COVID19 Prevalence in Germany – Chances and Challenges
Übner T, Bohn U, Hartmann M, Schragner K, Nestle E, Arends H, Eitelhuber A, Krieger F, Diez G*
Institut Virion\Serion GmbH, Würzburg, Germany
*Corresponding author: Diez G, Institut Virion\Serion GmbH, Friedrich-Bergius-Ring 19, 97076 Würzburg, Germany
Received: 26 March 2021; Accepted: 07 April 2021; Published: 28 April 2021
Article Information
Citation: Übner T, Bohn U, Hartmann M, Schragner K, Nestle E, Arends H, Eitelhuber A, Krieger F, Diez G. COVID19 Prevalence in Germany – Chances and Challenges. Archives of Clinical and Biomedical Research 5 (2021): 285-293.
View / Download Pdf Share at FacebookAbstract
Even one year after declaring a pandemic, the number of infections is still increasing. In order to prevent the further spread of SARS-CoV-2, public health interventions have been introduced globally. One major target is to avoid the overload of health care systems, especially for the most severely affected patients. According to the Robert-Koch-Institut (RKI), approximately 2.4 Mio people were infected with SARS-CoV-2 in Germany by the end of February 2021. This represents 2.92% of the German population. However, asymptomatic courses are underestimated. Studies so far focused mainly on identifying acute infected individuals as well as the clinical course of patients and vaccine development. As a consequence, in Germany, as in many other regions knowledge about the seroprevalence of SARS-CoV-2 and thus about the immune status of the population is lacking. Nevertheless, such data is urgently needed to understand the dynamics of the pandemic and hence to design interventions such as vaccination campaigns appropriately. In this work the anti-SARS-CoV-2-IgG immune status of 140 healthy blood donors from southern Germany, collected at the end of February 2021, was determined using the SERION ELISA agile SARS-CoV-2 IgG. In this cohort, 5.7 % were evaluated positive for SARS-CoV-2 specific IgG antibodies, suggesting a higher prevalence as expected.
Keywords
<p>SARS-CoV-2, SERION ELISA agile SARS-CoV-2, Prevalence, Spikeprotein, Nucleoprotein, Magnetic beads, SERION pro-BIND</p>
Article Details
1. Introduction
In December 2019, a new coronavirus (CoV) emerged to cause an acute respiratory disease known as coronavirus disease 19 (COVID-19). The virus was identified to be a betacoronavirus related to severe acute respiratory syndrome coronavirus (SARS-CoV) and thus was named SARS-CoV-2. The virus spread worldwide within 1 month after the first identification, and can be transmitted via aerosol related human-to-human contact. As of March 14th 2020, the virus has infected over 130,000 individuals in 122 countries, approximately 3% of which had a fatal outcome. Due to the rapid rise in number of cases and uncontrolled and vast worldwide spread, the WHO has officially declared SARS-CoV-2 a pandemic. Even twelve month after declaring a pandemic, the number of infections is not decreasing [1-3].
SARS-CoV-2 belongs to the Coronaviridae family [2, 4]. The first two-thirds of the positive sense single-stranded RNA genome contain non-structural proteins (NSP). The NSPs form the main part of the transcription/ replication machinery, including the RNA polymerase [4]. The last third of the genome mainly encodes the four structural proteins: nucleo- (N), membrane- (M), envelope- (E) and spike protein (S) [2, 4, 5]. While the N protein is associated with the RNA genome, the S, E and M proteins together form the virus envelope. Generally, the S protein is functionally divided into S1 with its receptor binding domain (RBD), responsible for the contact with the host cell, and the S2 domain, responsible for cell membrane fusion [2, 5]. Similar to other CoVs, SARS-CoV-2 also uses Angiotensin converting enzyme II (ACE2) as an entry receptor [2]. Among the four coronavirus structural proteins, N and S protein are major immunogenic proteins and are very well suited for detection of anti-SARS-CoV-2 antibodies [3, 5]. The major route of transmission of COVID-19 is droplet and close contact [2, 4]. The mean incubation period is about 5 days, ranging from 1-14 days. Symptoms such as fever, dry cough, breathing difficulties, headache, pneumonia or loss of smell and taste are observed to varying degrees in many patients. The course of the disease varies greatly in its symptoms and severity, ranging from asymptomatic progression (up to 80% of the infections) to severe pneumonia with lung failure and death [1-3, 6]. Therapy currently consists mainly of supportive measures, although great efforts have been made in the development of antiviral drugs. First vaccines were approved in December 2020 [7-10].
