Antibiotic Resistance and Phenotypic and Genotypic Detection of Colistin Resistance among Enterobacter Species Isolated from Patients of a Tertiary Care Hospital, Bangladesh
Nazmun Nahar Munny1,*, S.M. Shamsuzzaman2, Tamzeed Hossain3
1Assistant Professor, Department of Microbiology, East West Medical College, Dhaka, Bangladesh
2Professor and Head, Department of Microbiology, Dhaka Medical College, Dhaka, Bangladesh
3Fellowship trainee, Department of Internal Medicine, Dhaka Medical College Hospital, Dhaka, Bangladesh
*Corresponding author: Nazmun Nahar Munny, Assistant Professor, Department of Microbiology, East West Medical College, Dhaka, Bangladesh.
Received: 17 August 2021; Accepted: 26 August 2021; Published: 01 September 2021
Article Information
Citation: Nazmun Nahar Munny, S.M Shamsuzzaman, Tamzeed Hossain. Antibiotic resistance and phenotypic and genotypic detection of colistin resistance among Enterobacter species isolated from patients of a tertiary care hospital, Bangladesh. Archives of Microbiology and Immunology 5 (2021): 337-352.
View / Download Pdf Share at FacebookAbstract
The prevalence of antimicrobial resistance among Enterobacter species is alarmingly high. Colistin as a last resort antibiotic reintroduced in the treatment pipeline to combat the drug resistant Enterobacter infection has led to emergence of colistin resistance. The study was designed to determine the prevalence of antibiotic resistance among isolated Enterobacter and to assess the occurrence of colistin resistance both phenotypically and genotypically in a tertiary care hospital, Bangladesh. This cross-sectional study was conducted during the period July2018 to June 2019. A total 350 different samples were processed. Enterobacter were identified phenotypically and were processed for antimicrobial susceptibility test (AST) by using modified disk diffusion method. Colistin resistance was determined by minimum inhibitory concentration (MIC) using agar dilution method. Colistin resistant strains were further processed for colistin resistance genes using polymerase chain rection (PCR). Among the total samples processed, 65.14% (228/350) yielded culture positive growth of which 12.28% (28/228) were Enterobacter species. Among isolated Enterobacter 22 (9.64%) were E. cloacae and 6 (2.63%) were E. aerogenes. In AST, 89.29%, 57.14% and 42.86% were resistant to ciprofloxacin, amikacin and imipenem respectively. Tigecycline and colistin were most effective antibiotics. Among colistin resistant Enterobacter isolates, 85.71%, 71.42%, 42.85% and 57.14% were positive for PmrC, PmrA, PhoP and PmrB respectively detected by PCR. Our study reports identification and distribution of drug resistant Enterobacter strains in a major tertiary care hospital. High prevalence of drug resistance and emergence of colistin resistance is indicative of deteriorating situation. Therefore, further extensive surveillance is mandatory.
Keywords
<p>Enterobacter; Antimicrobial susceptibility; Colistin resistance; Polymerase chain reaction; Agar dilution method; E.cloacae, E.aerogenes</p>
Article Details
1. Introduction
Enterobacter spp., including Enterobacter cloacae and E. aerogenes, are opportunistic pathogens that cause infections in the blood, respiratory, urinary, and gastrointestinal tracts [1] and are a member of the ESKAPE group of significant bacterial pathogens in humans [2]. Enterobacter is associated with hospital acquired infection that has been ranked as the third most frequent isolate following Escherichia coli and Klebsiella species [3].
These organisms seem to have innate resistance to older anti-microbial agents and have the propensity to rapidly develop resistance to newer anti-microbial agents [4]. Treatment of infections with Enterobacter spp. is difficult and broad resistance to third generation cephalosporins, penicillin and quinolones is an increasing problem [5]. Gram negative bacteria, mainly Enterobacterales, Acinetobacter baumannii and Pseudomonas aeruginosa are able to produce enzymes such as extended-spectrum beta-lactamases (ESBLs), AmpC beta-lactamases, carbapenemase and metallo-beta-lactamases (MBL), which enable the host bacteria to develop resistance to most classes of antibiotics in use [6]. Enterobacter spp. is increasingly associated with multidrug resistance, including the resistance to the last-resort carbapenems [7]. Enterobacter spp. is second most common carbepenem resistant Enterobacteriaceae (CRE) in the United States [8]. The rapid increase of serious infections by resistant Enterobacter necessitates the availability of an alternative antimicrobial agent for treatment [9].
As multidrug resistance and carbapenem resistance increase in Gram-negative pathogens including Enterobacter spp., colistin, a polycationic antimicrobial peptide, is often recommended as a last-line therapeutic option [10]. Although colistin resistance rate is relatively low worldwide, resistance has begun to emerge [11]. In addition to the steady emergence of colistin resistance due to chromosomal mutations, plasmid-borne colistin resistance has recently been reported, which is a great threat to public health [12]. Implementation of effective surveillance programs and infection controls are considered as the two pillars to check the growth and spread of AMR [13]. Accurate and timely information on local and national patterns of organisms causing infection and their drug-resistance profile is therefore paramount for good clinical management and benchmarking [14].
However, few studies have investigated on the colistin resistance of Enterobacter spp. in comparison with studies of other Gram-negative pathogens. Therefore, our study was designed to determine the prevalence of drug resistant strains ofEnterobacter species from clinical isolates to recognize the possible therapeutic options for combating these ICU bugs. At the same time, we also report colistin resistant Enterobacter cloacae and E. aerogenes and presence of colistin resistant gene among them in Bangladesh.
