Could the Omicron Variant be the last Variant of Concern of the COVID-19 Pandemic? - Global Immunity is Key
*Yves Muscat Baron
1Mater Dei Hospital, Medical School of Malta, University of Malta, Malta
*Corresponding Author: Yves Muscat Baron. Mater Dei Hospital, Medical School of Malta, University of Malta, Malta
Received: 12 December 2022; Accepted: 19 December2022; Published: xxxx.
Article Information
Citation: Yves Muscat Baron. Could the Omicron Variant be the last Variant of Concern of the COVID-19 Pandemic? Global Immunity is Key. Journal of Environmental Science and Public Health. 7 (2023): 07-13.
View / Download Pdf Share at FacebookAbstract
The Omicron variant was designated a Variant of Concern (VoC) due to its increased transmissibility and antibody evasion. Data from several countries however suggested a milder clinical outcome for the Omicron variant compared to the previous VoCs. The clinical outcome in the coming year (2023) is however uncertain due to Omicron’s persistent evolution, developing variants with increased immune escape attributes in the presence of populations that may not possess adequate immunological defences. The Omicron variant utilizes the endosomal route of cell entry unlike previous VoCs. This may be due to Omicron’s superior spike protein receptor binding domain (RBD)’s adhesion to the host cell’s angiotensin converting enzyme II (ACE2) receptor. Efficient cell entry may have increased Omicron’s tropism to rapidly infect the extensive surface area of the nasopharyngeal mucosa and its adjacent sinuses. The endocytic mode of cell invasion may result in a more efficient recruitment of several contemporaneous RBD-ACE2 complexes of the same virus and other viruses to the attached host cell, suggesting a correlation between viralhost cell binding and transmissibility and a negative correlation with clinical severity. The nasopharyngeal region acting as a buffer, would have gained time with the initial containment of the Omicron infection providing immunological protection, preceding significant seeding into the lungs. The combination of previous waves of natural infection, uneven global vaccination efforts and widespread Omicron infection and its most recent sub-variants (BF7 and XXB), may elude worldwide immunity, exacerbating the pathogenic effects of future SARS-CoV-2 outbreaks. Emulating the pattern of waves of infection during the devastating 1918 Spanish influenza, the current COVID-19 Pandemic may have approached an upended immunological equilibrium, due to adverse immunological, anthropogenic and environmental factors, swaying in favour of
Keywords
<p>variant of concern; COVID-19 pandemic; immunity; RBDACE2 binding; transmissibility; global immunity.</p>
Article Details
3. Introduction
In early November 2021, the first sequenced Omicron case was reported from Botswana and a few days later another case was reported from Hong Kong in an individual travelling from South Africa. Later in the same month, scientists from South Africa, observed a cluster of COVID-19 cases in the Gauteng Province prompting the World Health Organization to designate a newly sequenced variant of concern (VoC) named the Omicron variant[1].
Epidemiological studies rapidly indicated that the Omicron variant was highly transmissible. The reproduction number (Ro) of the Omicron variant at its peak was calculated to be at least 10 compared to Delta’s Ro 7.5 and the original variant Wuhan Hu 1 Ro of 2.5 [2]. Moreover genomic studies revealed an unprecedented constellation of mutations on the spike protein and viral surface coating, prompting fear of immune escape from both natural infection and current vaccines. Contrasting with previous variants of concern, it became apparent that hospitalizations and deaths in South Africa did not surge following the emergence of the Omicron variant [3]. A similar pattern was seen in Denmark and the U.K. [4, 5]. Over the past year, over 300 variants of the Omicron Virus have been detected, with a recent identification of 130 subvariants in China raising the prospect of the emergence of a more pathological variants as the BF7 and XBB subvariants [GISAID].
Viral adhesion and incorporation into the host cell requires an extensive array of synchronized complex network of salt bridges, hydrophobic sites, hydrogen bonding and electrostatic interactions between the viral RBD and the host cell ACE-2 receptor.[6] Omicron’s host cell invasion is executed by endosomal cellular incorporation, independent of TMPRSS-2 (transmembrane protease serine type 2) and furin priming, as was enacted with previous VoCs.[7,8] Binding to the ACE2 receptor appears to be stronger than previous VoCs, possibly encouraging contemporaneous multiple RBD-ACE-2 receptor complexes between a single virus and other viruses to the host cell to occur simultaneously. The various stages of multiple viral invasions of the host cell may also interfere with syncythial cell formation, the hallmark of COVID-19 alveolar pathology.