During the first week after onset of symptoms, qRT-PCR is used as a reliable method for detecting SARS-CoV-2 infection and as the infection progresses, the combination of qRT-PCR and antibody tests is optimal for accurate diagnosis [3, 11-14]. After an infection, specific IgG, IgA and IgM antibodies can be found [13-15]. Among the acute phase marker IgM and IgA, specific IgA antibodies may persist longer in comparison to IgM [15]. The sole use of antibody detection is particularly important in the later stages of infection, when the virus has already been eliminated by the hosts immune response [16]. Furthermore, since several spike based vaccines are now available, antibody tests can be used to determine the immunestatus before and after immunization [11-14]. In addition, antibody tests are suitable for epidemiological purposes to identify individuals who have developed immunity after infection that may protect against subsequent reinfection or who may be considered as potential plasma donors for therapeutic purposes [17]. One year after declaring a world wide emergency, the number of infections remain on a high level. According to the Robert-Koch-Institut (RKI), approximately 2.4 Mio people were infected with SARS-CoV-2 in Germany by the end of February 2021.
In this work the immune status of 140 healthy blood donors from southern Germany without any preselection, collected at the end of February 2021 from the Bavarian red cross, were screened for the presence of anti-SARS-CoV-2 IgG using the SERION ELISA agile SARS-CoV-2 IgG (Institut Virion/Serion GmbH). To further confirm wether seropositive results were due to natural infection and not caused by prior vaccination a multiplex bead-based assay (Institut Vrion/Serion GmbH) with nucleoprotein and whole spike antigen was executed. Further anti-SARS-CoV-2 IgA and IgM was determined to clarify wether recent exposure to the pathogen was repsonsible for the IgG titer, using SERION ELISA agile IgA and SERION ELISA agile IgM assay. Eight samples were tested positive for SARS-CoV-2 specific IgG antibodies due to natural infection, representing a positive rate of 5.7%. This suggests a higher prevalence as officially determined.
2. Material and Methods
2.1 Anti-SARS-CoV-2 antibody ELISA
SERION ELISA agile SARS-COV-2 IgG, SERION ELISA agile SARS-COV-2 IgA, SERION ELISA agile SARS-COV-2 IgM (Institut Virion-Serion GmbH, Würzburg, Germany) All tests were conducted strictly following the IFU. The classification of the results were excecuted as follows; in case of the quantitative IgG & IgA assays, all borderline/gray zone results were counted as reactive (reactive >10U/ml), for the IgM assys the
ratio from individual serum activity (s) and cut-off
control (co) was determined.
2.2 Blood donors
All plasma samples were provided by the Bavarian red cross (Wiesentheid). The 136 pre-COVID-19 samples were purchased in December 2019 (calendar week 50). The 140 samples representing the blood donors during the pandemic were collected February 2021 (calender week 8).
2.3 Antigens
The recombinant “SARS-CoV-2 Spike Ectodomain (S1-S2)” (BA400R03) used in this study contains the ectodomain of the SARS-CoV-2 Spike glycoprotein including S1, S2 and the RBD. The recombinant “SARS-CoV-2 Nucleoprotein” (BA400R04) contains the full-length sequence of the SARS-CoV-2 Nucleoprotein. The sequences of BA400R03 and BA400R04 are based on the reference Wuhan-Hu-1 strain and were expressed in insect cells.
2.4 Multiplex bead-based SARS-CoV-2 immunoassay
2.4.1 Coupling of antigens to magnetic beads: Proprietary recombinant antigens SARS-CoV2 Nucleoprotein (BA400R04) and Spike (S1-S2)-Ectodomain (BA400R03) were coupled on proprietary magnetic amino microparticles approximately 5 µm in size with different intrinsic fluorescent levels. Coupling was performed automatically in 96well plates (Maelstrom 8 Autostage, TANBead, Taiwan) at room temperature. For each antigen 4.7 x 106 beads per microplate well were washed in 100µL washing buffer (50 mM MES, 0.02% Tween® 20, pH 7.4), thereafter prepared for coupling by 20 minutes incubation in 100µL per well of a proprietary metal chelate coordinating buffer (SERION pro-BIND, CRC01). After a second washing step, antigens diluted in coupling buffer (50 mM MES, pH 6.1) were irreversibly immobilized on bead surface by 60 minutes incubation time. After another washing step, beads were blocked in 100 µL per well blocking buffer (10 mg/mL Blocking Reagent (Roche, 11 112 589 001), 25 mM TRIS, 100 mM NaCl, 0.09% NaN3) for 30 minutes, washed again and transferred into storage buffer (StabilCoat™ Plus Stabilizer (Surmodics, SC02-0050)).