2. Methods
2.1 Study design
This cross- sectional study was conducted from July 2018 to June 2019 in the department of Microbiology, Dhaka Medical College Hospital Bangladesh. Samples was collected from adult patients having clinically suspected infections admitting in Dhaka Medical College Hospital or were received in the Microbiology department for culture and sensitivity after taking informed written consent irrespective of sex and antibiotic intake. Patients who did not give consent were excluded from this study.
2.2 Isolation and Identification of Enterobacter Species
All the samples were aseptically collected following the standard microbiological procedure. A total 350 clinical sample consisting of urine (n=124), wound swab (n=113), endotracheal aspirates (n=58), and blood (n=53) was collected. Clinical samples were cultured on blood agar and MacConkey agar media. Isolates were identified phenotypically by standard microbiological techniques (Colonial morphology, Gram stain, and biochemical test. Genus Enterobacter was identified by characteristics colonies (Lactose-fermenting, sometimes mucoid colonies), Gram staining pattern as Gram negative bacilli, motility as motile, and standard biochemical reactions (catalase, oxidase, indole production, citrate utilization, urease activity, reaction in triple sugar iron medium and lysine decarboxylase test). Common Enterobacter spp (E. cloacae and E. aerogenes) were isolated [15].
2.3 Antimicrobial Susceptibility Test
Susceptibility to antimicrobial agents of all isolated organisms were determined by Kirby-Baurer modified disc diffusion technique using Mueller-Hinton plates and zones of inhibition were interpreted according to CLSI guidelines [16]. The criteria for the United States Food and Drug Administration was used for the interpretation of zone of inhibition of tigecycline. Antibiotic discs were obtained from commercial sources (Oxoid Ltd, UK). Following antimicrobial discs were used: amikacin (30μg), piperacillin-tazobactam (100/10μg), imipenem (10μg), tigecycline (15µg), ciprofloxacin (30μg), cefepime (30μg), ceftazidime (30μg), ceftriaxone (30μg), cefoxitin (30μg), cefuroxime sodium (30μg), amoxiclav (amoxicillin 20μg & clavulanic acid 10μg) and aztreonam (10μg). In this method, broth culture of test bacteria (comparable to McFarland tube no.0.5; inoculums density 1.5× 108 bacteria/ml) was uniformly carpeted on the surface of Mueller Hinton agar (MHA). Twelve, antibiotics discs were placed onto the lawn culture of the test bacteria by sterile forceps. The inoculated and seeded MHA plates were incubated at 37°C for 24h. After incubation, zone of inhibition was measured and results were interpreted as sensitive, intermediate and resistant [17]. Colistin susceptibility was determined by agar dilution method.
2.3.1 Colistin Susceptibility Test
Agar dilution method was used to determine the susceptibility of colistin. As 0.5 McFarland turbidity standards contain 1×108cfu/ml [16] 10 times dilution of test inoculums was done to achieve 1× 107cfu/ml. Two concentrations of colistin 2µg/ml and 4µg/ml were prepared in the agar medium. Bacterial inoculums was applied readily onto the agar surface and the plates were incubated at 37°C up to 24h. Isolates having a MIC of≤2μg/mL is considered colistin susceptible while MIC of>2 μg/mL is considered colistin resistant [18].
2.4 Phenotypic Detection of ESBLs Producers
Using sterile cotton swab, test inoculums (compared with McFarland standard) were inoculated in Mueller-Hinton agar plate. Third generation caphalosporins (ceftriaxone, ceftazidime and cefotaxime) were placed 20 mm apart from center of the amoxiclav disc. The inoculated plate was incubated at 370C for 24 hours. A clear extension of the edge of the inhibition zone of cephalosporin discs towards amoxiclav disc was interpreted at ESBLs production [19].
2.5 Phenotypic Detection of MBL Producers
Confrmation of MBL production was made by inhibition method in which Ethylene Diamine Tetra Acetic Acid (EDTA) was used as an inhibitor. Two imipenem discs were placed on the inoculated Mueller-Hinton agar plate. One imipenem disc was supplemented with 5 µl of 0.5 M EDTA solution (containing approximately 750 µg EDTA) and incubated overnight at 370C. An increased zone of diameter of ≥ 6 mm around the disc containing imipenem supplemented with EDTA compared to the disc containing imipenem only was interpreted as MBL production [20].
2.6 Determination ofMIC ofColistin
Minimum inhibitory concentration of colistin was determined by agar dilution method. Different concentrations of colistin ranging from 2µg/ml to 256µg/ml were prepared in the agar medium. For each plate 50ml Mueller-Hinton media was prepared.50 ml of sterile Mueller-Hinton agar was impregnated with 12.5 µl, 25 µl, 50 µl, 100 µl, 200 µl, 400 µl, 800 µl, 1600 µl of colistin to achieve the concentration 2µg/ml, 4µg/ml, 8µg/ml, 16 µg/ml, 32µg/ml, 64µg/ml, 128 µg/ml and 256 µg/ml per plate respectively. Bacterial inoculum was applied readily onto the agar surface and the plates were incubated at 37°C overnight. The MIC end point was determined as the lowest concentration of antibiotics that completely inhibits the visible growth [21]. Escherichia coli ATCC 25922 was used as control strain [16]. Commercially available colistin injection vial (Forest pharma Limited) was used
2.7 Molecular Method [22]
Polymerase chain reaction (PCR) was done for the detection of colistin resistance genes incolistin resistant Enterobacter species.