This paper hypothesizes that the Omicron variant and its sub-variants, due to their stronger RBD-ACE2 binding, mode of cell invasion and tropism for the nasopharyngeal region increasing its transmissibility, may eventually result in a more uncertain course of the COVID-19 Pandemic. This uncertain course may be due to the uneven global anti-SARS-CoV-2 immunity through the combination of variable waves of SARs-CoV-2 infections, heterogenous global vaccination and widespread infection with the Omicron variant and its sub-variants in large inadequately protected populations.
3.1 Omicron a Variant of Concern
The Omicron variant was designated by the World Health Organization (WHO) in November 2020 as a Variant of Concern because of its greater transmissibility, evasion from antibody neutralization, and due to the overwhelming rate of infections, it was portrayed to be of serious consequence to the population at large. [2]
The clinical outcome following infection with the Omicron variant did not appear to be as severe as the previous Variants of Concern.[3,4,5] As a consequence, the Omicron variant presented itself as a mild, highly transmissible upper respiratory tract infection, without causing significant pulmonary pathology as the previous variants of concern. Accordingly, Omicron infection potentially provided large swathes of the population with the attribute of herd immunity against future SARS-CoV-2 viruses. Moreover Omicron’s sub-variants, such as the Stealth Omicron (BA.2), may supplant its progenitor due to evolutionary superior RBD-ACE2 binding and transmissible attributes.
3.2 Omicron has a greater tropism to the Upper Respiratory Tract Epithelium
The reason why the Omicron variant has been contained as an upper respiratory tract infection, instead of being highly invasive in lung tissue as the previous variants of concern, is as yet not clear. There appears to be a change in the mode of viral invasion of the host cell which may partially explain the change in viral target. The Omicron variant is far superior to the Delta virus at infecting and replicating in the bronchial epithelium (70-fold) and conversely is less infectious in lung tissue. [9]
The conventional thinking as regards host cell invasion by previous variants of concern, involved the bonding of SARS-CoV-2 spike (S) protein to the host cell’s angiotensin converting enzyme II (ACE2) receptor, which was followed by TMPRSS-2 and furin priming of the S protein for protease processing, allowing eventual viral cell entry.[10] In the case of the Omicron variant, it appears that this variant achieves entry into the host cell through binding with ACE2 receptor, followed by endosomal fusion, independent of TMPRSS and furin spike protein priming.[7,8] Omicron and its sub-variants appear to possess a stronger interface binding between the RBD and the ACE2 receptor which may explain the more efficient endosomal route of cell entry.[11]
Variation in energy binding has been postulated through extensive all-atom molecular dynamics simulations and advanced free energy calculations caused by the inclusion and exclusion of viral mutations. [12] Possibly this increased binding property provides the possibility that multiple RBD’s of the infecting virus (size 0.07μm to 0.09μm) together with other viruses, to contemporaneously attach to several ACE2 receptors of the same cell (mean size 12 μm). This latter process may be more efficient than the ACE2, TMPRSS-2 and furin cell entry pathway. Moreover the different stages of multiple viral invasion of the cell, may interfere with syncythial cell development.
The increased binding and initial containment of the Omicron virus in the upper respiratory tract, including the nasopharynx, the frontal and maxillary sinuses may have an important immunological role. Through their characteristic extensive surface mucosal area, the nasopharynx, the frontal and maxillary sinuses may mount a rapid and adequate antibody reaction. The extensive inflammatory processes occurring in the frontal sinus epithelium may be the cause of the severe headaches typified by the Omicron variant infection. The time gained during nasopharyngeal infection, may be sufficient to mount an adequate antibody response, preceding significant seeding of infected particles deeper in the lungs, diminishing the risk of severe lung infection. The time factor is crucial as inadequate antibody response will increase the risk to allow invasive lung disease to set in.
3.3 Origin of the Omicron Variant
It is interesting to note, that an in-depth analysis of the proportion of variants from world genomic data, indicates that countries where variants of concern (VoC) and variants of interest (VoI) were detected, had a differential proportion of variants designated as “others”, contemporaneous with the VoC/VoI emergence (Table I). Variants designated as “others” may have offered the appropriate substrate from which a variant of concern or variant of interest was naturally selected.
In vitro studies have shown that certain mutations namely S477N, E484K and N501Y increase transmissibility, however other mutations may be epistatic to the phenotypic expression of these genes.[13] This may suggest, that emulating the process of natural selection, it is unlikely that a “highly successful” variant will undergo a large number of mutations to produce a “more highly successful” one, as a significantly elevated number of mutations may detract from its virulent efficiency – why change a winning formula. This was the situation with the Delta variant obtaining dominance in India over the two other sister variants – Kappa and 1617.3 variants.
