Unnatural Nucleotides: Exploring the Information Storage System of DNA
Aarti Yadav1*, Khushi Khera1, Rekha Mehrotra1, Kohinoor Kaur1, Yamini Agrawal2, Pratima Srivastava3 and Suresh Thakur4
1Shaheed Rajguru College of Applied Sciences for Women, University of Delhi, Vasundhara Enclave, Delhi-110096, India
2Department of Botany, University of Delhi, Vasundhara Enclave, Delhi-110096, India
3Arka Jain University, Mohanpur, Amaria Jamshedpur, Jharkhand-832108, India
4Trivitron Healthcare Pvt Ltd, AMTZ Campus, Pragati Maidan, Visakhapatnam-530031, India
*Corresponding author: Aarti Yadav, Shaheed Rajguru College of Applied Sciences for Women, University of Delhi, Vasundhara Enclave, Delhi-110096, India.
Received: 05 October 2022; Accepted: 27 October 2022; Published: 04 November 2022
Article Information
Citation: Aarti Yadav, Khushi Khera, Rekha Mehrotra, Kohinoor Kaur, Yamini Agrawal, Pratima Srivastava and Suresh Thakur. Unnatural Nucleotides: Exploring the Information Storage System of DNA. Archives of Clinical and Biomedical Research 6 (2022): 916-932.
View / Download Pdf Share at FacebookAbstract
The conventional genetic alphabet has been expanded now with the development of several synthetic unnatural base pairs. The unnatural base pairs formed by recognition and interaction of these unnatural bases are able to maintain the standard Watson and crick structure of DNA as well. Furthermore, these unnatural nucleotides developed by various groups have the ability to store and transcribe the information just like the natural DNA. The review describes the efficient replication, transcription and translation of Unnatural Base Pairs (UBPs) highlighting the possibility of the expanded genetic alphabets. The review also outlines the vast applications of UBPs as novel information storage components as well as in creation of semi-synthetic organisms expressing non canonical amino acids, high affinity aptamer generation, PCR based diagnostics and sitespecific labelling of RNAs.
Keywords
<p>AEIGS; Semi-Synthetic Organisms; Unnatural nucleotides; Unnatural basepairs; Unnatural Base Pairs</p>
Article Details
Abbreviation
: AEIGS- Artificially Expanded Genetic Information System; DNA- Deoxyribonucleic Acid; RNA- Ribonucleic Acid; SSO-Semi-Synthetic Organisms; UBPs- Unnatural Base Pairs; NMR- Nuclear Magnetic Resonance; dNTP- Deoxy Nucleotide Triphosphate; PCR- Polymerase Chain Reaction1. Introduction
The genetic information of life on Earth have been conserved in natural nucleotides-A, G, C, T(U) forming two exclusive sets of base pairs A-T(U) & G-C, that complementing to the fundamental processes of DNA replication & RNA transcription by several polymerases and translation to functional proteins via ribosomal reactions. In due course, several attempts have been made in order to expand the genetic code by addition of Unnatural Base Pairs (UBPs) with the aim to increase biomolecular functionality and the potential to create Semi-Synthetic Organisms (SSOs) containing more than four letter bases [1–4]. These unnatural nucleotides are chemically synthesized to bind each other via non-standard hydrogen bonding, hydrophobic interactions and other shape complementing patterns [2,5]. The standard Watson-Crick DNA model justifies the size-complementarity and the hydrogen bonding patterns responsible for the crystal structure which is necessary for the amplification of genetic information and specificity to the formation of base pairs [6]. Thus, while inculcating the UBPs in a DNA structure, their aromatic scaffolds must exhibit hydrophobic packing and must have hydrogen bond donor groups for the artificial nucleotides to interact with polymerases for their efficient incorporation in the non-template strand to bind with their UB partner in template strand [1–3].
2. Groups of Unnatural Base Pairs
Several groups have developed synthetic unnatural nucleotides that can form unnatural base pairs with their complementary synthetic bases.
2.1 Benner’s UBPs
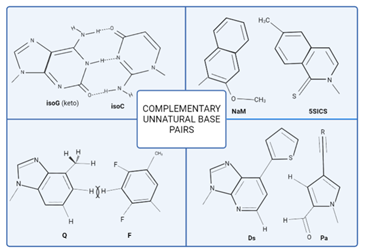
In 1990, Steven Benner and his group were successful in creating structural analogues of the G-C pair by exchanging the carbonyl and amino positions and forming non-standard hydrogen bonding patterns, yielding the isoG-isoC base pair (Figure 1). However, the tautomerization of isoG under physiological pH and instability of isoC under alkaline pH lowered the incorporation efficiencies which further led to the development of P-Z pair i.e., 6-amino-5- nitro-2(1H)-pyridone (Z) and 2-aminoimidazo[1,2-a]-1,3,5-triazin-4(8H)-one (P) pair [5,7]. The P-Z pair depicted the standard Watson-Crick geometry and differed from G-C pair in the configuration of hydrogen bond donor and acceptor groups. These practices led to the development of an artificially expanded genetic information system (AEGIS) [7]. X-ray crystallography revealed that the P-Z pair showed edge-to-edge similarity to standard Watson-Crick model when present in A-form and B-form duplexes and considerably high fidelity in DNA replication using Taq DNA Polymerase. The P-Z pair was then used as 6-letter DNA aptamers to target specific cells and after the discovery of B-S base pair too, they were used as a series of 8-letter DNA and RNA [1,2]. But due to the ability of P base pair to interact and mis incorporate C by forming two hydrogen bonds, Hirao and his co-workers introduced the concept of steric hindrance to hydrogen bonding in order to reduce the possibility of disincorporation. Based on this concept, they developed the x-y pair and s-y pair. Base pairs x and s, having large residues at the 6th position of 2-amino purine scaffold sterically hindered the 4-keto group of T. Base pair y, due to the presence of relatively small hydrogen at position corresponding to T nucleotide, could not bond with it. However, the inability to exclude y base pair to mismatch with A decreases its selectivity of incorporation with s or x and thus, ruling it out to perform replication process [1,8] (Figure 1).
Figure 1: Complementary unnatural base pairs. Chemical structures of unnatural base pairs. Benner’s team created the isoG–isoC base pair. Romesberg’s team created the NaM–5SICS base pair. Kool’s team developed the Q–F base pair. Hirao’s team created the Ds-Pa base pair. Dotted lines represent H-bonds and R represents functional groups. This figure is recreated from "Kimoto et al. (2020) using biorender software [1].
2.2 Kool’s UBPs
Around the same time, Eric Kool’s team reported non-hydrogen bonded A-T analogues, Q-F and Z-F artificial nucleotides bonded by hydrophobic interactions. The Z-F pair, however, lacked the N3 moiety of purines and the 2-keto group of pyrimidines, which interact with polymerases and thus lowered the efficiency of replication (Figure 1). In the Q-F pair, N3 of purine interacted with the side chains of polymerase amino acids and thus, showed compatible replication like the natural A-T pair [1,2]. The study highlighted the importance of shape complementarity between base pairs rather than directional interactions.
2.2 Hirao’s UBPs
Hirao and his coworkers noticed that the shape complementarity of the Q-F pair can be improved by pairing Q with a 5-membered ring analogue instead of a 6-membered ring for F base. This led to the development of Pa (pyrrole-2-carboaldehyde) with a pyrrole scaffold and an aldehyde group, which interacted with polymerases (Fig 1). The selectivity in replication was higher for Q-Pa than Q-F pair [1,2,5]. NMR analysis revealed complementing planar geometry and precise accommodation in the B-DNA duplex. It was also discovered that Pa also pairs with s base and can easily be incorporated in RNA, thus increasing the transcription efficiency of the pair [1,2]. During further attempts to of improve ing the shape complementarity of the s-Pa pair, Hirao and his coworkers removed the hydrogen binding 1-Nitrogen and 2-amino groups from s (or substituted methyl group of Q base with thienyl moiety) and designed hydrophobic Ds base. The Ds-Pa pair had a higher replication and transcription efficiency. Although, there was a high chance of Ds-Ds dimer formation and stacking due to high hydrophobicity and misincorporation of Pa with A in the DNA duplex. This problem was solved by employing modified triphosphates, γ-amidotriphosphates which reduced the mispairing significantly [1,2,8]. The replication fidelity of the Ds-Pa pair, using Vent DNA polymerase was found to be 99.0% but after 20 cycles of PCR the unnatural base pair selectivity reduced to 96%-97%. To solve this problem, the aldehyde group of the Pa base was replaced by a nitro group and a propynyl group was conjugated to design the Px base [1,9]. This slightly decreased the hydrophobicity, and improved the interaction with polymerases, and the better shape complementarity which gave rise to better selectivity of dPxTP opposite Ds. The selectivity thus increased to 99.9% per cycle and the misincorporation reduced to less than 0.01% per cycle per base pair. Since, the Px base could be modified with any functional group, such as amino, diols, dyes, biotin, azides, or ethynyl, it resulted in the development of qPCR, better DNA aptamers and RNA labelling methods [1, 2, 9, 10].
2.3 Romesberg’s UBPs
In 1999, Romesberg and his co-workers reported a hydrophobic self-base pair propynyl isocarbostyril (PICS-PICS) having high incorporation efficiency along with better selectivity and high duplex stability. The dPICSTP could be inserted opposite PICS in replication as a natural base by using exo- Klenow fragments of E. coli Polymerase I [1, 2]). However, due the partial stacking nature of PICS-PICS pair, the extension process, after incorporation, was extremely poor. This problem was resolved by using new polymerases and extensively studying structure-activity relationship (SAR) through chemical approaches. During screening nucleotide insertions and PCR techniques, they discovered the MMO2-5SICS pair, in which they further modified MMO2 base scaffold to give NaM. The NaM-5SICS pair showed higher than 99.9% PCR amplification selectivity and 88% transcription fidelity with T7 RNA Polymerase [1,2,11–14]. Marx et al. determined the prechemistry of 5SICS-NaM pair and its crystal structure revealed that with Klen Taq DNA Polymerase the 5SICS nucleoside and dNaM present in the template gives an open edge-to-edge conformation, similar to those as the Watson-Crick pairing and forms a cross-strand intercalated structure in a DNA duplex [1,2,15].