2.7.1 Procedure of Bacterial Pellet Formation and DNA Extraction
A loop full of bacterial colonies from MHA media was inoculated into a micro centrifuge tube having sterile TSB and incubated overnight at 37ºC. Incubated tube was centrifuged at 4000g for 10 minutes. Supernatant was discarded and tubes containing bacterial pellets were kept at -20ºC for DNA extraction. Three hundred microlitre of sterile distilled water was added to micro centrifuge tubes having pellets and vortexed until mixed well. Then the mixture was heated at 100° C for 10 minutes in a heat block. After heating, tubes were immediately placed on ice for 5 minutes and centrifuged at 14000 g for 6 minutes at 4ºC. Finally, the supernatant was taken into another micro centrifuge tube. This extracted DNA was preserved at 4ºC for 7-10 days and -20ºC for a long time.
2.7.2 Mixing of Master Mix with Primer and DNA Template
PCR was performed in a final reaction volume 25 µl in a PCR tube, containing 12.5 µl of master mix (mixture of dNTP, taq polymerase, MgCl2and PCR buffer), 2 µl forward primer, 2 µl reverse primer (Promega Corporation, USA, 2 µl of extracted DNA and 6.5 µl of nuclease free water. After a brief vortex, the tubes were centrifuged. The pair of primers were used to yield PCR products depicted in (Table 1).
Table I: Primers used in this study [23]
|
Genes |
Sequence (5’-3’) |
Size (bp) |
|
|
1.PmrA |
F |
CGC AGG ATA ATC TGT TCT CCA |
808 |
|
R |
GGT CCA GGT TTC AGT TGC AA |
||
|
2.PmrB |
F |
GCG AAA AGA TTG GCA AAT CG |
659 |
|
R |
GGA AAT GCT GGT GGT CAT CTG A |
||
|
3.PmrC |
F |
CTC TCG CCT CGT TCT GAA |
140 |
|
R |
CGG AGT GGT GTC GAG GAT A |
||
|
4.mgrB |
F |
ACC ACC TCA AAG AGA AGG CGT T |
347 |
|
R |
GGC GTG ATT TTG ACA CGA ACA C |
||
|
5.PhoP |
F |
GAG CGT CAG ACT ACT ATC GA |
942 |
|
R |
GTT TTC CCA TCT CGC CAG CA |
||
|
6.PhoQ |
F |
CCA CAG GAC GTC ATC ACC A |
1594 |
|
R |
GCA GGT GTC TGA CAG GGA TT |
||
|
7.mcr1 |
F |
CGG TCA GTC CGT TTG TTC |
309 |
|
R |
CTT GGT CGG TCT GTA GGG |
||
|
8.mcr2 |
F |
TGTTGCTTGTGCCGATTGGA |
567 |
|
R |
AGATGGTATTGTTGGTTGCTG |
||
2.7.3 Amplification in Thermal Cycler (Gene Atlas, Master cycler gradient, Japan, Model482)
PCR assays were performed in a DNA thermal cycler. After amplification products were processed for gel documentation or kept at -20ºC till tested.
2.7.4 Agarose gel Electrophoresis and Visualization
PCR products were detected by electrophoresis on 1.5% agarose gel. Gel was prepared with 1 X TBE buffer (Tris EDTA). For 1.5% agarose gel preparation, 0.18 gram agarose powder (LE, analytic grade, Promega, Madison, USA) was mixed with a 1.25 ml TBE buffer. A comb was placed in a gel tray, the gel was poured. After solidification, 1 µl of loading dye and 5 µl of amplicon was mixed on parafilm and was loaded in agarose well. Similarly, 2 µl of 100bp DNA ladder was mixed with 1µl loading dye and was loaded. Gel electrophoresis was done in 230 voltages for 30 minutes. After electrophoresis, the gel was stained with ethidium bromide (20µl ethidium bromide in 200 ml distilled water). The gel was observed under UV transilluminator (Gel Doc, Major Science, Taiwan) for DNA bands. The DNA bands were identified according to their molecular size by comparing with the molecular weight marker (100bp DNA ladder) loaded in a separated lane.
3. Result
Among the total samples processed, 65.14% (228/350) had bacterial growth. Out of 228 culture positive samples, 28 (12.28%) were Enterobacter species of which 22(9.64%) were E. cloacae and 6 (2.63) were E. aerogenes. Among 61 urine samples 11(18.03) were Enterobacter species, among 93 wound swab and pus 9(9.68%) were Enterobacter species (Table II).
Table II: Distribution of isolated Enterobacter cloacae and Enterobacter aerogenes from different culture positive samples (N=228)
|
Samples |
E.cloacae |
E.aerogenes |
|
Urine (N=61) |
8 (13.11) |
3 (4.91) |
|
Wound swab and pus (N=93) |
7 (7.53) |
2 (2.15) |
|
Endotracheal aspirates (N=53) |
6 (11.32) |
1 (1.07) |
|
Blood (N=21) |
1 (4.16) |
0 (0.00) |
|
Total (N=228) |
22 (9.64) |
6 (2.63) |
Among the 28 isolated E. cloacae and E. aerogenes, all were resistant to amoxiclav and cefoxitin, 89.29% were resistant to ciprofloxacin and cefuroxime, 57.14% were resistant to amikacin, 50% were resistant to piperacillin-tazobactam, 42.86% were resistant to imipenem, 25% were resistant colistin and 3.57% were resistant tigecycline (Table III).