At the initial stages of the pandemic, a single amino acid substitution (aspartic acid displaced by glycine) on the original template, resulted in the G614 mutation improving SARS-CoV-2 infectivity and pathogenicity over the progenitor Wuhan Hu 1.[14,15] The converse may be occurring with Omicron variant, whereby the highly successful Delta variant could only be out-competed by the Omicron variant, because it appears to have been derived from a completely different lineage and infecting a different target organ (Table 2.).[7]
This selective process may also have occurred with the inception of the alpha variant. The 20A.EU1 variant (designated as an “other variant”) was dominant in Spain in mid-August 2020 and was transferred to the U.K. in mid-September 2020, possibly by tourists returning from their holidays in Spain. Two clusters of the 20A.EU1 variant were noted in mink farmers in North East region of Spain suggesting reverse zoonosis. [16] A similar process of reverse zoonosis may also have occurred with the Omicron variant utilizing a mouse host.[17] The B.1.1.7 variant and the 20A.EU1 variant appear to evolve from the same 20A progenitor clade[18] and possibly the 20A.EU1 variant, may have atypically reversed its mutation to the progenitor, which later went on to produce the Alpha B.1.1.7 variant.[19]
|
Variant |
Average number of mutations |
|
Alpha |
29.7 |
|
Gamma |
29.1 |
|
Beta |
28.4 |
|
Delta |
35.4 |
|
Omicron |
53.3 |
Table1: The average number of mutations per sample in each variant of concern. Adapted from: CoVariants.org and
GISAID.
3.4 Immunity following Vaccination and Natural Infection
The current vaccines were engineered on the original genotype Wuhan Hu 1. Following the progenitor Wuhan Hu 1, a number of variants promulgated the pandemic. These variants involved the G614 mutation, the Alpha, Beta, Gamma, Delta and now the Omicron variants.
Despite the variation in spike protein mutations, the vaccines still provide a measure of protection from severe disease. This is due to the activity of antibodies derived from the vaccine, which evade the variants’ mutations, attacking the epitopes belonging to the template of the original viral phenotype. Undoubtedly long-term T-cell immunity will provide additive protection derived from previous infections with VoCs and vaccination. [20]
|
Variant of Concern/Interest VoC/VoI |
Country of Origin |
Percentage of “Other” variants at point of increase in VoC/VoI gradient |
|
Beta |
South Africa |
87.60% |
|
Alpha |
UK |
87.30% |
|
Epsilon |
USA |
82% |
|
Gamma |
Brazil |
77.80% |
|
Lambda |
Peru |
67% |
|
Mu |
Columbia |
64.30% |
|
Iota |
USA |
64% |
|
Eta |
Nigeria |
56.60% |
|
Kappa |
India |
55% |
|
Delta |
India |
28.90% |
|
Omicron |
Botswana |
4.80% |
Table 2: Percentage of genomic detection of Variants of Concern/Interest/”Others” with Country of origin in descending order.
Percentage of variants designated as “Others” when there was a significant gradient in the increase of VoC/VoI genomic detection. The variants Delta and Omicron with the greatest transmissibility had the lowest percentage of “Other” variants possibly suggesting that their inception and successful dominance was determined by superior transmission/selective pressure out-competing existing variants. The vaccination drive during the emergence of the Delta and Omicron variants may have also significantly increased the competitive inhibition, limiting the survival of “Others” variants.[Adapted from: CoVariants.org and GISAID 5th January 2022].
Severe pathology of COVID-19 emanates from viral invasion of the pneumocyte I and II cells in the alveoli.[21] Necrosis and anatomical devastation is wrought by the viral infection, the reactive cytokine storm and secondary bacterial infections.[22,23] Contrasting with the Delta variant, the Omicron variant does not appear to elicit a severe intracellular interferon response, diminishing the risk of a severe reactive cytokine storm.[24]
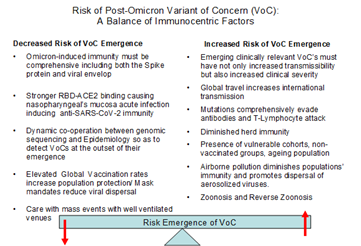

Figure 1: The possibility of the emergence of a Post-Omicron Variant of Concern depends on a number of contrasting risk factors. The main theme delineating these risks, encompasses the global population’s comprehensive immunity. The organization of the global response to the emergence of a variant will also determine whether the variant will develop into a Variant of Concern.