Dhami et al. transferred the d5SICS-dNaM pair into a plasmid present in E. coli and created the first Semi-Synthetic Organism (SSO) with increased genetic alphabet. Additional analogues were further created to increase the optimization of the base pair. Romesberg’s team modified the distal ring of 5SICS base and developed NaM-TPT3 pair. The NaM-TPT3 pair depicted higher selectivity and better replication than 5SICS-NaM pair, when incorporated in a DNA duplex [1, 16–18]. Malyshev and his team in 2014 incorporated the 5SICS-NaM and NaM-TPT3 base pairs into E. coli genome and reported efficient replication with high selectivity rate and high transcription fidelity. This was the first successful creation of Semi-Synthetic Organism (SSO) i.e., E. coli with an expanded Six-letter genetic alphabet system [1,17–19]. In further attempts to create a more reliable SSO, Feldman and Romesberg embedded the NaM-TPT3 base pair in constructive plasmids to increase UBP retention and developed CNMO-TPT3 pair with more than 90% retention. In order to further enhance the storage and retrieval of information, NaM-TAT1 base pair was constructed. The development of these UBPs made some significant progress towards fluent expansion of the genetic alphabet and creation of SSOs. This, further, helped in the understanding of protein synthesis with higher density ncAAs (non-canonical amino acids) and their site-specific incorporation [1,20].
2.4 Replication and DNA Polymerases
For life to reproduce and harness genetic inheritance from one generation to another, enzymes such as DNA polymerases, RNA polymerases, and reverse transcriptases are essential indispensable [21–25]. DNA polymerases are especially important in this respect, particularly their interaction with peculiar nucleotides. Even in the absence of interbase hydrogen bonding, few of the wild type bacterial polymerases can replicate DNA containing UBPs that interact via steric complementarity. This discovery of non-canonical base pairs has been used to generate tailored Escherichia coli strains that can more efficiently express proteins containing non-canonical amino acids [26–28].
2.5 Modified DNA
The conventional nucleotides- A, T, G, and C can be altered empirically to provide insights of the Watson-crick DNA in terms of replication and storage of additional biological information. The isocytidine: isoguanine (S:B) pair, filed by Alexander Rich in 1962 may be employed as an additional information storage unit in DNA. Synthetic biologists are also trying to understand how far the underlying principle of WC pairing may be modified. For example, Eric Kool, Floyd Romesberg, and Ichiro Hirao's laboratories have worked to eliminate inter-base hydrogen bonding. Steven Benner and colleagues devised a new method for synthesizing UBPs that take advantage of orthogonal hydrogen bonding patterns [2,12,29–31]. This ultimately led to numerous discoveries about the chemical alterations that can (and cannot) be tolerated in replicating units, particularly how protonation, deprotonation, and various tautomeric forms of nucleobases might affect replication fidelity [32]. These efforts resulted in the development of an artificially expanded genetic system (AEGIS), “hachimoji” DNA, in which the letters A, G, T, and C are replaced by two of the unnatural pyrimidine analogs and their size and hydrogen bond complementing counterparts, the purine analogs [33].
2.6 Altered DNA Polymerases for UBPs
Families A–D of replicative DNA polymerases has been identified, with family D being the most recently discovered DNA polymerase family from Archaea [34]. The family names showed homology with the polA, polB, and polC genes which translate into three canonical polymerases from Escherichia coli: DNA polymerase I, DNA polymerase II, and DNA polymerase III alpha subunit, respectively. DNA repair is performed by polymerases belonging to families X and Y. The potential to fill tiny gaps allows the family X to accomplish base excision repair and double-strand break repair. Some polymerases in the X family can act as polymerases without using a template. Eukaryotic polymerases are grouped in family Y, which has less homology with the previously recognized families. The majority of polymerases in the family Y lack proofreading. They lack proofreading exonuclease domains and have a more accessible active site to absorb base damage, allowing them to avoid DNA lesions [35]. The Klenow fragment produced the first crystal structure for family A polymerase. The crystal exhibited a "right hand" form, with the active site in the "palm," which contains the catalytic amino acids, a "thumb," which binds double-stranded DNA, and "fingers," where the incoming dNTPs bind and interact with the template. Crystallographic studies of the Pyrococcus furiosus (family B) revealed that it has five different domains: the finger, palm, thumb, N-terminal, and exonuclease domains [35]. Altered variants of DNA polymerase with a wide variety of biophysical and catalytic properties will certainly be required for upcoming emerging technologies based on the canonical and other extended genetic alphabets [36]. Despite the advancements in the field, the detailed mechanism of the functionality and incorporation of UBPs by the DNA polymerases is yet to be deciphered [34,37–41]. Further studies are required to fully understand how various factors such as Nucleobase pair complementarity, nucleobase tautomerism, hydrogen bond free energy fluctuations, and the conformational dynamics of the polymerase interact to determine the efficiency and integrity of UBP inclusion [36, 37, 42, 43]. One of the approaches to study the structural and functional properties of polymerases is the directed evolution. It is the most effective way to discover DNA polymerase variations that can integrate UBPs with high efficiency and fidelity [35, 44, 45]. Directed evolution procedures such as compartmentalized self-replication (CSR) or compartmentalized self-tagging (CST) is often used to generate modified DNA polymerases [44, 46–49].
2.7 DNA polymerases Catalyse Crucial Steps; Therefore, they Must Balance Three Opposing Requirements
- High Specificity with no more than one error per billion turnovers. Specificity is imperative at both the replicative (5‘-3’ polymerase activity) as well as proof reading (3’-5’ exonuclease activity) step as it is critical for cell survival. Too many errors can be deleterious to the cell. Reverse transcriptases lack the proofreading domain and are hence more prone to errors. DNA polymerases that are predominantly involved in DNA repair, such as trans-lesion synthesis, also have a lower fidelity. The exocyclic C=O groups of T and C, as well as the N-3 nitrogen in the A and G purine rings, present electron density in the minor groove of the DNA duplex. As a result, the "minor groove scanning theory" puts forth the argument that polymerases donate hydrogen bonds to this electron density in the primer, template, and incoming triphosphate in order to enforce Watson–Crick geometry, and therefore fidelity, on base pair identification.
- Acceptance of four different substrates: To present to the polymerase, four substrates have few common molecular characteristics such as their sizes and geometries. The possible existing combination can be- template dG, dC, dA, dT and dCTP, dGTP, dTTP, and dATP respectively. The four bases in the main groove have distinct functionalities. T has a methyl group, C has hydrogen, G has two hydrogen bond acceptors, and A has one donor and one acceptor.
- High processivity: DNA polymerases must be able to incorporated nucleotides rapidly. coli DNA polymerase copies 4000 nucleotides per second to duplicate its whole genome.
DNA synthesis proceeds in primer dependent manner in 5’ to 3’ direction, where magnesium ions act as cofactor. Initially a complex is formed between the enzyme and DNA duplex consisting of a template strand and complementary primer providing the 3’ hydroxyl group for commencing the replication. dNTPs associate with this complex forming pre-incorporation complex. Processivity is a feature found in replicative DNA polymerases, particularly those in which DNA binding is the rate-limiting step. The count of dNTPs, which are integrated into the growing strand right before template-primer duplex dissociates from the enzyme, is referred to as processivity. Perfectly paired dNTPs stay in the active site longer and thus allows the enzyme to adopt a “closed” conformation enabling the 3’ hydroxyl group in the primer to attack the dNTP's alpha-phosphate in presence of magnesium ions. If the complex includes mismatched dNTPs, kinetics indicates that the active site remains in an "open" conformation which allows breakup of the incoming nucleotide to occur before creation of the new P–O bond. In the event that a mismatched dNTP is used, most replicative polymerases have a 3’–5’ exonuclease domain that can correct the mismatch. The 3’ hydroxyl end of the primer transfers from the polymerase to the exonuclease site, allowing the mismatched nucleotide to be removed. This proofreading phase is necessary to ensure fidelity of replication. After incorporating the correct dNTP, following the release of pyrophosphate, post-incorporation complex is formed, with the primer extended by one nucleobase and serving as the substrate for the next template dependent addition of dNTP. These processes are repeated until the template strand contains no unpaired nucleobases. Several experimental and computational studies on the incorporation of UBPs by natural DNA polymerases indicate that electronic restriction in the minor groove is significant for nonstandard base pairing, even when there is no inter-base hydrogen bonding, as in the Romesberg pair. Furthermore, Z:P has proven to be the most straightforward AEGIS UBP to insert into DNA by polymerase-catalyzed processes, being the only pair with shuffled hydrogen-bonding patterns for which both components present electron density to the minor groove (Table 1). The kinetics of nucleobase incorporation, fidelity and processivity can be measured using a variety of assays. Incorporation efficiency, for example, can be evaluated using nested PCR wherein primers labeled with numerous successive UBPs at the 5’end are used. Only if the polymerase is capable of replicating the UBPs are amplicons produced in the reaction mixture. To test fidelity, the DNA polymerase must duplicate a sequence that contains UBPs. For this PCR reaction is performed. If the UBP is removed during PCR cycles, two distinct restriction sites are produced in the amplification products, based on incorporation of natural nucleotide to replace the UBP. In putative transition mutations, incubating the PCR products with two restriction endonucleases results in cleaved products for the amplicons from which the UBP has been removed. As a result, the degree to which polymerase fidelity differs from 100 percent is reported by this assay. Furthermore, conducting the PCR with various doses of the UBP components allows for a quantitative evaluation of fidelity. Structure based mutagenesis experiments have shown some success, however limited, in changing the substrate choices of DNA polymerases. The sophistication in the conformational changes in polymerase throughout each catalytic cycle indicates a need for a variant which can efficiently manage many residue substitutions. For specific purposes, numerous Taq polymerase variants such as ZP Klentaq, with modified catalytic and/or biophysical features have been generated (Table 2).