Table III: Antibiotic resistance pattern of isolated E. cloacae and E. aerogenes (N=28)
|
Antimicrobial drugs |
E. cloacae N=22 n(%) |
E. aerogenes N=6 n(%) |
Total N=28 n(%) |
|
Amoxiclav |
22 (100.00) |
6 (100.00) |
28 (100.00) |
|
Cefoxitin |
22 (100.00) |
6 (100.00) |
28 (100.00) |
|
Ciprofloxacin |
21 (95.45) |
4 (66.67) |
25 (89.29) |
|
Cefuroxime |
22 (100.00) |
3 (50.00) |
25 (89.29) |
|
Ceftriaxone |
21 (95.45) |
3 (50.00) |
24 (85.71) |
|
Ceftazidime |
19 (86.36) |
3 (50.00) |
22 (78.57) |
|
Aztreonam |
15 (53.57) |
2 (33.33) |
17 (60.71) |
|
Amikacin |
13 (59.09) |
3 (50.00) |
16 (57.14) |
|
Imipenem |
11 (50.00) |
1 (16.67) |
12 (42.86) |
|
Piperacilin-tazobactam |
11 (50.00) |
3(50.00) |
14 (50.00) |
|
Cefepime |
7 (31.82) |
2 (33.33) |
9 (32.14) |
|
*Colistin |
6 (27.27) |
1 (16.67) |
7 (25.00) |
|
Aztreonam |
15 (53.57) |
2 (33.33) |
17 (60.71) |
|
Tigecycline |
1 (4.54) |
0 (0.00) |
1 (3.57) |
N= Total number of isolated bacteria
n =Total number of resistant bacteria
*Colistin resistance was determined by MIC by agar dilution method.
Out of 28 isolated E. cloacae and E. aerogenes, 15 multidrug resistant strains were detected. Of which, 5 (45.45%) were detected from urine, 5 (55.55%) from wound swab, 4 (57.14%) from endotracheal aspirates, one (100%) from blood sample (Table IV).
Table IV: Distribution of multidrug resistant (MDR) E. cloacae and E.aerogenes isolated from different samples
|
Sample |
Total isolates N=28 |
MDR isolates n =15 (%) |
|
Urine |
11 |
5 (45.55) |
|
Wound swab and pus |
9 |
5 (55.55) |
|
ETA |
7 |
4 (57.14) |
|
Blood |
1 |
1 (100.00) |
|
Total |
28 |
15 (53.57) |
ETA= Endotracheal aspirate
N= Total number of isolated E. cloacae and E. aerogenes
n = Total number of MDR isolates.
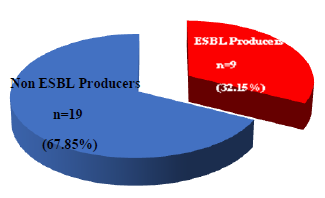
Among isolated Enterobacter cloacae and E. aerogenes 32.15% were ESBLs producer detected by double disc synergy test (Figure 1).
N= Total number of E.cloacae and E.aerogenes.
n= Number of ESBL producers & non ESBL producers.
Table V: Phenotypic detection of carbapenemase producers by combined disc method among imipenem resistant E. cloacae and E. aerogenes (N=12)
|
Organisms |
Carbapenemase producers CD assay n (%) |
|
E. cloacae |
8 (66.67) |
|
E. aerogenes |
1 (8.33) |
|
Total |
9 (75.00) |
N= Total number of imipenem resistant bacteria.
n = Number of carbapenemase producers by phenotypic test
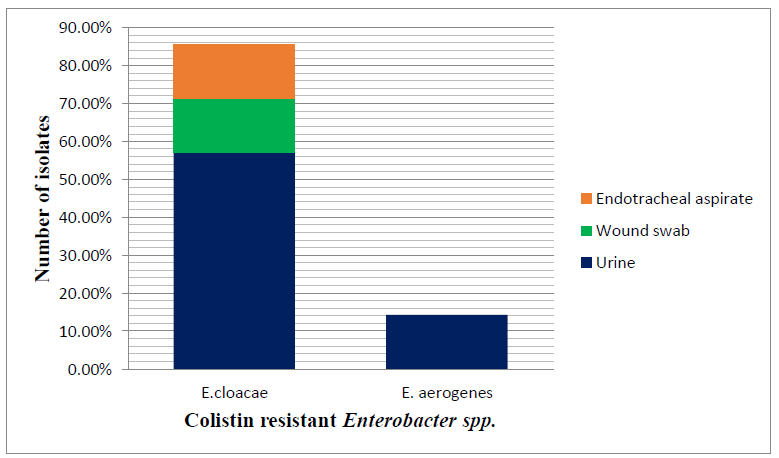
Among the colistin resistant Enterobacter spp., 6 (85.71%) were E.cloacae and 1 (14.28%) were E.aerogenes isolated from different clinical samples (Figure 2)
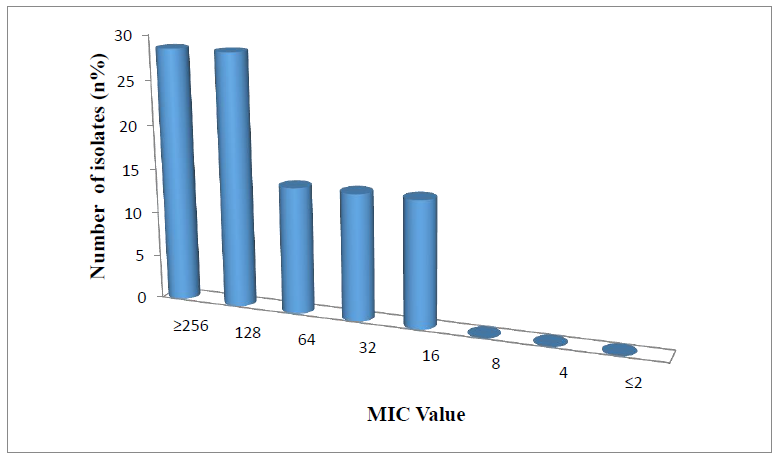
Among 7 colistin resistant Enterobacter Species (6, E. cloacae and one, E. aerogenes), 2 (28.57) had MIC 256 µg/ml, 3 (14.29) had MIC 128 µg/ml (Figure 3).