The situation now may mirror that encountered early in 2020. Progressive waves of the pandemic in different parts of the world, with a variety of variants of concern/interest, allied to the global vaccination drive, may have provided a substantial proportion of the world’s populations with an adequate level of comprehensive immunity against SARS-CoV-2, but other populations with different immunological profiles may not have this protection. In the alveoli, circulating antibodies and T-lymphocytes are a few micrometres away from the line of viral invasion. The concentration and proximity of these defences, allow rapid and effective containment of infection, before syncythial development (Figure 2.).[21]
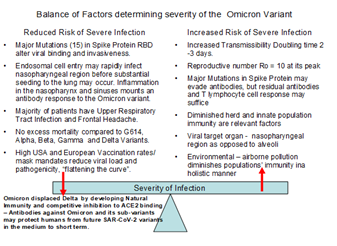
Figure 2: Immunological equilibrium of the population is balanced by factors decreasing severity of Omicron infection contrasting with factors increasing risk of severe infection by the Omicron variant. Omicron variant rapidly displaced Delta by developing natural immunity to its own epitopes and those of the Delta variant – The antibody response mounted against Omicron is comprehensive, unlike the vaccines which concentrate on the spike protein. This comprehensive antibody response may protect humans from future SAR-CoV-2 variants in the medium to short-term. This protection may not have occurred in populations with different immunological profiles as may have happened in China and India.
3.5 COVID-19 and the Influenza Pandemics
The critical importance of an immunological memory on the scale of the global human population may be inferred from the Influenza pandemics. In the 1918 Spanish influenza pandemic, the mortality (40-50 million worldwide) curve simulated a “W” as opposed to the usual “U” curve, indicating that not only the very young and elderly were effected, but atypically, 20-40 year old adults also succumbed to the pandemic.[25] A suggestion had been proposed that the influenza pandemic 30 years earlier, in 1889 (called the Russian flu), may have provided the 1918 middle-aged (older than 40years) and elderly populations with antibody and T-cell immunity derived from the immunological memory of the preceding pandemic.[26]. Moreover the symptomatology and the epidemiology of the Russian influenza suggest a possible zoonotic aetiology.[27]
Following the 1918 the devastating Spanish influenza pandemic, the subsequent next pandemic, called the Asian influenza occurred 38 years later, in 1957.[28] The 1957 Asian influenza is thought to have a zoonotic origin when viral mutations in wild foul shared genes with a pre-existing human influenza strain.[29] The Asian influenza is estimated to have caused significant mortality approximating 1.1 million excess deaths (95% CI, 0.7 million-1.5 million excess deaths)[30] prompting the following quote:
‘Although we have had 30 years to prepare for what should be done in the event of an influenza pandemic, I think we have all been rushing around trying to improvise investigations with insufficient time to do it properly. We can only hope that people will have taken advantage of their opportunities and at the end it may be possible to construct an adequate explanation of what happened”. [31]
3.6 Threats to the Global Immunological Equilibrium
As suggested by the influenza and SARS pandemics, there are several threats to the global populations’ immunological equilibrium. Worldwide travel may be one factor that may breach a population’s immunological shell, as its immunity may not be aligned to that of another distant population. An outbreak can become a pandemic within a few days, as global travel is much more efficient than in the 1918. An Indian regiment recruited during the First World War took approximately one month to travel from the Western Front in France to Bombay. [32] There is a suggestion that this was the series of events whereby the Spanish influenza was introduced to the Indian subcontinent, where it was initially called the Bombay Fever, resulting in an estimated 17 million loss of lives.[33]
Population vulnerabilities come into further focus by the different national demographic characteristics, with immunological ancestry may also play a part, as would ethnic receptor isoforms as in case of the ACE2 receptor.[34,35] There also lies the increased risk of zoonosis and reverse zoonosis which may generate future variants of concern.[16] The unprecedented survival of immuno-suppressed individuals and the ageing population, provide more ideal circumstances for the development of variants, due to elevated viral loads. Hospitalization for these vulnerable groups even in the presence of mild disease, and especially in the setting of overwhelmed Healthcare services, present a threat to patient safety.