|
Family of DNA polymerase |
Found in |
Properties |
References |
|
A |
prokaryotes, eukaryotes and bacteriophages |
5’-3’ polymerization domain; 5’–3’ exonuclease domains |
[36] |
|
Taq |
5’-3’ polymerization domain; 5’–3’ exonuclease domains |
[36] |
|
|
Klentaq (Thermus aquaticus) |
Lacks N-terminal 5’–3’ exonuclease domain; have inactive 3’–5’ exonuclease domain |
[36] |
|
|
ZP Klentaq |
Improved fidelity for UBPs; have mutations at distal end M444V, P527A, D551E, and E832V |
[36] |
|
|
B |
prokaryotes, eukaryotes, archaea, and viruses |
5’-3’ polymerization domain; 3’–5’and 5’-3’ exonuclease domains |
[36] |
|
Deep-Vent |
|||
|
Pfu |
|||
|
Therminator DNA Polymerase |
Inactivate the 3′-5′ exonuclease activity exonuclease deficient; contains mutations in the conserved exonuclease domain (separate from the polymerase active site domain) (D141A/E143A) and a mutation (A485L) in the conserved polymerase active site Region III. |
[25] |
|
|
C |
Bacteria |
Leading strand synthesis when associated with DnaE; 5’-3’ polymerization domain; 3’–5’ and 5’-3’ exonuclease domains. |
[50] |
|
D |
Archaea except Crenarchaea |
Large catalytic subunit (DP2) and a smaller subunit with 30 –50 proofreading exonuclease activity (DP1). |
[34] |
Table 1: Details of the different families of DNA polymerases.
|
DNA polymerase variant |
Amino Acid Substitution |
Properties |
References |
|
Taq variant |
(E602V, A608A, I614M, E615G) |
Incorporates both NTPs and dNTPs with the same efficiency |
[51,52] |
|
(F73V, R205K, K219E, M236T, E434D, A608V) |
Functions at higher temperatures |
[48] |
|
|
M1 (G84A, D144G, K314R, E520G, F598L, A608V, E742G) |
Extend substrates that have 3′-mismatches such as C:C or A:G |
[53] |
|
|
M4 (D58G, R74P, A109T, L245R, R343G, G370D, E520G, N583S, E694K, A743P) |
Extend substrates that have 3′-mismatches such as C:C or A:G |
[53] |
|
|
A597T, L616A, F667Y, E745H |
Improved ability to accept dNTP-ONH2 substrates. (L616A), not previously identified, allows Taq to incorporate both reversible and irreversible terminators. Modeling showed how L616A might open space behind Phe-667, allowing it to move to accommodate the larger 3′-substituent. |
[36,54] |
|
|
E520G, K540I, L616A |
Improved ability to accept dNTP-ONH2 substrates. (L616A), not previously identified, allows Taq to incorporate both reversible and irreversible terminators. Modeling showed how L616A might open space behind Phe-667, allowing it to move to accommodate the larger 3′-substituent. |
[36,54] |
|
|
Δ(1-279) Taq variant M444V/P527A/D551E/E832V |
Pause less when challenged in vitro to incorporate dZTP opposite P in a template. |
[35,36,55] |
|
|
Δ(1-279) Taq variant N580S/L628V/E832V |
Pause less when challenged in vitro to incorporate dZTP opposite P in a template |
[35,36,55] |
|
|
Thermococcus gorgonarius DNA polymerase variant |
D141A, E143A |
replicate xeno-nucleic acids; inactivate the 3’–5’ exonuclease activity |
[36,56] |
|
RT521K (Tgo: V93Q, D141A, E143A, E429G, F445L, A485L, I521L, E664K, K726R) |
enhance template binding and promiscuous RT activity across a range of chemistries (A485L, E429G, I521L, E664K, K726R |
[56] |
|
|
(V93Q) |
abrogate stalling at template uracil |
[56] |
|
|
RT-TKK (RT521K: I114T, S383K, N735K) |
improved 2’OMe-RNA RT activity; reverse transcribe XNA chemistry |
[56] |
|
|
Thermococcus litoralis (Vent DNA Polymerase exo-) |
A488L |
mutating an active site alanine 488 to a larger, more bulky side chain increased the efficiency of modified nucleotides including ddNTPs, rNTPs, and 3′-dNTPs (Cordycepin) |
[25] |
|
Pfu exo-/A486Y (Evans et al., 2000); KOD exo-/A485L (Hoshino et al., 2016); Tgo exo-/A485L (Pinheiro et al., 2012 |
[25] |
||
|
DNA polymerase from Thermococcus sp. 9ºN resulting in the commercial Therminator DNA Polymerase. |
[25] |
||
|
Vent(exo-) and KOD XL are capable of adapting 5-Indolyl-AAdUTP (TAdUTP) |
[57] |
Table 2: Variants of DNA Polymerases.
2.8 Structural Details of the DNA Polymerase
A completely manufactured genome would probably contain consecutive UBPs, just as natural genomes; organisms with that kind of a genome would need a DNA polymerase capable of replicating them. Table 1 states the details for different classes of DNA polymerases. Till date, single UBPs have been successfully replicated using naturally occurring family A and B DNA polymerases. Family a polymerases are mostly found in bacteria, and their peptide fold is defined as having fingers, palm, and thumb domains that are involved in polymerase action. When the corresponding dNTP binds to the fingers domain of family A DNA polymerase, a substantial conformational change takes place in the fingers domain, leading to the creation of a closed complex enabling incorporation. Correctly paired nucleotides are selected in the closed conformation of the polymerase through hydrogen bonding and size complementarity. The fingers domain closure angle of the Geobacillus kaustophilus enzyme for incorporating nucleotides is 37°. Unlike the Geobacillus enzyme, WT Klentaq's incorporation of natural dNTPs necessitates a substantially bigger conformational change, with the fingers domain rotating by 59°. Z:P pairs are more effectively and accurately absorbed by ZP Klentaq, with fingers close down by 64°. Pre- and post-incorporation, protein-nucleic acid interactions for ZP Klentaq are identical to those seen in analogous WT complexes, thus, implying that the evolved polymerase closely resembles the WT enzyme. Family A polymerases have 3’–5’ and 5’–3’ exonuclease domains, the latter of which removes the RNA primers needed for lagging strand production. The N-terminal 5’–3’ exonuclease domain is missing in Klentaq, the Klenow or big fragment of DNA polymerase from Thermus aquaticus DNA polymerase, and has inactive the 3’–5’ exonuclease domain. Family B DNA polymerases, on the other hand, are present in all archaea and have a 3’–5’ exonuclease domain as well as a polymerase domain that is similar to that of family A polymerases. Deep-Vent DNA polymerase (family B) or its combination with Taq DNA polymerase (family A) were used to reproduce hydrophobic non-hydrogen-bonding pairs created by Romesberg (NAM:TPT3) and Hirao (Ds:Px) respectively (Table 1). Replicating templates with consecutive hydrophobic nucleobases, on the other hand, has been significantly more challenging. Interruptions to the DNA double helix produced by the inclusion of hydrophobic UBPs seem to be part of the issue. In duplex DNA, the dNaM: d5SICS pair, for example, stacks in an intercalative fashion, as demonstrated in NMR structures and binary template, primer, Klentaq complexes. If replication is to be successful, at least 6 natural base pairs must separate Ds-Px pairs; to yet, no DNA containing successive NaM: PTP3 (or structurally related hydrophobic UBPs) has been replicated. WT Klentaq polymerase can integrate up to four successive Z:P pairings, and these UBP tetrads have no effect on duplex DNA's capacity to acquire A and B DNA conformations. Conversely, WT Klentaq's Z:P absorption accuracy is only 99.8% per theoretical PCR cycle, referring to easy loss of UBP during PCR-based applications. However, an engineered Klentaq version (ZP Klentaq) with increased fidelity and Z:P integration efficiency was developed. Despite Z:P pairs are easily incorporated in both B- and A-form of DNA and retain standard properties such as groove widths, base stacking parameters; that are consistent with both helical forms, modeling of a P: Z coupled to the WT reveals in order to accommodate the UBP, Klentaq requires significant structural changes within the active site. The hydrophobic core of the palm domain, serving as the enzyme's command centre, contains residue 444. The palm domain connects both the fingers and the thumb domains. M444V was chosen as the best candidate among the amino acid changes in ZP Klentaq for enabling access to greater relative domain mobility when compared to the WT. The palm domain also contains crucial catalytic residues Asp-610 and Asp-785, which coordinate Mg ions with the incoming dNTP in the enzyme's active site. The use of a Val instead of a Met in this position produces open space within the core, which could lead to enhanced motion in the fingers and thumb domains.