Among 7 colistin resistant isolates, 3 (42.85) were positive for phoP, 5 (71.42) were positive for pmrA, 4 (57.14) were positive for pmrB, 6 (85.71) were positive for pmrC genes detected by PCR. No mgrB, mcr 1 and mcr 2 genes were detected. Multiple gene were detected from urine sample. None of these gene were detected from blood sample (Table VI).
Table VI: Detection of phoP, phoQ, mgrB, pmrA, pmrB, pmrC, mcr-1 and mcr-2 genes from different isolates among colistin resistant E. cloacae and E. aerogenes by PCR (N=7)
|
Genes |
Urine n(%) |
Wound swab n(%) |
ETA n(%) |
Total n(%) |
|
phoP |
3 (42.85) |
0 (0.00) |
0 (0.00) |
3 (42.85) |
|
phoQ |
1 (14.29) |
1 (14.29) |
0 (0.00) |
2 (28.57) |
|
pmrA |
4 (57.14) |
1 (14.29) |
0 (0.00) |
5 (71.42) |
|
pmrB |
2 (28.57) |
1 (14.29) |
1 (14.29) |
4 (57.14) |
|
pmrC |
4 (57.14) |
1 (14.29) |
1 (14.29) |
6 (85.71) |
|
mgrB |
0 (0.00) |
0 (0.00) |
0 (0.00) |
0 (0.00) |
|
mcr 1 |
0 (0.00) |
0 (0.00) |
0 (0.00) |
0 (0.00) |
|
mcr 2 |
0 (0.00) |
0 (0.00) |
0 (0.00) |
0 (0.00) |
N= Total number of colistin resistant E. cloacae and E. aerogenes
n = Number of colistin resistant gene in different samples.
ETA= Endotracheal aspirates.
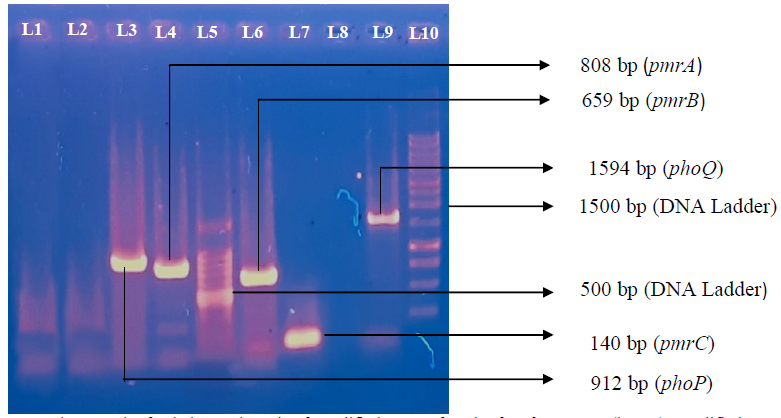
Figure 4: Photograph of gel electrophoresis of amplified DNA of 912bp for phoP, Gene (lane 3), amplified DNA of 808bp for pmrA (lane 4), 100 bp DNA ladder (lane 5), amplified DNA of 659 bp pmrB (lane 6), amplified DNA of 140 bp for pmrC (lane 7), amplified DNA of 1594 bp phoQ (lane 9). One kbp DNA ladder (lane 9), negative control without DNA (TE buffer) (lane one), negative control Escherichia coli ATCC 25922 (lane 2), blank (lane 8).
4. Discussion
Enterobacter has been identified as an important pathogen causing many types of hospital-acquired infections (HAIs) [24]. The emergence of Enterobacter as a worrying resistant pathogen is an important health concern, especially when the scarcity of new antibiotics active against Gram-negative bacteria is considered .As surveillance of AMR and early response to the infection control are crucial steps to bypass the issues, this study aimed to determine the antimicrobial resistance pattern of Enterobacter species isolated from different clinical specimens and also to find out colistin resistant Enterobacter spp. along with resistance gene in Bangladesh.
Out of 350 samples, 228 (65.14 %) samples were culture positive of which 28 (12.28%) were Enterobacter species. These finding are in agreement with the recent study [25] in DMCH who reported that 63.20% of samples (urine, wound swab, ETA, blood, sputum, pus) were culture positive. A study [26] in India reported that a National Nosocomial Infection Surveillance System (NNIS) showed that Enterobacter accounts for 5 to 11% of all nosocomially acquired blood, wound, respiratory tract infection and urinary tract infection. These findings are nearly close to the present finding. In the present study, among 28 isolated Enterobacter species, 22 (78.57%) were identified as Enterobacter cloacae and 6 (21.43%) were identified as Enterobacter aerogenes by biochemical tests. These findings were similar to a study [26] in India who reported that 77.94% were E. cloacae and 22.05% were E. aerogenes. This similarity may be attributed to the fact that these two studies were conducted in same geographic area.