2.4.2 Specific antibody detection: The immunoassay was performed automatically with bead handling (KingFisher Flex, Thermo Scientific) in 96 well microtiter plates: Nucleoprotein- and spike-beads were vortexed, sonicated and 2000 beads of each bead set were added per microplate well in 100µL storage buffer. Beads were transferred into 120µL per well of diluted samples: For IgA and IgM assay, samples were diluted 241-fold in dilution buffer 1 (Virion\Serion, B231), in case of IgG detection, sample dilution was 481-fold in dilution buffer 2 (Virion\Serion, B431). After 20 minutes of incubation at 37°C, beads were washed in 200µL per well washing solution (Virion\Serion, B232). Binding of specific antibodies was confirmed using 50µL per well of 900 fold diluted R-Phycoerithrin-conjugated AffiniPure F(ab’)2 goat anti-human IgG (Jackson ImmunoResearch Lab. Code 109-116-098), 2001 fold diluted R-Phycoerithrin-conjugated AffiniPure F(ab’)2 Fragment Donkey Anti-Human IgM (Jackson ImmunoResearch Lab. Code 709-116-073), or 75 fold diluted Fluorescein (FITC)-conjugated AffiniPure F(ab’)2 goat-anti human IgA (Jackson ImmunoResearch Lab. Code 109-096-0119) in conjugate buffer (50mM NaH2PO4 H2O, 150mM NaCl, 0,4% BSA, 0.09% NaN3, 0.02% Tween® 20 pH 7.4 (Virion\Serion, B266). Unbound conjugate was removed by another washing step after 20 minutes incubation at 37°C. Finally beads were transferred into conjugate-buffer.
A flow cytometer (CytoFLEX S, Beckman Coulter) was used for readout, the assay signal is expressed in median fluorescence intensity (MFI). Different quantities of intrinsic fluorescent dye allows distinction of both bead-populations (red laser/ filter 712/25). Bound specific IgA antibodies were detected by FITC-Signal (blue laser /filter 525/40), IgG and IgM antibodies by R-PE-Signal respectively (yellow laser/ filter 585/42).
3. Results
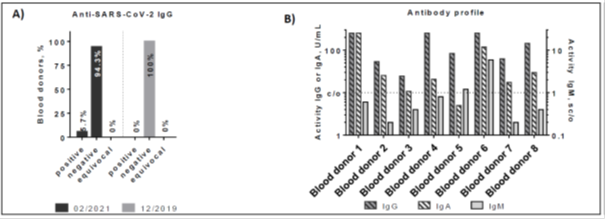
Anti-SARS-CoV-2 IgG reactivity from 140 plasma samples from healthy individuals obtained at the end of February 2021 were determined using the SERION ELISA agile SARS-CoV-2 IgG assay (see Figure 1). Out of 140 individuals, eight revealed a positive IgG titer (5.7%). In comparison, none of the 136 blood-donors, obtained in December 2019 gave a positive result for IgG, demonstrating a 100% naïve German population before 2020. In order to verify whether the IgG titer was generated as a consequence of recent exposure to SARS-CoV-2 components, all IgG-positive blood donors were also examined for SARS-CoV-2 specific IgA and IgM antibodies. Of eight IgG positive samples, six revealed a positive and one showed an equivocal result in the SERION ELISA agile SARS-CoV-2 IgA test. In addition, two samples were also rated positive in the SERION ELISA agile SARS-CoV-2 IgM. Therefore, each IgG titer correlates with an IgA- or IgM-specific acute titer, suggesting that the IgG antibody response occurred due to recent exposure to SARS-CoV-2.