2.9 Retrieving Information from Unnatural Nucleotides: Transcription, Translation and Site-Specific Incorporation
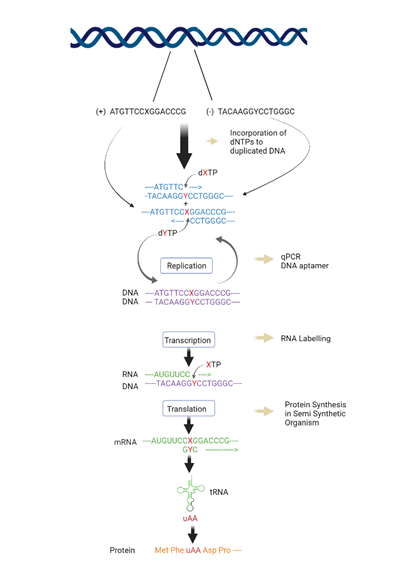
After creation and replication of the UBPs, the genetic information stored in them is meant to be retrieved by transcribing the UBPs into mRNA’s and tRNA’s codons and anticodons and further producing non-canonical amino acids (nCAAs) containing proteins [8]. For successful transcription and translation process to occur, the incorporation of triphosphates of unnatural nucleotides in the template is necessary. In 2014, UBPs were incorporated in the genome of E. coli using a nucleoside triphosphate transporter (PtNTT2) extracted from Phaedactylum tricornutum which imported d5SICSTP and dNaMTP present in media to transcribe unnatural nucleotide containing mRNA and tRNA and translate proteins containing non-canonical amino acids nCAAs from cognate unnatural codons and anticodons [18,20]. In 2019, Eggert et al. identified some Reverse transcriptases (RTs) allowing quantification of rNaM and rTPT3 in RNA transcript and employed four RTs commercially available viz. Avian Myeloblastosis Virus (AMV) RT, Moloney Murine Leukemia Virus (MMLV) RT, SuperScript II (SS II) RT, and SuperScript IV (SS IV) RT. Later, they employed Taq DNA Polymerase with RT activity and replicated NaM:TPT3 pair efficiently. Incorporation efficiency of TPT3 is quantitively high in RNA. Therefore, using RTs and UBP triphosphates, the RNA was reverse transcribed into cDNA product containing the same UBPS [58]. To retrieve the increased genetic information, Zhang et al. (2017) [59] incorporated dNaM-dTPT3 sequence on 151 position of super fold green fluorescent protein of template codon such that sequences AXC and GYT were positioned according to the anticodon template M. mazei Pyl tRNA known to be selectively nCAA charged N6 -(2-azidoethoxy)-carbonyl-L-lysine and were transferred to E. coli. The colonies were left to transform via T7 RNAP. Later, the plasmids were amplified using the biotinylated UBP triphosphates to measure the retention and transcription fidelity. It was discovered that at low concentrations, ncAA incorporation efficiencies are lower and higher in high concentrations [17, 20]. The transcription of UBPs to make RNA transcripts can also facilitate the site-specific incorporation of some unnatural nucleotide triphosphates into the transcribed mRNA and tRNA molecules. In 2004, Endo et al. reported the incorporation of a nucleoside 5’ -tri-phosphate of a hydrophobic y analogue, 5-phenylethynyl-3-(β-D-ribofuranosyl) pyridin-2-one 5-triphosphate (Ph-yTP) in RNA fragments by using T7 RNAP by replacing a uridine residue. The Ph-yTP full-length 17-mer transcript, using a template having s base pair was generated, indicating the fact that Ph-yTP was successfully incorporated in the RNA containing’s. Fluorescent labelling of RNA can also do site-specific incorporation into the RNA at desired positions. Using fluorescent cap analogues like coumarin-derivatized GTP at 5’ termini of RNA by employing T7 transcription or by T7 transcription under T7 2.5 promoter using N6- or 5′-fluorescein-derivatized AMP [60]. Moreover, several biotinylating methods have also been reported. RNA can be biotin labelled at 5’ end via transcription using N6- biotin derivatives of AMP. For example, unnatural base pair bio- yTP can be incorporated by employing T7 RNAP in RNA opposite s or v in template strands [61]. The site-specific incorporation of UBPs by methyl cyclopropene-modified unnatural triphosphate in DNA template provides the sequential information required to incorporate into an unnatural RNA nucleotide which is in-vitro transcribed. This technique creates sequence-specifically label long non-coding RNAs (lncRNAs) which is a hundred nucleotide long [62]. In 2006, Hirao et al. reported T7 transcription of templates consisting of Ds-Pa pair by site-specific radiolabelled incorporation of PaTP and DsTP via T7 RNP. After examining continuous transcription results of the UBPs, 32P-labelled transcripts were run through electrophoresis and found out the transcription fidelity to be 94%. Later, biotinylated 52-mer RNA was used and DNA6 was transcribed containing Ds-Pa pair and natural DNA fragments to yield 47-85% transcription efficiencies [8]. Another way is mediated by extra base pairs via transcription to incorporate UBP TP at desired positions. For instance, to maintain shape complementarity, in 2011, Ohtsuki et al. incorporated x-y pair into a DNA enzymatically and the fragments were left to anneal in the presence of yTP by using T7 Polymerase and partially double stranded DNA fragments. A 35-mer template strand including the promoter sequence were used along with T7 polymerase and a short sequence consisting C and T. This resulted in a 17-mer full length product along with natural nucleotides and yTP. The incorporation of x opposite yTP was proved by the fact that a 15-mer product was yielded in the presence of ATP, GTP and yTP. After analysing the products, it was observed that x was incorporated opposite yTP but a slight incorporation of UTP (5%) opposite x was also took place. This misincorporation was handled by increasing the concentration of yTP [60] (Figure 2).
Figure 2: Transcription, translation and site-specific incorporation in Un-natural base pairs. Expansion of genetic code by creation of unnatural base pairs and their incorporation into fundamental processes like replication, transcription and translation allowing expanded storage and retrieval of expanded genetic information with varied application. This figure is recreated from "Kimoto et al. (2020) using biorender software [1].
3. Applications of UBPs
3.1 qPCR using UBPs
qPCR technique utilized the UBPs developed by Benner and his co-workers i.e., isoG-isoC pair. A primer with a fluorescently labelled T at its 5’ end besides isoC in the hanging part and an isoG NTP conjugated with a quencher dabcyl group (Dab-disoGTP) are used in this technique. The Dab-isoG and isoC primers are incorporated opposite to each other and quenching of the fluorophore in the primer is done by adding Dab-isoG in close proximity to increase the fluorescence intensity during the PCR amplification process [1,63]. Strong fluorescence was also reported in some of the Ds derivatives like s and Dss bases. Some other unnatural bases like y, Pa and Ps bases can also be conjugated with a fluorophore or quencher with the help of a linker. Dss and Px pair can be used as a fluorophore or quencher pair as 2-nitropyrrole moiety of Px acts as a fluorescent quencher. It was also discovered that the due to stacking interactions between the Px base and fluorophores, the free triphosphates quench the fluorescence of the fluorophores. However, when the Px base is incorporated into a DNA duplex, the fluorescence is enhanced as the fluorophore is exposed outside the duplex. On this basis, another qPCR method using specific detection methods with Ds-Px pair was developed [1] Multiplex-based diagnosis kits for the surveillance of emerging mosquito-borne pathogens, such as dengue (DENV1–4), chikungunya (CHIKV), and Zika viruses (ZIKV) utilized the UBPs developed by Benner’s team. The Luminex xMAP DHA protocol with ‘‘self-avoiding molecular recognition systems (SAMRS)’’ is used in RT-PCR techniques to produce single-stranded biotinylated amplicons containing Z by mispairing the G-Z pair. Through a reverse-primer extension/transliteration reaction. Hybrid product with target specific probe using P-Z pair having Luminex beads containing P are obtained [1,64,65].
3.2 DNA Aptamers using UBPs
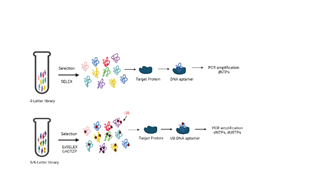
DNA Aptamers are in-vitro generated via an evolutional engineering method known as SELEX, (Systematic Evolution of Ligands by Exponential enrichment) single-stranded DNA fragments having the ability to bind specifically to target molecules (Figure 3). They are generated by repetitive cycles of targeted selection and PCR amplification techniques using DNA libraries [1]. However, the affinity of DNA Aptamers is limited to target proteins. Hydrophobic Ds bases can be incorporated into DNA libraries to generate high affinity DNA aptamer [66]. Further, these enriched DNA libraries are amplified with dPaTP via PCR to incorporate Pa with Ds and further A with Pa to determine DNA base library via deep sequences. Barcodes sequence can be assigned to tell us the position of Ds in a DNA aptamer [67,68]. Besides deep sequencing by replacement PCR, to determine UB position in aptamers, each DNA can also be isolated from libraries by hybridization using immobilized DNA probes or by performing Sanger gap-sequencing of each isolated DNA in which a gap appears in the sequencing peak patterns of the natural bases when Ds position appears [1,69]. Using this method, a DNA aptamer which binds to von Willebrand Factor (vWF) A1 with high affinity was isolated [1]. A series of Ds DNA aptamers were generated targeting three breast cancer cell lines i.e., MCF7, MDA-MB-231, and T-47D using ExSELEX with diverse specificity. These DNA aptamers such as 14A-MCF7, generated by targeting MCF7, and 05-MB231, generated by targeting MDA-MB-231, binds to numerous cancer cells at once without binding to normal cell lines, such as MCF-10A and HUVEC [1,70,71]. Ds-DNA aptamers are made stable against exonucleases and heat due to the conjugation of a mini hairpin DNA with GCGAAGC sequence at 3’ end. This conjugation does not affect the modification site i.e., the central A of mini-hairpin structure where the Ds-DNA aptamer is labelled [72]. The Ds-DNA aptamers, when combined with antibodies can be used for selection of highly sensitive target protein in a Sandwich-type ELISA method [73]. Benner’s and Tan’s team, in 2014, also reported a 6-letter DNA aptamer made of randomized Z and P sequences that bind to the MDA-MB-231 breast cancer cell line, HepG2 liver cancer cells, and glypican-3-overexpressing tumor cells. A long 6-letter GACTZP DNA aptamer, intercalated with doxorubicin was also conjugated which exhibited anti-cancer activity to target liver cancer HepG2 cells [74]. Benner’s team also developed GACTZP DNA aptamers targeting anthrax protective antigen. Recently, a unique method of aptamer generation called ligand-guided selection (LIGS) is reported in which incubation of GACTZP DNA aptamers and target cells is done. The competitive binding of monoclonal antibodies to the target in the same region releases the aptamers. These methods, when combined are used to generate and isolate aptamers binding to CD3 T cell receptors [75,76] (Figure 3).