In the present study, thirteen commonly used antibiotics were used on isolated Enterobacter spp, to observe the antimicrobial resistance pattern by disc diffusion method. Among the antibiotics used, Enterobacter spp. was 100% resistant to cefoxitin and amoxiclav. These finding can be correlated with a study [27] who reported that Enterobacter species are intrinsic resistance to aminopenicillins, amoxicillin/ clavulanic acid, cefoxitin and first generation cephalosporins because of the production of chromosomally encoded, inducible AmpC β-lactamase. In the present study, ceftriaxone and ceftazidime had resistance rate 85.71% and 78.57% a study in palestine [28] reported 68.3% resistance to ceftriaxone and 63.14% to ceftazidime which are lower than the present study. CLSI states that, Enterobacter may develop resistance during prolong therapy by third generation cephalosporins as a result of derepression of AmpC β-lactamase. Therefore, isolates that are initially susceptible may become resistant within 3 to 4 days after initiation of therapy [17], which might be true for higher proportion of resistance for Enterobacter spp. to cefuroxime in the present study. In the present study, 57.17% Enterobacter spp. were resistant to amikacin. In contrast to present findings a study in Iran [29] reported 48.60% Enterobacter resistant to amikacin in a study in Iran which is lower than the present findings. The reason behind the higher resistance rate in the present study might be due to the fact that, there was a significant effect of selection pressure of primary antibiotic in the following cases: Amikacin in E. cloacae and E. coli, piperacillin/tazobactam in K.pneumoniae [30]. Among the isolated Enterobacter spp. 32.14% were detected as ESBL producers. A study in India [31], reported the prevalence of ESBL were 33.33% which are in agreement with the present findings. The prevalence of ESBLs among the Enterobacter spp. in Pakistan from different hospital ranged from 14.93% -79% [32]. In case of other members of Enterobacteriacae ESBL production rate has decreased. The prevalence of the ESBL producers had been decreasing over the last few years. Exact reason of such reduction of ESBL is not clear but it can be explained by the fact that the use of ampicillin and cephalosporin has been decreased due to resistant strains and the use of carbapenems and polymyxin has been increased by the physicians which might have some role in decreasing ESBL production. Moreover, the prevalence of ESBL producer varies with time as well as from country to country and even hospital to hospital [33].
In the present study, imipenem resistance in Enterobacter spp. was 42.86%. Imipenem resistance among the species varies widely in different parts of the world. A study [34] in India reported 53.8% imipenem resistance where as a study [28] from Palestine showed resistance rates as low as 12.2%. The frequency of imipenem resistance Enterobacter spp. is increasing in Bangladesh which is reflected by these studies. Risk factors for the acquisition of carbapenem resistant Enterobacteriaceae in a tertiary care hospital may include, prior exposure to antimicrobials, long term hospital stay especially in ICU, high dependency units, renal, hematology, oncology unit, presence of indwelling devices such as central line, urinary catheter, endotracheal tube and enteral feeding tube, co-morbid or immune compromised patients [35]. In the present study, among 12 imipenem resistant Enterobacter spp. 75% carbapenemase producers were detected by CD assay. In a previous study [36] in Dhaka Medical College hospital, 71.43% carbapenamase producing Enterobacter spp. was detected by CD assay which is close with the present findings.
Among the isolated Enterobacter spp. 7 (25%) colistin resistant Enterobacter spp were identified. A study in Korea [37] reported that 16% of Enterobacter spp were resistant to colistin. Another study [38] reported 13.9%- 20.1% colistin resistances in Enterobacter spp. However, colistin resistance is rare upon first isolation but often arises during treatment via mutations [39]. Now a day, the use of colistin has increased due to increased prevalence of CRE which might be the reason for emergence of resistance of this reserve drug. In the present study, 15 (53.57%) multidrug resistant Enterobacter spp. were isolated from different samples (urine, wound swab, ETA, blood). A study [40] in Nepal reported that 52.90% Enterobacter spp. were MDR.
Among colistin resistant Enterobacter species, 6 (85.71%) E. cloacae and 1 (14.28%) E. aerogenes were isolated from different clinical samples. A study in Korea[37] reported among colistin resistant Enterobacter 23.9% were E. cloacae and 4.2% were E.aerogenes which is similar to our study. MIC range of colistin among colistin resistant Enterobacter species were 16µg/ml- ≥ 256µg/ml. This result suggests that colistin resistance in Enterobacter might be associated with chromosomal mutations in mgrB, phoP/phoQ, pmrA, pmrB, pmrC and crrABC which also described in K. pneumoniae [41]. High MIC may also be due to strong selective pressure in the isolates [42].
Among the colistin resistant Enterobacter species, 85.71% were positive for pmrC, 71.42% were positive for pmrA, 57.14% were positive for pmrB, 47.61% were positive for phoP. In the present study, mgrB, mcr-1 and mcr-2 genes were also searched but among colistin resistant Enterobacter spp. none of the isolates was positive for any of the genes. In Enterobacter spp. mutations of the pmrAB and phoPQ genes are mainly involved in colistin hetero resistance [43]. To best our knowledge no available data was found to compare the prevalence of colistin resistance genes among colistin resistant Enterobacter spp.