3.7 Atmospheric Pollution and the Emergence of variants of Concern
The incessant atmospheric pollution, instigating climate change, undoubtedly is holistically impacting the populations’ immunity in an adverse manner in both the short and long-term. Airborne pollution with particulate matter has been well established as a prime factor in the propagation of SARS-CoV-2 port of entry, the ACE2 receptor and possibly may also act as a vector for viral transmission.[18,36] Airborne pollution may also have been responsible in SARS-CoV-2’s evolution.[14,37] Similar to the emergence of other VoCs and VoIs, the emergence of the Omicron variant in South Africa and its sub-variant Stealth Omicron in Kolkata appear to have been preceded by elevated levels of airborne pollution including particulate matter PM2.5.[38,39]. A crucial environmental factor that may have led to the emergence and spread of more virulent subvariants in the Chinese population, areF the widespread smoking habits in China, whereby 66% of Chinese males indulge in tobacco smoking, which is replete with PM2.5.[15].
During national and regional lockdowns there was a significant perceptible reduction in atmospheric pollution. Despite these significant reductions, airborne pollution returned to its former levels after lockdowns were lifted.[40] It is biologically plausible and scientifically proven that this environmental factor, profoundly influencing respiratory health, may be a crucial co-factor in the genesis of the COVID-19 Pandemic and its prolongation due to the emergence of subsequent VoCs. It is a sobering thought that all the global effort to contain the COVID-19 pandemic which caused such huge morbidity and mortality, at incalculable economic and social cost, could be undone because of the inertia of the international community to harness escalating atmospheric pollution.
3.8 Actions that need to be undertaken:
- Genomic surveillance closely interconnected with dynamic epidemiology has to be maintained to detect variants in a timely manner.
- More basic science is required to elucidate the positive correlation of viral RBD binding to epithelial receptor and resultant viral transmissibility
- More basic science is required to examine the possible negative correlation between RBD binding to epithelial receptor and invasiveness of the respiratory tract, impacting clinical severity.
- Physical and social distancing is critical to “flatten the curve”, as overwhelmed Healthcare services even with mild disease do pose a threat to patient safety especially to the vulnerable.
- Respiratory health has to be fostered by encouraging less air pollution and further curbing of tobacco smoking.
- A healthier atmospheric environment can only be attained with a determined reduction on fossil fuel reliance.
4. Conclusion
As intimated in this paper, the outcome of a pandemic is a delicate balance dependent on the factors governing the global holistic immunity and the virulence of the infecting agent. The Omicron variant may be the last variant of concern, until such time the balance sways in favour of a more virulent virus able to circumvent the immunological equilibrium attained. As demonstrated in this paper, the emergence of another Variant of Concern, very much depends on a number of caveats which to a major extent may be determined by anthropogenic activity and environmental health determinants.
References
- Technical Advisory Group on SARS-CoV-2 Virus Evolution (2020).
- Khan Burki T. Omicron variant and booster COVID-19 vaccines 2021 Lancet Respiratory Medicine. 10(2)(2021): (21)00559-2.
- Wolter N, Jassat W, Walaza S, et al. Early assessment of the clinical severity of the SARS-CoV-2 Omicron variant in South Africa. medRxiv 399(10323)( 2022): 437-446.
- Plesner Lyngse F, Mortensen LH, Denwood MJ, et al. SARS-CoV-2 Omicron VOC Transmission in Danish Households. medRxiv 10(2022).
- Meng B, Ferreira IA, Abdullahi A, et al. SARS-CoV-2 Omicron spike mediated immune escape, infectivity and cell-cell fusion bioRxiv 3(2022).
- Taka S,Yilmaz Z, Golcuk M, et al. Critical Interactions Between the SARS-CoV-2 Spike Glycoprotein and the Human ACE2 Receptor E. bioRxiv 125(21)(2021): 5537-5548.
- Willett BJ, Grove J, MacLean OA, et al. The hyper-transmissible SARS-CoV-2 Omicron variant exhibits significant antigenic change, vaccine escape and a switch in cell entry mechanism. medRxiv 7(2022): 1161–1179.
- Peacock TP, Brown JC, Zhou J, et al. The SARS-CoV-2 variant, Omicron, shows rapid replication in human primary nasal epithelial cultures and efficiently uses the endosomal route of entry. bioRxiv 10(2021).
- Nancy Lapid. Omicron thrives in the airways no in the lungs. New data on asymptomatic cases. 7(2021).
- Torre-Fuentes L, Matías-Guiu J, Hernández-Lorenzo L. ACE2, TMPRSS2, and Furin evariants and SARS-CoV-2 infection in Madrid, Spain. J Med Virol 93(2)( 2021): 863-869.