Even though a vaccine has been available since December 2020, the number of infections are still increasing. According to the RKI approximately 2.4 million people in Germany had been infected by the end of February 2021 (see also Table 1). This is approximately 2.92% of the German population. In the Bavarian population, as many as 3.31% were tested positive by the end of February 2021 (see also Table 1). This fact, together with the in Figure 1 determined positive rate of 5.7%, suggests a higher incidence than actually calculated.
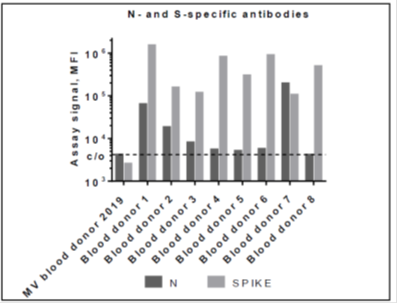
Since vaccination programs have already started, and current approved vaccines encode the full-length SARS-CoV-2 S protein a positive anti-SARS-CoV-2 S protein IgG titer may also be due to prior immunization. In order to discriminate whether the positive SARS-CoV-2 specific IgG titer is due to natural infection, the antibody response to the SARS-CoV-2 N protein, which is not detectable after vaccination, is determined. For this approach, a bead-based multiplex assay with two SARS-CoV-2 antigens (whole spike and N protein) is used. (see also Figure 2). Consistent with the SERION ELISA agile SARS-CoV-2 IgG results, all eight samples showed specific reactivity with the whole spike antigen coated particle. Seven of the eight healthy blood donors tested positive for SARS-CoV-2 IgG in the SERION ELISA agile SARS-CoV-2 IgG also showed N specific IgG reactivity. All signals were above the cut-off determined by the mean value of healthy pre-COVID-19 blood donors.
Figure 1: Reactivity of anti-SARS-CoV-2 specific antibodies in healthy blood donors. A) Anti-SARS-CoV-2 S protein IgG reactivity in plasma samples from healthy blood donors collected in February 2021 and December 2019. B) Anti-SARS-CoV-2 IgG, IgM and IgA reactivity of eight individuals identified as seropositive in Figure 1A. Threshold: IgG and IgA: >10 U/ml = positive Threshold IgM: >0.9 = positive.
|
2021-02-25 |
Germany |
Bavaria |
|
Population |
83.2 Mio |
13.1 Mio |
|
COVID-19 cases (n) |
2.427.615 |
447.230 |
|
COVID-19 cases (%) |
2.92 |
3.31 |
Table 1: SARS-CoV-2 positively tested individuals; Source RKI.
4. Discussion
Anti-SARS-CoV-2 serological assays are urgently needed for contact tracing, epidemiological and vaccine evaluation studies. The N and the S proteins are the main immunogenic CoV proteins. While N proteins are prone to cross-reactivity, S protein-specific antibodies correlate strongly with virus-neutralizing antibodies [1, 3, 5]. Since the majority of the human population has antibodies against endemic human CoV, it is crucial to verify the specificity of these assays to avoid false-positive results. Additionally, the two zoonotic CoV, SARS-CoV and MERS-CoV, are also beta-coronaviruses, increasing the potential for cross-reactivity. For the SERION ELISA agile SARS-CoV-2 IgG the whole spike protein is used exclusively to achieve a very specific detection and a high correlation with protective neutralizing antibodies that are mainly produced against the S protein. Since none of the blood donors in the pre-pandemic panel from December 2019 showed a positive result in the SERION ELISA agile SARS-CoV-2 IgG, cross reactivity is reduced to a minimum, suggesting a highly specific serological assay. This was recently verified [18]. Strömer and co-workers demonstrates a high specificity of >99% and a sensitivity of 96.2% for the test used. Furthermore, it can be assumed, that the determined antibody titers are due to the exposure of the individuals to SARS-CoV-2 pathogens and not caused by vaccination. Almost all anti-SARS-CoV-2 IgG positive individuals show antibodies directed against S and N proteins (Figure 2). All vaccines currently available on the market use S protein exclusively for immunization [11-14]. In that case, only an anti-S-specific titer would be expected. In contrast to the anti-S-IgG concentration, the N specific IgG titers are moderate in most cases. This might be due to the fact, that SARS-CoV-2 N specific IgGs disapear soon after the infection [18]. Supportive for the hypothesis of natural infections are the additional IgA titers of the individuals. It is known, that intramuscular vaccination does not induce specific IgA or IgM titer in general. Conventional injected vaccines usually induce specific T-cell responses in the bloodstream and serum IgG reactivity [19]. In respiratory infections, IgA titers arise from mucosal contact with the pathogen, suggesting an IgA titer due to infection rather than vaccination.