5. Conclusion
Our study result shows Enterobacter species seem to be emerged increasing resistance to multiple antibiotics may be due to the pervasive and irrational practices of antibiotic prescription and use. In our study we have also identified a high colistin resistance rate in E. cloacae but not in E. aerogenes. Therefore, prompt action is recommended for proper infection control. More emphasis should be given on routine AMR surveillance and antibiotic stewardship programs for the better management of patients.
6. Acknowledgements
Department of Microbiology, Dhaka Medical College, Bangladesh provided laboratory support to perform this study.
7. Financial Support and Sponsorship
Nil
8. Conflict of Interest
There are no conflicts of interest in this study
9. References
- Langley JM, Hanakowski M, Leblanc JC. 2001. Unique epidemiology of nosocomial urinary tract infection in children. Am. J. Infect. Control 29 (2001): 94–98.
- Boucher HW, Talbot GH, Bradley JS, Edwards JE, Gilbert D, Rice LB, Scheld M, Spellberg B, Barlett J. Bad bugs, no drugs: no ESKAPE! An update from the Infectious Diseases Society of America. Clin. Infect Dis 48 (2009): 1–12.
- Bae S, Kim T, Kim MC, Chong YP, Kim SH, Sung H et al. Clinical Characteristics and outcomes of spontaneous bacterial peritonitis caused by Enterobacter Species versus Escherechia coli: a matched case-control study. Bio Med Infect Dis 252 (2016): 1-6.
- Patel KK, Patel S. Enterobacter spp: An emerging nosocomial infection. Intl J Appl Res 2 (2016): 532-538.
- Chavda KD, Chen L, Fouts DE, Sutton G, Brinkac L, Jenkins SG et al. Comprehensive genome analysis of carbapenemase producing Enterobacter spp: new insights into phylogeny, population structure and resistance mechanisms. mBio 7 (2016).
- Shad A. MCR-1 colistin resistance in Escherichia coli wildlife: a continental mini-review. J Drug Metab Toxicol 9 (2018): 243.
- Gallet AVC, Ocampo AM, Chavda K, Chen L, Barry N. Kreiswirth BN et al. Molecular epidemiology of carbapenem-resistant Enterobacter cloacae complex infections uncovers high frequency of non-carbapenemase-producers in five tertiary care hospitals from Colombia. bioRxiv preprint first posted online (2018).
- Wilson BM, El Chakhtoura NG, Patel S, Saade E, Donskey CJ, Bonomo RAet al. Carbapenem-Resistant Enterobacter cloacae in Patients from the US Veterans Health Administration, 2006-2015. Emerg Infect Dis 23 (2017): 878-880.
- Jia X, Dai W, Ma W, Yan J, He J, Li S, et al. Carbapenem-resistant E. cloacae in Southwest China: molecular analysis of resistance and risk factors for infections caused by NDM-1-producers. Front Microbiol 9 (2018): 658.
- Nation RL, Li J. Colistin in the 21st century. Curr Opin Infect Dis 22 (2009): 535 -543.
- Norgan AP, Freese JM, Tuin PM, Cunningham SA, Jeraldo PR, Patel R. Carbapenem- and colistin-resistant Enterobacter cloacae from Delta, Colorado, in 2015. Antimicrob Agents Chemother 60 (2016): 3141-3144.
- Liu YY, Wang Y, Waslsh TR, Yi LX, Zhang R, Spender J et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: a microbiological and molecular biological study. Lancet Infect Dis 16 (2016): 161-68.
- Acharya KP, Wilson RT. Antimicrobial resistance in Nepal. Front Med (2019).
- Gould IM. Hospital antibiotic stewardship to control resistance—how should it be done? In: Antibiotic Policies. New York, NY: Springer (2012): 39–52.
- Chessbrough M. Microscopial techniques used in Microbiology, culturing bacterial pathogens, biochemical test to identify bacteria. In: Chessbrough M, ed. District Laboratory Practice in Tropical Countries, Part 2, 2ndedn: Cambridge University Press, India (2009): 35-70.
- Cheesbrough M. Antimicrobial susceptibility testing. In: Cheesbrough M, (editor). District Laboratory Practice in Tropical countries, part 2, 2nd Cambridge University press (2009):132-143.
- Clinical Laboratory Standards Institute. Performance Standard for Antimicrobial Susceptibility Testing: Twenty-seventh Informational Supplement M200-S28. Wayne, PA, USA: CLSI; (2018).
- European Committee on Antimicrobial Susceptibility Testing. Breakpoint tables for interpretation of MICs and zone diameters.Version 8.0, valid from 2018-01-01. EUCAST: Clinical breakpoints eucast.org/clinical breakpoints/ (accessed on- 13.02.2019).
- Jarlier V, Nicolas M, Fournier G, Philippon A. Extended broad-septrum β-lactamase conferring transferable resistance to newer β-Lactum agents in Enterobacteriaceae hospital prevelance and susceptibility patterns. Rev Infect Dis 10 (1988): 867-878.
- Qu TT, Zhang JL, Wang J, Tao J, Yu YS, Chen YG et al. Evaluation of phenotypic tests for detection of Metallo-β-Lactamase producing Pseudomonas aeruginosa strains in chaina. J Clin Microbiol 47 (2009): 1136-1142.
- Andrews JM. Determination of minimum inhibitory concentrations. J Antimicrob Chemother 48 (2001): 5-16.
