1) in the PD mouse model germ-free animals developed fewer plaques and almost no neurological deficits as compared to conventionally colonized controls, (2) treatment of PD mice with antibiotics resulted in an improvement of the neurological deficits and (3) fecal transplantation from patients with PD to germ-free mice resulted in neurological deficits resembling PD [19].
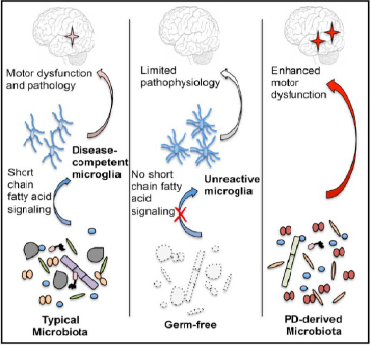
Figure 4: Contribution of short chain fatty acids (SCFA) and fecal tansplantation from patients with Parkinson’s disease (PD) to PD pathogenesis and its prevention by elimination of the intestinal microbiota [19].
The underlying concept is the central contribution of the microbial community to a defect of the microglia via short-chain fatty acids (SCFAs) that represent bacterial fermentation products. While germ-free mice showed a reduced microglia mice, SCFAs modulate microglia and enhance PD pathophysiology [19]. The identification of the disease-causing bacteria and the mechanism leading to the deposition of the neurotoxic AS plaques await further clarification.
3. Conclusions and Perspectives
Basic biomedical research has made major advances in recent years and holds the promise to increasingly provide individual diagnostic, preventive as well as therapeutic options for patients with inherited or acquired, malignant or non-malignant diseases. Apart from an increasing number of host genetic susceptibility loci and environmental factors, the individual microbial community is central for the barrier between microbes and hosts. In particular, the intestinal microbial community is involved in a large number of normal biological functions in health (Table 1) as well as in numerous common, gastrointestinal and non-gastrointestinal diseases, such as obesity/ metabolic syndrome, atherosclerosis/ thrombosis or neurodevelopmental, psychiatric and neurodegenerative diseases (Table 2). Dietary interventions targeting intestinal microbiota, such as calory restricted diets rich in fiber and vegetables as well as fecal microbial transplantation are examples for health benefits in humans. In recent years, the intestinal microbiome thus has become one of the most dynamic areas of biomedical research that holds an enormous potential for interventions regarding human health and disease.
Acknowledgement
The excellent secretarial assistance of Mrs. Mariette Gutgsell is gratefully acknowledged.
Conflict of Interests
The author declares no conflict of interest.
Financial Disclosure
The author has no financing to disclose.
References
- Manolio TA, Collins FS. The HapMap and genome-wide association studies in diagnosis and therapy. Annu Rev Med 60 (2009): 443-456.
- Hou L, Heilbronner U, Degenhardt F, et al. Genetic variants associated with response to lithium treatment in bipolar disorder: a genome-wide association study. Lancet 387 (2016):1085-1093.
- Human Microbiome Project C. A framework for human microbiome research. Nature 486 (2012): 215-221.
- Morgan XC, Segata N, Huttenhower C. Biodiversity and functional genomics in the human microbiome. Trends Genet 29 (2013): 51-58.
- Rossen NG, Fuentes S, van der Spek MJ, et al. Findings from a randomized controlled trial of fecal transplantation for patients with ulcerative colitis. Gastroenterology 149 (2015): 110-118.
- Lynch SV, Pedersen O. The human intestinal microbiome in health and disease. N Engl J Med 375 (2016): 2369-2379.
- Li J, Jia H, Cai X, et al. An integrated catalog of reference genes in the human gut microbiome. Nat Biotechnol 32 (2014): 834-841.
- Fujimura KE, Sitarik AR, Havstad S, et al. Neonatal gut microbiota associates with childhood multisensitized atopy and T cell differentiation. Nat Med 22 (2016): 1187-1191.
- Ramanan D, Bowcutt R, Lee SC, et al. Helminth infection promotes colonization resistance via type 2 immunity. Science 352 (2016): 608-612.
- Moschen AR, Gerner RR, Wang J, et al. Lipocalin 2 protects from inflammation and tumorigenesis associated with gut microbiota alterations. Cell Host Microbe 19 (2016): 455-469.
- Cleynen I, Boucher G, Jostins L, et al. Inherited determinants of Crohn's disease and ulcerative colitis phenotypes: a genetic association study. Lancet 387 (2016): 156-167.
- Konishi H, Fujiya M, Tanaka H, et al. Probiotic-derived ferrichrome inhibits colon cancer progression via JNK-mediated apoptosis. Nature communications 7 (2016): 12365.
- Ridaura VK, Faith JJ, Rey FE, et al. Gut microbiota from twins discordant for obesity modulate metabolism in mice. Science 341 (2013): 1241214.
- Pedersen HK, Gudmundsdottir V, Nielsen HB, et al. Human gut microbes impact host serum metabolome and insulin sensitivity. Nature 535 (2016): 376-381.
- Subramanian S, Huq S, Yatsunenko T, et al. Persistent gut microbiota immaturity in malnourished Bangladeshi children. Nature 510 (2014): 417-421.
- Zhu W, Gregory JC, Org E, et al. Gut microbial metabolite TMAO enhances platelet hyperreactivity and thrombosis risk. Cell 165 (2016): 111-124.
- Wang Z, Roberts AB, Buffa JA, et al. Non-lethal inhibition of gut microbial trimethylamine production for the treatment of atherosclerosis. Cell 163 (2015): 1585-1595.
- Fung TC, Olson CA, Hsiao EY. Interactions between the microbiota, immune and nervous systems in health and disease. Nature neuroscience 20 (2017): 145-155.
- Sampson TR, Debelius JW, Thron T, et al. Gut microbiota regulate motor deficits and neuroinflammation in a model of Parkinson's disease. Cell 167 (2016): 1469-1480.
- Honda K, Littman DR. The microbiota in adaptive immune homeostasis and disease. Nature 535 (2016): 75-84.
- Vatanen T, Kostic AD, d'Hennezel E, et al. Variation in microbiome LPS immunogenicity contributes to autoimmunity in humans. Cell 165 (2016): 842-853.
- Thaiss CA, Zmora N, Levy M, Elinav E. The microbiome and innate immunity. Nature 535 (2016): 65-74.
- Ijssennagger N, Belzer C, Hooiveld GJ, et al. Gut microbiota facilitates dietary heme-induced epithelial hyperproliferation by opening the mucus barrier in colon. Proc Natl Acad Sci 112 (2015): 10038-10043.
- Cho I, Yamanishi S, Cox L, et al. Antibiotics in early life alter the murine colonic microbiome and adiposity. Nature 488 (2012): 621-626.
- Reinhardt C, Bergentall M, Greiner TU, et al. Tissue factor and PAR1 promote microbiota-induced intestinal vascular remodelling. Nature 483 (2012): 627-631.
- Yano JM, Yu K, Donaldson GP, et al. Indigenous bacteria from the gut microbiota regulate host serotonin biosynthesis. Cell 161 (2015): 264-276.
- Spor A, Koren O, Ley R. Unravelling the effects of the environment and host genotype on the gut microbiome. Nat Rev Microbiol 9 (2011): 279-290.


 Impact Factor: * 5.8
Impact Factor: * 5.8 Acceptance Rate: 71.20%
Acceptance Rate: 71.20%  Time to first decision: 10.4 days
Time to first decision: 10.4 days  Time from article received to acceptance: 2-3 weeks
Time from article received to acceptance: 2-3 weeks 