3.3 RNA Labelling using UBPs
Site-specific incorporation of unnatural bases into RNA transcript allots novel functionality to it. Kimoto et al., in 2002, reported site-specific incorporation via T7 transcription by incorporating y substrate into the transcript opposite s base. Certain functional groups such as biotin for immobilization, iodine for photo-crosslinking and dyes for fluorescent labelling are attached to the y base with the help of propynyl linker to use in T7 transcription for RNA labelling [1, 77, 78]. By fluorescent labelling of the s base, site specific fluorescent RNA labelling can be done opposite Pa base with 97% selectivity. Using this technique, the dynamic structure of a stable GAAA-loop hairpin forming a shared GA pair with two a protruding opposite each other and third stacked above them [79]. Either of the protruding A(s) were replaced with s and with the increase in temperature, the fluorescence and absorbance of these hairpin loops containing s base were measured. After receiving, assuring results, these techniques were used for dynamic analysis of tRNA molecule [73]. Modified Pa base substituted with biotin, dyes, and ethynyl and azide groups are added opposite Ds base with high selectivity [8, 80]. While post transcriptional modification processes, ethynyl and azide derivatives are used for click reactions with any other functional groups [81]. With Ds-Pa, s-Pa and s-y pairs, combining s, modified y and Pa bases for triple-labelling is also possible [1]. Romesberg’s team also reported the incorporation of NaM and MMO2 substrates opposite 5SICS into RNA via T7 RNA Polymerase. However, modified derivatives of NaM are not acceptable for T7 transcription. Clickable 5SICS and amino-acid linked 5SICSand MMO2 were synthesized for post transcriptional processes which were incorporated in 243-mer 16S ribosomal RNA fragment of Thermus thermophilus having modified azide-Cy5 and NHS-Cy3 reagents [82,83]. Kath-Schorr’s team is also known to have synthesized TPT3 substrates labelled with nitroxyl spin for T7 transcription, which was incorporated opposite NaM. 185-mer glmS ribozyme and the 377-mer non-coding Xist RNA were introduced with modified TPT3 bases to measure the inter-spin distance distributions by pulsed electron para-magnetic resonance spectroscopy. By using T7 transcripts containing NaM or TPT3 as templates, reverse transcription of NaM-TPT3 pair was also studied by them. The incorporation efficiency of NaM opposite TPT3 was much higher than the incorporation efficiency of TPT3 opposite NaM [58,84,85].
3.4 UBP Components for SSOs
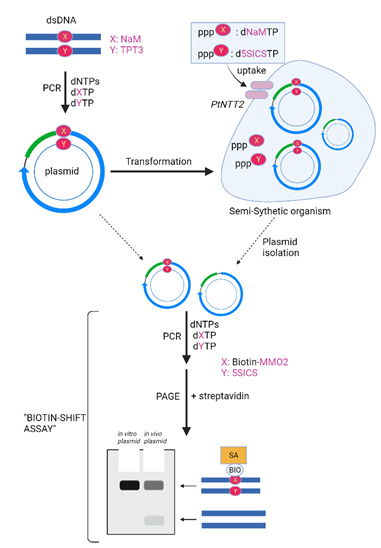
Unnatural bases can also be incorporated in the genetic system of Semi-Synthetic Organisms (SSOs) to create an organism with expanded information to increase functionality. For creation of such cells, triphosphates of UB nucleotides must be continuously supplied in sufficient concentrations as required for DNA replication. Thus, the growth media are often supplemented with the UB nucleosides which are converted into triphosphates with the help of certain kinases as in the nucleoside salvage pathways [86]. Romesberg’s team, in 2002, reported unnatural nucleotides like d5SICS and dNaM were phosphorylated by nucleoside kinases of Drosophila melanogaster (DmdNK) by using ATP, in vitro [18]. Benner was also found to report that DmdNK and its Q81N mutant can phosphorylate dP more effectively than dZ. An assay was done to confirm the broad specificity of E. coli nucleoside diphosphate kinase towards dNDPs (deoxy-ribonucleoside diphosphates) like dZDP and dPDP. It also,less efficiently, phosphorylated dDS and dPx nucleotides [87]. Another method by which the NTPs can be transferred inside the cell was by targeting nucleoside triphosphate transporters (NTTs). Activities of eight different NTTs were tested and it was discovered that Phaeodactylum tricornutum NTT called PtNTT2 which could accumulate d5SICS and dNaM triphosphates in cells. The triphosphates were stabilized in growth media provided to PtNTT2-expressing E. coli which resulted in accumulation of d5SICSTP and dNaMTP in the cytoplasm (Figure 4) [1,18]. A plasmid DNA with a NaM-TPT3 pair was transformed into pCDF-1b expressed E. coli C41(DE3) harbouring PtNTT2 with a continuous supply of d5SICSTP and dNaMTP via liquid media to the transformants. The retention of the plasmid in the transformants was studied by (1) isolation of propagated plasmid for the detection of the corresponding d5SICS peak by direct monitoring of nucleotide compositions by LC-MS/MS, (2) Sanger sequencing patterns and (3) labelling PCR amplification products of plasmid with biotin. It was concluded that the replication fidelity of NaM–5SICS pair in DNA was 99.4% in vivo [1, 18]. Further improvisations were made by Romesberg’s team transporter engineering using codon-usage optimization, chemical optimization of UBPs and usage of CRISPR-Cas9 system to remove errors UBP retention [73] (Figure 4).
Figure 4: Transformation in the Un-natural base pairs by using by Nam and TPT3. Romesberg’s team transformed a plasmid containing NaM-TPT3 pair (prepared by culturing in inorganic phosphate-rich growth media in the presence of the NaM and 5SCIS triphosphates), prepared by PCR into E. coli C41(DE3) by employing a PtNTT2 overexpression system. The retention of NaM-TPT3 pair was examined by biotin-shift assay. This figure is recreated from "Kimoto et al. (2020) using biorender software (1).
3.5 Protein Synthesis using UBPs
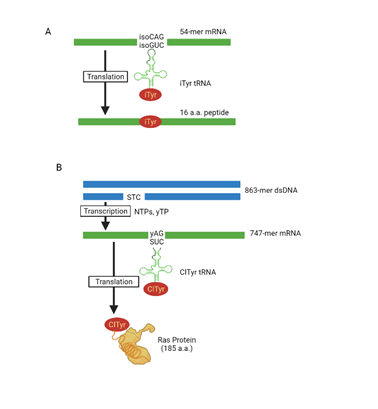
Unnatural amino acids (uAAs) are integrated into proteins via the enlarged codon and anticodon interactions between mRNA and tRNA (Figure 5) In the present three-base codon system, this allows 64 four-letter codons to be expanded to 216 with introduction of just 2 codons making it six-letter codons [1] (Figure 5). Benner and his colleagues published an in vitro rabbit reticulocyte lysate translation system based on the isoG–isoC pair. Chemical and enzymatic procedures were used to chemically synthesis a 56-mer mRNA with an isoCAG codon and create 3-iodotyrosyl-tRNA with a CUisoG anticodon and a 16-amino acid peptide containing 3-iodotyrosine at the specified location was generated [1]. Similarly, Hirao et al, 2002 reported the incorporation of ClTyr (halogenated tyrosine) into position 32 of the Ras protein (s–y pair) using T7 RNA polymerase, E. coli extracts for protein synthesis, the DNA template, and ribonucleoside triphosphates of E. coli in in vitro transcription and translation [1]. Even though semi-synthetic organisms store more information than natural organisms, retrieving it requires in vivo transcription of the unnatural base pair into mRNA and tRNA, aminoacylation of the tRNA with a non-canonical amino acid, and efficient participation of the unnatural base pair in ribosome decoding [59]. Further Zhang et al, successfully carried out the in vivo transcription using semi-synthetic DNA with dNaM and dTPT3 into mRNAs (two different unnatural codons; sfGFP codon 151 (TAC) replaced by the unnatural codon AXC (sfGFP(AXC)151; X denotes NaM); sfGFP(GXC)151) and tRNAs with cognate unnatural anticodons (tRNASer(GYT); Y denotes TPT3 and tRNAPyl(GYC)), and efficient decoding of these mRNAs and tRNAs at the ribosome to direct the site-specific incorporation of natural or non-canonical amino acids into super folder green fluorescent protein in YZ3 strain [59]. They used a super folder green fluorescent protein (sfGFP) 188 as a model translation system and targeted position Y151, which accepts a wide variety of natural and synthetic amino acids. These findings emphasize that interactions other than hydrogen bonding can play a role in information storage and retrieval at every stage. The resulting semi-synthetic creature can both store and retrieve more information, and it might be used to develop the above-mentioned semi-synthetic organisms and functionalities [59]. This technique significantly increases the quantity of data that can be genetically encoded. According to Yorke Zhang, the unnatural base pair makes 152 additional codons available. The approach eventually allows for high-efficiency expression of proteins including several non-natural amino acids, it could be critical for future protein therapies and biotechnologies.
Figure 5: Protein synthesis using UBPs. (A) In vitro translation of isoG-isoC pair allowing incorporation of 3-iodotyrosine (iTyr) in the peptide. (B) In vitro transcription and translation using s-y pair allowing incorporation of 3-chlorotyrosine (ClTyr) into Ras protein. This figure is recreated from "Kimoto et al. (2020) using biorender software [1].