- Franco MR, Caiaffa-Filho HH, Burattini MN, Rossi F. Metallo-beta-lactamases among imipenem-resistant Pseudomonas aeruginosa in a Brazilian university hospital. Clinics (Sao Paulo) 65 (2010): 825-829.
- Haeili M, Javani A, Moradi J, Jafari Z, Feizabadi MM, Babaei E et al.MgrB alterations mediated colistin resisteance in Klebsiella pneumoniae isolates from iran. Front Microbiol 8 (2017): 2470.
- Lockhart SR, Abramson MA, Beekmann SE, Gallagher G, Riedel S, Diekema DJ, et al. Antimicrobial resistance among Gram-negative bacilli causing infections in intensive care unit patients in the United States between 1993 and 2004. J Clin Microbiol 45 (2007): 3352–9.
- Jahan S. Efficacy of the antibiotic combinations in the colistin resistant Klebsiella pneumoniae both in vitro and in vivo. DMC (2018).
- Sujatha B, Shobha KL, Rao AS, Rao GS. Antibacterial Susceptibility Pattern of Uropathogenic Enterobacter Species from a Tertiary Care Hospital. JKIMSU 7 (2018): 1-6.
- Dimitrova D, Stoeva T, Markovska R, Stankova P, Bozhkova M et al. Antimicrobial susceptibility of clinically significant isolates of Enterobacter spp., obtained from patients, hospitalized in Varna University the period 2014-2016. J of IMAB 23 (2017): 1828-1833.
- Adwan G, Rabaya D, Adwan K, Al-Sheboul S. Prevalence of β-lactamases in clinical isolates of Enterobacter cloacae in the West Bank-Palestine. International Journal of Medical Research & Health Sciences 5 (2016): 49-59.
- Mortazavi SM, Mansouri F, Azizi M, Alvandi A, Karbasfrushan A, Goli NM et al. Prevalence of Class I and II Integrons among MDR Enterobacter cloacae Isolates Obtained from Clinical Samples of Children in Kermanshah, Iran. Journal of Clinical and Diagnostic Research 12 (2018): 13-16.
- Sedlakova MH, Urbanek K, Vojtova V, Suchankova H, Imwensi P, Kolar M. Antibiotic consumption and its influence on the resistance in Enterobacteriaceae. BMC Res Notes 7 (2014): 1-10.
- Wadekar MD, Anuradha K, Venkatesha D. Phenotypic detection of ESBL and MBL in clinical isolates of Enterobacteriaceae. Int J Current Res Acad Rev 1 (2013): 89-95.
- Amin H, Zafar A, Ejaz H, Jameel NU. Phenotypic characterization of ESBL producing Enterobacter cloacae among children. Pak J Med Sci 29 (2013): 144-147.
- Farzana R, Shamsuzzaman SM, Mamun KZ, Shears P. Antimicrobial susceptibility pattern of extended spectrum β-lactamase producing gram-negative bacteria isolated from wound and urine in a tertiary care hospital, Dhaka City, Bangladesh. The Southeast Asian journal of tropical medicine and public health 44 (2013): 96-103.
- Khajuria A, Praharaj AK, Kumar M, Grover N. Carbapenem resistance among Enterobacter Species in a tertiary care hospital in central India. Chemotherapy Research and Practice (2014): 1-6.
- Ling ML, Tee YM, Tan SG, Amin IM, How KB et al. Risk factors for acquisition of carbapenem resistant Enterobacteriaceae in an acute tertiary care hospital in Singapore. Antimicrob Resist Infect Control 4 (2015): 26 (1-7).
- Tafneen FA. Detection of AMPC β-lactamases, extended spectrum β-lactamases and carbapenemases in common Enterobacter species by phenotypic and genotypic methods with their antibiotic resistance pattern. [M.Phil thesis] DMC (2018).
- Ko KS, Hong YK, Lee JY. Colistin resistance in Enterobacter isolates in Korea. Journal of Microbiology 56 (2018): 435-440.
- Afonso LV, Dalmolin TV, Daiana LM. Plasmid-mediated Colistin Resistance: What Do We Know? J Infectiology 1 (2018): 16-22.
- Wyres KL, Holt KE. Klebsiella pneumoniae population genomics and antimicrobial-resistant clones. Trends Microbiol 24 (2016): 944-956.
- Adhikari RP, Shrestha S, Rai JR, Amatya R. Antimicrobial Resistance Patterns in Clinical Isolates of Enterobacteriaceae from a Tertiary Care Hospital, Kathmandu, Nepal. Nep Med J 1 (2018): 74-78.
- Poirel L, Kiefer N, Nordmann P. In vitro study of ISApl1-mediated mobilization of the colistin resistance gene mcr-1. Antimicrob Agents Chemother (2017).
- Dalmolin TV, Lima-Morales DD, Barth AL. Plasmid-mediated colistin resistance: what do we know? J Infectiol 1 (2018): 16–22.
- Uechi K, Tada T, Shimada K, Nakasone I, Kirikae T, Fujita J. Emergence of a carbapenem-resistant and colistin-hetero resistant Enterobactercloacae clinical isolate in Japan. J Infect Chemother 25 (2018): 285-288.



 Impact Factor: * 3.5
Impact Factor: * 3.5 Acceptance Rate: 71.36%
Acceptance Rate: 71.36%  Time to first decision: 10.4 days
Time to first decision: 10.4 days  Time from article received to acceptance: 2-3 weeks
Time from article received to acceptance: 2-3 weeks 