According to the RKI nearly 2.5 Mio infections were registered in Germany by the end of February 2021. This represents approximately 3% of the entire German society. However, the calculated incidence does not reflect the determined prevalence of this work. That rate is almost twice as high. This finding is in accordance with other studies, investigating the prevalence [20, 21]. In a small German town, the seroepidemiological study demonstrated a 5-fold higher infection rate than calculated on base of the RT-PCR tests [20]. Preliminary results from the KoCo19 study also confirm this finding. Here, the specific IgG level of more than 5.000 inhabitants was determined. The positive rate was 4-fold higher than expected [21]. This might be caused by asymptomatic courses of the disease [2, 12]. Serological assays are urgently needed to supplement the diagnostic repertoire in identifying patients with past SARS-CoV-2 infection. SARS-CoV-2 specific antibodies are detectable more than one week after onset of symptoms, limiting the role of serology for identification of acute infection [1, 3, 22]. But serological assays might allow for the detection of patients during a later stage of the disease when viral clearance may precede the disappearance of symptoms. Furthermore, neutralizing antibodies are known to be produced after infection, indicating that reconvalescent individuals have developed immunity against SARS-CoV-2 [23, 24]. More than 90% of anti-SARS-CoV-2 specific neutralizing antibodies are directed against the S protein. Vaccines based on the immunogenically active S protein are available, but production capacity is limited [11-14]. Anti-S-specific IgG antibodies are detectable after immunization as well as after infections with SARS-CoV-2 [1, 3, 25, 26]. It might be helpful to distinguish reconvalescent individuals from the serological naïve by specific IgG determination. Immunization is not necessarily needed for that reconvalescent group, so that vaccines might be applied more efficiently in non-immune individuals. For that purpose S-specific serological assays could be helpful and guarantee the efficient application of vaccines in serologically naive populations. In addition, serological assays, based on the S protein could also be helpful to identify potential plasma donors expressing neutralizing antibodies for therapeutic transfusion [22, 27].
Acknowledgements
Thank you to C. Loewel and Dr. T. Schumacher for fruitful and constructive discussions.
Conflicts of Interest
None, all authors are affiliated with the Institut VIRION\ SERION GmbH.
References
- Peeri NC, Shrestha N, Zaki R, et al. The SARS, MERS and Novel Coronavirus (COVID-19) Epidemics, the Newest and Biggest Global Health Threats: What Lessons Have We Learned?. International Journal of Epidemiology 49 (2020): 717-726.
- He F, Deng Y, Li W. Coronavirus Disease 2019 (COVID-19): What We Know?. Journal of Medical Virology (2020).
- Nisreen MA Okba, Marcel A Muller, Wentao Li, et al. SARS-CoV-2 Specific Antibody Responses in COVID-19 Patients. preprint (Infectious Diseases (except HIV/AIDS) (2020).
- Shanmugaraj B, Malla A, Phoolcharoen W, et al. Emergence of Novel Coronavirus 2019- nCoV: Need for Rapid Vaccine and Biologics Development. Pathogens 9 (2020): 148.
- Meyer B, Drosten C, and Müller M. Serological Assays for Emerging Coronaviruses: Challenges and Pitfalls. Virus Research 194 (2014): 175-183.
- Zhou P Yang XL, Li shi Z. A Pneumonia Outbreak Associated with a New Coronavirus of Probable Bat Origin. Nature 579 (2020): 270-273.
- Fernando P Polack, Stephen J Thomas, Nicholas K, et al. Safety and Efficacy of the BNT162b2 mRNA Covid-19 Vaccine. n engl j med 383 (2020): 2603-2615.
- Ugur Sahin, Alexander Muik, Ozlem Tureci, et al. COVID-19 vaccine BNT162b1 elicits human antibody and TH1 T cell responses. Nature 586 (2020): 594-599.