3.6 Chemical Optimization
Through the expressed PtNTT2 encoded in a plasmid under the T7 promoter, Romesberg's team's nascent semi-synthetic organism (SSO), DM1, allowed effective replication of a plasmid harboring a single NaM–5SCIS pair by incorporating its UB triphosphates from the medium within the cells (PT7). The SSO, on the other hand, expanded slowly and easily lost the UBP. As a result, Romesberg improved the SSO by taking into account three points using both genetic/biological and chemical methods: (1) transporter engineering, (2) UBP chemical improvement, and (3) mistake elimination using the CRISPR–Cas9 system. In vitro, Romesberg's group discovered that the NaM–TPT3 pair has superior replication fidelity than the NaM–5SCIS pair [16]. They compared the retention of the NaM–TPT3 pair to that of the NaM–5SCIS pair in 16 distinct sequence contexts (50-NXN-30; N = A, G, C, or T; X = NaM) using the YZ3 strain. They confirmed that the NaM–TPT3 pair retained more than the NaM–5SCIS pair in all of the sequence contexts investigated by biotin-shift assay, [59] though retention of the NaM–TPT3 pair was still modest or poor in some sequence contexts. They used the NaM–TPT3 combination as a reference for further development of their UBPs in subsequent trials.
3.7 Probing
The expansion of the genetic code by an artificial base-pair system is one of the appealing techniques for site-specific fluorescence tagging of RNA. This technique allows for the enzymatic insertion of extra components into RNA at specific locations via transcription mediated by extra base pairs [79]. Unnatural base pairs have recently been produced, such as 7-(2-thienyl) imidazo[4,5-b] pyridine (symbolized as Ds) and pyrrole-2-carbaldehyde (denoted by Pa), 2-amino-6-(2-thienyl) purine (denoted by s) and 2-oxopyridine (denoted by y), and imidazolin-2-one (symbolized as z). In replication and transcription, each base pair has a discrete and distinct selectivity [8, 60, 61]. The Ds–Pa pair complements each other in replication and transcription, allowing polymerases to incorporate Ds and Pa into DNA and RNA at specified sites. In addition, the s–y pair can be employed unidirectional in transcription to incorporate y and modified y bases into RNA opposite s in DNA templates, such as fluorophore-linked y bases [79].
3.8 FRET Characterization
In combination with the Ds–Px pair (Ds analogues, such as the s and Dss bases (a Ds base analogue with an extra thienyl group), have a strong fluorescence in their natural state; Other UBs, such as the y, Pa, and Px bases, can be conjugated with a fluorophore or quencher via a linker without disrupting the UB pairs), the fluorescent s base (s–Pa pair) allows for visible PCR amplification [10]. While the s base emits blue fluorescence at 434 nm whenever excited at 365 nm, two consecutive s bases (ss) in DNA cause self-quenching, with no apparent fluorescence. On the other hand, the quenched ss still works as a FRET donor, allowing visual detection when combined with a FRET acceptor like FAM or Cy3 [23]. As a result, Cy3-dPxTP (excitation: 450–550 nm, emission: B570 nm) was created and integrated it in a proximal site opposite Ds near the position of as in the template. The Cy3 fluorescence was detectable to the naked eye after PCR amplification with Cy3-dPxTP and a primer including ss and Ds at 365 nm excitation. Using this approach, a single nucleotide polymorphism in quinolone resistant genes was identified in E. coli and Streptococcus pneumoniae using visual PCR.
3.9 Codon Optimization using UBP
The four natural nucleotides control all the information in the cell with addition of the above mentioned UBPs such as dNaMTP, d5SICSTP to this genetic information we can further broaden the stored information in these nucleotides [88]. Using molecular biology engineering methods, the unnatural nucleotides have elaborated the traditional genetic codes from 64 codes (61 sense codons and 3 nonsense codons) to the extent which is beyond the limits one can imagine. For these special UBPS therefore it is essential to use the cognate anticodons in the semisynthetic organisms for efficient translation, thereby the desired novel protein production. Using the codon optimized UBPs strains the recombinant proteins of industrial importance and nucleic acid therapies can be exploited to their maximum extent including the gene, messenger ribonucleic acids therapy or the nucleic acid-based vaccines [88, 89]. Further optimization, routine strains creation, transporters, vector designing needs to be carried out for making the process cost effective, durable to stand out in the global markets [90, 91].
3.10 Amplification of Near Conserved Region-using UBP
Unnatural nucleotides have expanded the stored genetic information both in vivo as well as in various invitro PCR based assays. For in vivo semisynthetic organisms have been created which extrapolated the new stored information using the engineered metabolisms as stated in this review. Further for the invitro assays both natural and unnatural nucleotides mixtures have been used with recombinant DNA polymerases, which are well suited for this task [92, 93].
3.11 Probable Role in the Therapeutics and DNA/RNA based Vaccines
Nucleic acid-based vaccines (deoxyribonucleic acid and messenger ribonucleic acid based) development has been critical part of the COVID-2019 pandemic and owing to the fastest growing technology over the conventional vaccine development as they are comparatively cheap and easy to scale up. The recombinant nucleic acid containing additional unnatural nucleotides might prove to be efficacious for vaccines by widening the scope information encoded by that nucleic acid for diseases like cancer, malaria, HIV infection; providing more stability, enhanced expression and may be adding on to the immunogenicity. The usage of these unnatural nucleotides may also help us to combat the problem of anti-microbial resistance as the new information now being coded by them uses the cognate codons and anticodons that can be exploited to overcome the problem. However, the efficacy, dosage, carrier material, route of administration of these nucleic acid-based vaccines with unnatural nucleotides needs to be further examined [94, 95]. These unnatural modified nucleic acids can be developed for therapeutic imaging also as stated above by coupling them with PCR or a reporter dye that could be visualized in a real time fashion in the cell. Further these unnatural nucleotides can also be part of genome editing techniques such as clustered regularly interspaced short palindromic repeats (CRISPR) /CRISPR associated (Cas), antisense technology including peptides and proteins, and small interfering ribonucleic acids, in future to create the genomes with novel properties as well revolutionizing the diagnostics for future outbreaks especially for viruses [96]. Currently numerous drugs are in pipeline containing modifies oligonucleotides and several have been approved by Food and Drug Agency for treatment of diseases like acute hepatic porphyris, Duchene muscular dystrophy [97]. In near future in places of these modified bases these unnatural nucleotides can be used to develop the treatment of such life limiting conditions.
3.12 The Role of UBPS in Therapeutics miRNA synthesized using UBPs/UNs
Traditionally, noncoding regulatory micro ribonucleic acids (miRNA) like other genome modifying techniques was exploited to downregulate the gene of interest by its interaction with 3’ untranslated region and thus facilitating its degradation via ribonucleic acid interference mechanism [98]. The miRNA has been reported in the extracellular matrix also apart from intracellular fluids and their levels can be analysed for the diseased status, subcellular localization, and state of the cell. The levels of the miRNA directly affect the post translational repotire existing in the cell thereby controlling the cellular dynamics [99]. With incorporation of these UBPs in these miRNAs like modified oligonucleotide might prove to be an efficacious edition to the nucleic acid pool for target-based therapeutics.
4. Conclusion
The development and introduction of unnatural nucleotides and amino acids can revolutionize the domain of molecular biology and therapeutics to a great extent and increase the functionalities of genetic material. Synthesis of SSOs has also proven to be successful with an expanded genetic alphabet and extended practical applications. Modification of the genetic makeup of an organism facilitates the analysis of protein functions and interactions. The successful incorporation of synthetic base pairs offers deeper insights into structures and bonding between different bases as well as effect of modification on organisms’ metabolism. Replication of UBPs with the help of special polymerases produced by point mutations and chemical alterations demonstrating high specificity and fidelity presents the possibility of generation of successful SSOs. Easy retrieval of information by transcription and generation of uAAs by translation can be vastly applied to the field of high-affinity DNA aptamer generation, antibody generation, rapid diagnostic kits, RNA labelling, CRISPR technology, transporter engineering etc. The evolution of four-letter bases to six- and eight-letter base libraries have immensely contributed to the development of SELEX technology and enhanced chemical optimization. [1, 6]. Further advancements, in terms of, sequence authentication and data storage in order to augment the functionality of UBPs is still in its infancy [92]. The progress made through the development of unnatural bases can prove to be evolutionary in the field of synthetic biology.
5. Future Prospects
UBPS needs to be further validated and expanded for eukaryotic model organisms leading to the creation of eukaryotic semi-synthetic organisms, which can provide insight into the creation of life. UBPs may play a pivotal role in developing therapeutics for various diseases such as cancers and autoimmune disorders. A single modifiable change led to the development of interleukin-2 (IL-2) variant THOR-707 that may be used for solid cancers. Similarly, other UBP combinations can be exploited and used especially with emerging drug resistance cases. However, the efficacy of such a combination will require clinical trials. UBPs can also be crucial for the development of DNA modification systems such as Restriction enzymes, CRISPER-Cas9, DNA sequencing using next-generation platforms but these areas need to be further explored.
Acknowledgement
The authors would like to thank the management of Shaheed Rajguru College of Applied Sciences for Women, University of Delhi, for their contributions. The manuscript was written through contribution of all authors. All authors have given approval for the final version of the manuscript.
Conflict of Interest
There is no conflict of interest.
References
- Kimoto M, Hirao I. Genetic alphabet expansion technology by creating unnatural base pairs, Chemical Society Reviews 49 (2020): 7602-7626.
- Hirao I, Kimoto M, Yamashige R. Natural versus artificial creation of base pairs in DNA: Origin of nucleobases from the perspectives of unnatural base pair studies, Accounts of Chemical Research 45 (2012): 2055-2065.
- Hamashima K, Kimoto M, Hirao I. Creation of unnatural base pairs for genetic alphabet expansion toward synthetic xenobiology, Current Opinion in Chemical Biology 46 (2018): 108–114.
- Lee KH, Hamashima K, Kimoto M, et al. Genetic alphabet expansion biotechnology by creating unnatural base pairs, Current Opinion in Biotechnology 51 (2018): 8-15.
- Galindo-Murillo R, Barroso-Flores J. Hydrophobic unnatural base pairs show a Watson-Crick pairing in micro-second molecular dynamics simulations, Journal of Biomolecular Structure & Dynamics 38 (2020): 4098-4106.
- Benner SA, Hutter D, Sismour AM. Synthetic biology with artificially expanded genetic information systems. From personalized medicine to extraterrestrial life, Nucleic Acids Research (2003): 125-126.
- Wang W, Sheng X, Zhang S, et al. Chen, Theoretical characterization of the conformational features of unnatural oligonucleotides containing a six nucleotide genetic alphabet, Physical Chemistry Chemical Physics 18 (2016): 28492-28501.
- Hirao I, Kimoto M, Mitsui T, et al. An unnatural hydrophobic base pair system: site-specific incorporation of nucleotide analogs into DNA and RNA, Nature Methods 3 (2006): 729-735.
- Kimoto M, Sato A, Kawai R, et al. Site-specific incorporation of functional components into RNA by transcription using unnatural base pair systems, Nucleic Acids Symposium Series 53 (2009): 73-74.
- Yamashige R, Kimoto M, Okumura R, et al. Visual Detection of Amplified DNA by Polymerase Chain Reaction Using a Genetic Alphabet Expansion System, Journal of the American Chemical Society 140 (2018): 14038-14041.
- Leconte AM, Hwang GT, Matsuda S, et al. Discovery, characterization, and optimization of an unnatural base pair for expansion of the genetic alphabet, Journal of the American Chemical Society 130 (2008): 2336-2343.
- Malyshev DA, Romesberg FE. The expanded genetic alphabet, Angewandte Chemie (International Ed. in English) 54 (2015): 11930-11944.
- Matsuda S, Leconte AM, Romesberg FE. Minor groove hydrogen bonds and the replication of unnatural base pairs, Journal of the American Chemical Society 129 (2007): 5551-5557.
- Leconte AM, Matsuda S, Romesberg FE. An efficiently extended class of unnatural base pairs, Journal of the American Chemical Society 128 (2006): 6780-6781.
- Betz K, Malyshev DA, Lavergne T, et al. Structural insights into DNA replication without hydrogen bonds, Journal of the American Chemical Society 135 (2013): 18637-18643.
- Dhami K, Malyshev DA, Ordoukhanian P, et al. Systematic exploration of a class of hydrophobic unnatural base pairs yields multiple new candidates for the expansion of the genetic alphabet, Nucleic Acids Research 42 (2014) 10235-10244.
- Dien VT, Holcomb M, Feldman AW, et al. Progress toward a Semi-Synthetic Organism with an Unrestricted Expanded Genetic Alphabet, Journal of the American Chemical Society 140 (2018): 16115-16123.
- Malyshev DA, Dhami K, Lavergne T, et al. A semi-synthetic organism with an expanded genetic alphabet, Nature 509 (2014): 385-388.
- Feldman AW, Romesberg FE, In Vivo Structure-Activity Relationships and Optimization of an Unnatural Base Pair for Replication in a Semi-Synthetic Organism, Journal of the American Chemical Society 139 (2017): 11427-11433.
- Feldman AW, Dien VT, Karadeema RJ, et al. Optimization of Replication, Transcription, and Translation in a Semi-Synthetic Organism, Journal of the American Chemical Society 141 (2019): 10644-10653.
- Goodman MF, Tippin B. The expanding polymerase universe, Nature Reviews. Molecular Cell Biology 1 (2000): 101-109.
- Young RA. RNA polymerase II, Annual Review of Biochemistry 60 (1991): 689-715.
- Autexier C, Lue NF. The structure and function of telomerase reverse transcriptase, Annual Review of Biochemistry 75 (2006): 493-517.
- Aschenbrenner J, Marx A. DNA polymerases and biotechnological applications, Current Opinion in Biotechnology 48 (2017): 187-195.
- Gardner AF, Jackson KM, Boyle MM, et al. Therminator DNA Polymerase: Modified nucleotides and unnatural substrates, Frontiers in Molecular Biosciences 6 (2019): 28.
- Betz K, Malyshev DA, Lavergne T, et al. Marx, KlenTaq polymerase replicates unnatural base pairs by inducing a Watson-Crick geometry, Nature Chemical Biology 8 (2012): 612-614.
- Fischer EC, Hashimoto K, Zhang Y, et al. New codons for efficient production of unnatural proteins in a semisynthetic organism, Nature Chemical Biology 16 (2020): 570-576.
- Liu CC, Schultz PG. Adding new chemistries to the genetic code, Annual Review of Biochemistry 79 (2010): 413-444.
- Kool ET. Replacing the Nucleobases in DNA with Designer Molecules, Accounts of Chemical Research 35 (2002): 936-943.
- Feldman AW, Romesberg FE. Expansion of the Genetic Alphabet: A Chemist’s Approach to Synthetic Biology, Accounts of Chemical Research 51 (2018): 394-403.
- Yamashige R, Kimoto M, Takezawa Y, et al. Highly specific unnatural base pair systems as a third base pair for PCR amplification, Nucleic Acids Research 40 (2012): 2793-2806.
- Eberlein L, Beierlein FR, van Eikema Hommes NJR, et al. Tautomeric Equilibria of Nucleobases in the Hachimoji Expanded Genetic Alphabet, Journal of Chemical Theory and Computation 16 (2020): 2766-2777.
- Hoshika S, Leal NA, Kim MJ, et al. SantaLucia, A.J. Meyer, S. DasGupta, J.A. Piccirilli, A.D. Ellington, J. SantaLucia, M.M. Georgiadis, S.A. Benner, Hachimoji DNA and RNA: A genetic system with eight building blocks, Science (New York, N.Y.) 363 (2019): 884-887.
- Raia P, Delarue M, Sauguet L. An updated structural classification of replicative DNA polymerases, Biochemical Society Transactions 47 (2019): 239-249.
- Laos R, Thomson JM, Benner SA. DNA polymerases engineered by directed evolution to incorporate non-standard nucleotides, Frontiers in Microbiology 5 (2014).
- Ouaray Z, Benner SA, Georgiadis MM, et al. Building better polymerases: Engineering the replication of expanded genetic alphabets, The Journal of Biological Chemistry 295 (2020): 17046-17059.
- Raper AT, Reed AJ, Suo Z. Kinetic Mechanism of DNA Polymerases: Contributions of Conformational Dynamics and a Third Divalent Metal Ion, Chemical Reviews 118 (2018): 6000-6025.
- Steitz TA. DNA polymerases: structural diversity and common mechanisms, The Journal of Biological Chemistry 274 (1999): 17395-17398.
- Johnson KA. The kinetic and chemical mechanism of high-fidelity DNA polymerases, Biochimica et Biophysica Acta 1804 (2010): 1041-1048.
- Kropp HM, S.L. Dürr SL, Peter C, et al. Snapshots of a modified nucleotide moving through the confines of a DNA polymerase, Proceedings of the National Academy of Sciences of the United States of America 115 (2018): 9992-9997.
- Wu WJ, Yang W, Tsai MD. How DNA polymerases catalyse replication and repair with contrasting fidelity, Nature Reviews Chemistry 1 (2017): 1-16.
- Joyce CM, Benkovic SJ. DNA polymerase fidelity: Kinetics, structure, and checkpoints, Biochemistry 43 (2004) 14317-14324.
- Kunkel TA, Bebenek K. DNA replication fidelity, Annual Review of Biochemistry 69 (2000): 497-529.
- Houlihan G, Arangundy-Franklin S, Holliger P. Exploring the Chemistry of Genetic Information Storage and Propagation through Polymerase Engineering, Accounts of Chemical Research 50 (2017): 1079-1087.
- Chen T, Romesberg FE, Directed polymerase evolution, FEBS Letters 588 (2014): 219-229.
- Pinheiro VB, Taylor AI, Cozens C, et al. Synthetic genetic polymers capable of heredity and evolution, Science (New York, N.Y.) 336 (2012): 341-344.
- Pinheiro VB, Arangundy-Franklin S, Holliger P, Compartmentalized Self-Tagging for In Vitro-Directed Evolution of XNA Polymerases, Current Protocols in Nucleic Acid Chemistry 57 (2014): 9.9.1-9.9.18.
- Ghadessy FJ, Ong JL, Holliger P. Directed evolution of polymerase function by compartmentalized self-replication, Proceedings of the National Academy of Sciences of the United States of America 98 (2001): 4552-4557.
- Tawfik DS, Griffiths AD. Man-made cell-like compartments for molecular evolution, Nature Biotechnology 16 (1998): 652-656.
- Raia P, Delarue M, Sauguet L. An updated structural classification of replicative DNA polymerases, Biochemical Society Transactions 47 (2019): 239-249.
- Ong JL, Loakes D, Jaroslawski S, et al. Directed evolution of DNA polymerase, RNA polymerase and reverse transcriptase activity in a single polypeptide, Journal of Molecular Biology 361 (2006): 537-550.
- Singh I, Laos R, Hoshika S, et al. Snapshots of an evolved DNA polymerase pre- and post-incorporation of an unnatural nucleotide, Nucleic Acids Research 46 (2018): 7977-7988.
- Ghadessy FJ, Ramsay N, Boudsocq F, et al. Generic expansion of the substrate spectrum of a DNA polymerase by directed evolution, Nature Biotechnology 22 (2004): 755-759.
- Chen F, Gauchera EA, Leal NA, et al. Reconstructed evolutionary adaptive paths give polymerases accepting reversible terminators for sequencing and SNP detection, Proceedings of the National Academy of Sciences of the United States of America 107 (2010): 1948-1953.