- Merryn Voysey, Sue Ann Costa Clemns, Shabir A Madhi, et al. Safety and efficacy of the ChAdOx1 nCoV-19 vaccine (AZD1222) against SARS-CoV-2: an interim analysis of four randomised controlled trials in Brazil, South Africa, and the UK. Lancet 397 (2021): 99-111.
- Baden L R, Frey S, Diemert D, et al. Efficacy and Safety of the mRNA-1273 SARS-CoV-2 Vaccine. N Engl J Med 384 (2021): 403-416.
- To KK, Tsang OT, Leung WS, et al. Articles Temporal profiles of viral load in posterior oropharyngeal saliva samples and serum antibody responses during infection by SARS- CoV-2: an observational cohort study. Lancet Infect Dis 20 (2020): 565-574.
- Jia X, Zhang P, Tian Y, et al. Clinical Significance of IgM and IgG Test for Diagnosis of Highly Suspected COVID-19 Infection. preprint (Infectious Diseases (except HIV/AIDS) (2020).
- Guo L, Ren L, Yang S, et al. Profiling Early Humoral Response to Diagnose Novel Coronavirus Disease (COVID-19). Clinical Infectious Diseases 71 (2020): 778-785.
- Amanat F, Stadlbauer D, Krammer F, et al. A Serological Assay to Detect SARS-CoV-2 Seroconversion in Humans. preprint Allergy and Immunology 26 (2020): 1033-1036.
- Andrea Padoan, Sciacovelli L, Basso D, et al. IgA-Ab response to spike glycoprotein of SARS-CoV-2 in patients with COVID-19: A longitudinal study. Clinica Chimica Acta 507 (2020): 164-166.
- Zou L, Ruan F, Huang M, et al. SARS-CoV-2 Viral Load in Upper Respiratory Specimens of Infected Patients. N Engl J 382 (2020): 1177-1179.
- Bao L, Deng W, Gao H, et al. Reinfection could not occur in SARS-CoV-2 infected rhesus macaques (In Press).
- Anabelle Strömer, Rose R, Grobe O, et al. Kinetics of Nucleo- and Spike Protein- Specific Immunoglobulin G and of Virus- Neutralizing Antibodies after SARS-CoV-2 Infection. Microorganisms 8 (2020): 1572.
- Prosper N. Boyaka. Inducing Mucosal IgA: A Challenge for Vaccine Adjuvants and delivery Systems. J Immunol 199 (2017): 9-16.
- Hendrik Streeck, Schulte B, Beate M, et al. Infection fatality rate of SARS-CoV-2 infection in a German community with a super-spreading event. medRxiv (2020).
- Katja Radon, Saathoff E, Prits.ch M, et al. Protocol of a population-based prospective COVID-19 cohort study Munich, Germany (KoCo19). BMC 20 (2020): 1036.
- Alexander Krüttgen, Dreher M, Hornef M, et al. Comparison of four new commercial serologic assays for determination of SARS- CoV-2 IgG. Journal of Clinical Virology 128 (2020): 104394.
- Takuya Tada, Belinda M, Ramin S, et al. Neutralization of viruses with European, South African, and United States SARS-CoV- 2 variant spike proteins by convalescent sera and BNT162b2 mRNA vaccine-elicited antibodies. bioRxiv preprint (2021).
- Pengfei Wang, Manoj S Nair, Liu L, et al. Antibody Resistance of SARS-CoV-2 Variants 5 B.1.351 and B.1.1.7. bioRxiv preprint (2021).
- Mark J Mulligan, Kirsten E Lyke, Nicholas Kitchin, et al. Phase I/II study of COVID-19 RNA vaccine BNT162b1 in adults; Nature 8 (2020): 1572.
- Sadoff J, Shukarev G, Dirk Heerwegh, et al. Interim Results of a Phase 1-2a Trial of Ad26.COV2. S Covid-19 Vaccine. N Engl J Med 21 (2021).
- Graham C, Seow J, Huettner I, et al. Impact of the B.1.1.7 variant on neutralizing monoclonal antibodies recognizing diverse epitopes on SARS-CoV-2. bioRxiv (2021).


 Impact Factor: * 5.8
Impact Factor: * 5.8 Acceptance Rate: 71.20%
Acceptance Rate: 71.20%  Time to first decision: 10.4 days
Time to first decision: 10.4 days  Time from article received to acceptance: 2-3 weeks
Time from article received to acceptance: 2-3 weeks 