- Laos R, Shaw R, Leal NA, et al. Directed evolution of polymerases to accept nucleotides with nonstandard hydrogen bond patterns, Biochemistry 52 (2013): 5288-5294.
- Houlihan G, Arangundy-Franklin S, Porebski BT, et al. Discovery and evolution of RNA and XNA reverse transcriptase function and fidelity, Nature Chemistry 12 (2020): 683-690.
- Percze K, Mészáros T. Analysis of Modified Nucleotide Aptamer Library Generated by Thermophilic DNA Polymerases, ChemBioChem 21 (2020): 2939-2944.
- Eggert F, Kurscheidt K, Hoffmann E, et al. Towards Reverse Transcription with an Expanded Genetic Alphabet, Chembiochem: A European Journal of Chemical Biology 20 (2019): 1642-1645.
- Zhang Y, Ptacin JL, Fischer EC, et al. A semi-synthetic organism that stores and retrieves increased genetic information, Nature 551 (2017): 644-647.
- Kawai R, Kimoto M, Ikeda S, et al. Site-specific fluorescent labeling of RNA molecules by specific transcription using unnatural base pairs, Journal of the American Chemical Society 127 (2005): 17286-17295.
- Moriyama K, Kimoto M, Mitsui T, et al. Site-specific biotinylation of RNA molecules by transcription using unnatural base pairs, Nucleic Acids Research 33 (2005): 1-8.
- Eggert F, Kulikov K, Domnick C, et al. Iluminated by foreign letters – Strategies for site-specific cyclopropene modification of large functional RNAs via in vitro transcription, Methods 120 (2017): 17-27.
- Lee WM, Grindle K, Pappas T, et al. High-Throughput, Sensitive, and Accurate Multiplex PCR-Microsphere Flow Cytometry System for Large-Scale Comprehensive Detection of Respiratory Viruses, Journal of Clinical Microbiology 45 (2007): 2626.
- Sharma N, Hoshika S, Hutter D, et al. Recombinase-based isothermal amplification of nucleic acids with self-avoiding molecular recognition systems (SAMRS), Chembiochem: A European Journal of Chemical Biology 15 (2014): 2268-2274.
- Glushakova LG, Sharma N, Hoshika S, et al. Detecting respiratory viral RNA using expanded genetic alphabets and self-avoiding DNA, Analytical Biochemistry 489 (2015): 62-72.
- Kimoto M, Yamashige R, Matsunaga KI, et al. Generation of high-affinity DNA aptamers using an expanded genetic alphabet, Nature Biotechnology 31 (2013): 453-457.
- Kimoto M, Matsunaga KI, Hirao I. DNA Aptamer Generation by Genetic Alphabet Expansion SELEX (ExSELEX) Using an Unnatural Base Pair System, Methods in Molecular Biology (Clifton, N.J.) 1380 (2016): 47-60.
- Kimoto M, Matsunaga KI, Hirao I. Evolving Aptamers with Unnatural Base Pairs, Current Protocols in Chemical Biology 9 (2017): 315-339.
- Matsunaga KI, Kimoto M, Hirao I. High-Affinity DNA Aptamer Generation Targeting von Willebrand Factor A1-Domain by Genetic Alphabet Expansion for Systematic Evolution of Ligands by Exponential Enrichment Using Two Types of Libraries Composed of Five Different Bases, Journal of the American Chemical Society 139 (2017): 324-334.
- Futami K, Kimoto M, Lim YWS, et al. Genetic Alphabet Expansion Provides Versatile Specificities and Activities of Unnatural-Base DNA Aptamers Targeting Cancer Cells, Molecular Therapy. Nucleic Acids 14 (2019): 158-170.
- Hirao I, Kimoto M, Lee KH. DNA aptamer generation by ExSELEX using genetic alphabet expansion with a mini-hairpin DNA stabilization method, Biochimie 145 (2018): 15-21.
- Matsunaga KI, Kimoto M, Hanson C, et al. Architecture of high-affinity unnatural-base DNA aptamers toward pharmaceutical applications, Scientific Reports 5 (2015): 1-7.
- Kimoto M, Shermane Lim YW, Hirao I. Molecular affinity rulers: systematic evaluation of DNA aptamers for their applicabilities in ELISA, Nucleic Acids Research 47 (2019): 8362-8374.
- Zhang L, Wang S, Yang Z, et al. An Aptamer-Nanotrain Assembled from Six-Letter DNA Delivers Doxorubicin Selectively to Liver Cancer Cells, Angewandte Chemie 132 (2020): 673-678.
- Zumrut HE, Ara MN, Fraile M, et al. Ligand-Guided Selection of Target-Specific Aptamers: A Screening Technology for Identifying Specific Aptamers Against Cell-Surface Proteins, Nucleic Acid Therapeutics 26 (2016): 190-198.
- Zumrut H, Yang Z, Williams N, et al. Ligand-Guided Selection with Artificially Expanded Genetic Information Systems against TCR-CD3€, Biochemistry 59 (2020): 552-562.
- Endo M, Mitsui T, Okuni T, et al. Unnatural base pairs mediate the site-specific incorporation of an unnatural hydrophobic component into RNA transcripts, Bioorganic & Medicinal Chemistry Letters 14 (2004): 2593-2596.
- Kawai R, Kimoto M, Mitsui T, et al. Site-specific fluorescent labeling of RNA by a base-pair expanded transcription system, Nucleic Acids Symposium Series (2004): 35-36.
- Kimoto M, Mitsui T, Harada Y, et al. Fluorescent probing for RNA molecules by an unnatural base-pair system, Nucleic Acids Research 35 (2007): 5360.
- Morohashi N, Kimoto M, Sato A, et al. Site-specific incorporation of functional components into RNA by an unnatural base pair transcription system, Molecules (Basel, Switzerland) 17 (2012): 855-2876.
- Someya T, Ando A, Kimoto M, et al. Site-specific labeling of RNA by combining genetic alphabet expansion transcription and copper-free click chemistry, Nucleic Acids Research 43 (2015): 6665-6676.
- Lavergne T, Lamichhane R, Malyshev DA, et al. FRET Characterization of Complex Conformational Changes in a Large 16S Ribosomal RNA Fragment Site-Specifically Labeled Using Unnatural Base Pairs, ACS Chemical Biology 11 (2016): 1347-1353.
- Seo YJ, Hwang GT, Ordoukhanian P, et al. Optimization of an unnatural base pair toward natural-like replication, Journal of the American Chemical Society 131 (2009): 3246-3252.
- Domnick C, Eggert F, Wuebben C, et al. EPR Distance Measurements on Long Non-coding RNAs Empowered by Genetic Alphabet Expansion Transcription, Angewandte Chemie International Edition 59 (2020): 7891-7896.
- Domnick C, Eggert F, Kath-Schorr S. Site-specific enzymatic introduction of a norbornene modified unnatural base into RNA and application in post-transcriptional labeling, Chemical Communications 51 (2015): 8253-8256.
- Lee HC, Kim JH, Kim JS, et al. Fermentative production of thymidine by a metabolically engineered Escherichia coli strain, Applied and Environmental Microbiology 75 (2009): 2423-2432.
- Chen F, Zhang Y, Daugherty AB, et al. Biological phosphorylation of an Unnatural Base Pair (UBP) using a Drosophila melanogaster deoxynucleoside kinase (DmdNK) mutant, PLOS ONE 12 (2017): e0174163.
- Zhang Y, Lamb BM, Feldman AW, et al. A semisynthetic organism engineered for the stable expansion of the genetic alphabet. Proc Natl Acad Sci U S A 114 (2017): 1317-1322.
- Mauro VP, Chappell SA. A critical analysis of codon optimization in human therapeutics. Trends Mol Med 20 (2014): 604-613.
- Elena C, Ravasi P, Castelli ME, et al. Expression of codon optimized genes in microbial systems: current industrial applications and perspectives. Front Microbiol 5 (2014): 21.
- Manandhar M, Chun E, Romesberg FE. Genetic Code Expansion: Inception, Development, Commercialization . Journal of the American Chemical Society (2021).
- Kimoto M, Kawai R, Mitsui T, et al. An unnatural base pair system for efficient PCR amplification and functionalization of DNA molecules. Nucleic Acids Res 37 (2009): e14.
- Hirao I, Kimoto M. Unnatural base pair systems toward the expansion of the genetic alphabet in the central dogma. Proc Jpn Acad Ser B Phys Biol Sci 88 (2012): 345-367.
- Leitner WW, Ying H, Restifo NP. DNA and RNA-based vaccines: principles, progress and prospects. Vaccine 18 (1999): 765-777.
- Pardi N, Hogan MJ, Porter FW, et al. mRNA vaccines - a new era in vaccinology. Nat Rev Drug Discov 17 (2018): 261-279.
- Berber B, Aydin C, Kocabas F, et al. Gene editing and RNAi approaches for COVID-19 diagnostics and therapeutics. Gene Ther 28 (2021): 290-305.
- Duffy K, Arangundy-Franklin S, Holliger P. Modified nucleic acids: replication, evolution, and next-generation therapeutics. BMC Biol 18 (2020): 112.
- Jacob O, Heyam H, Yara Z, et al. (2018). Overview of MicroRNA Biogenesis, Mechanisms of Actions, and Circulation. Frontiers in Endocrinology 9 (2018): 402.
- Saiyed AN, Vasavada AR, Johar SRK. Recent trends in miRNA therapeutics and the application of plant miRNA for prevention and treatment of human diseases. Futur J Pharm Sci 24 (2022).


 Impact Factor: * 5.8
Impact Factor: * 5.8 Acceptance Rate: 71.20%
Acceptance Rate: 71.20%  Time to first decision: 10.4 days
Time to first decision: 10.4 days  Time from article received to acceptance: 2-3 weeks
Time from article received to acceptance: 2-3 weeks 