- Woo GWE, Shah M. Omicron: A heavily mutated SARS-CoV-2 variant exhibits stronger binding to ACE2 and potently escape approved COVID-19 therapeutic antibodies bioRxiv 12(2021): 830527.
- Aggarwal A, Naskar S, Maroli N. Mechanistic Insights into the Effects of Key Mutations on SARS-CoV-2 RBD-ACE2 Binding bioRxiv 23(46)(2021).
- Zahradník J, Marciano S, Shemesh M, et al. SARS-CoV-2 variant prediction and antiviral drug design are enabled by RBD in vitro evolution. Nat Microbiol 6(9)( 2021):1188-1198.
- Korber B, Fischer WM, Gnanakaran S, et al, Tracking Changes in SARS CoV-2 Spike: Evidence that D614G Increases Infectivity of the COVID-19 Virus. Cell 182(4)(2020): 812–827.
- Baron YM. Could changes in the airborne pollutant particulate matter acting as a viral vector have exerted selective pressure to cause COVID-19 evolution? Med Hypotheses. 146(2021): 110401.
- Muscat Baron Y. Intra and Interspecies Interaction Between Mass Confined Animals and Their Handlers – An Ideal ReserVoIr for Coronavirus Evolution. Canadian Journal of Infection Control. Spring Edition (2021): 39-40.
- Sun Y, Lin W, Dong W, et al. Origin and evolutionary analysis of the SARS-CoV-2 Omicron variant. Journal of Biosafety and Biosecurity 4(1)(2022): 33-37.
- Lauring AS, Hodcroft EB. Genetic variants of SARS-CoV-2—What do they mean? JAMA 325(6) (2021):529–31.
- Baron YM. Are there medium to short-term multifaceted effects of the airborne pollutant PM2.5 determining the emergence of SARS-CoV-2 variants? Med Hypotheses 158(2021): 110718.
- Keeton R, Tincho MB, Ngomti A, et al. SARS-CoV-2 spike T cell responses induced upon vaccination or infection remain robust against Omicron. 603(2022): 488–492.
- Bösmüller H, Matter M, Fend F, et al. The pulmonary pathology of COVID-19. Virchows Arch 478(1) (2021): 137-150.
- Nile SH, Nile A, Qiu J, et al. COVID-19: Pathogenesis, cytokine storm and therapeutic potential of interferons. Cytokine Growth Factor Rev 53(2020): 66-70.
- Langford BJ, So M, Raybardhan S, et al. Bacterial co-infection and secondary infection in patients with COVID-19: a living rapid review and meta-analysis. Clin Microbiol Infect 26(12)(2020): 1622-1629.
- Bojkova D, Widera M, Ciesek S, et al. Reduced interferon antagonism but similar drug sensitivity in Omicron variant compared to Delta variant SARS-CoV-2 isolates. bioRxiv 32(2022): 319–321.
- Luk J, Gross P, Thompson WW. Observations on mortality during the 1918 influenza pandemic. Clin Infect Dis 33(8)(2001): 1375-8.
- Pyle G. The Diffusion of Influenza: Patterns and Paradigms. Chapter 2 Precursors to the Conventional Wisdom: Some European Experiences 5(1986): 23-33.
- Brüssow H, Brüssow L. Clinical evidence that the pandemic from 1889 to 1891 commonly called the Russian flu might have been an earlier coronavirus pandemic. Microb Biotechnol 14(5)(2021): 1860-1870.
- Vynnycky E, Edmunds WJ. Analyses of the 1957 (Asian) influenza pandemic in the United Kingdom and the impact of school closures. Epidemiol Infect 136(2)(2008):166-79.
- Kilbourne ED. Influenza pandemics of the 20th century. Emerg Infect Dis 12(1)(2006): 9-14.
- Viboud C, Simonsen L, Fuentes R, et al. Global Mortality Impact of the 1957-1959 Influenza Pandemic. J Infect Dis 213(5) (2016): 738-45.
- RCGP Archives. Between Ourselves. ACE G3–4 (1957).
- Bartholomew JG. An Atlas of Economic Geography. Proceedings of the Royal Geographical Society. 15 (2021).
- Chandra S, Kuljanin G, Wray J. Mortality From the Influenza Pandemic of 1918–1919: The Case of India. Demography 49(2012): 857–865.
- Muscat Baron Y. Could the Denisovan Genes have conferred enhanced Immunity against the G614 Mutation of SARS-CoV-2?. Human Evolution 36(1-2) (2021):139-144.
- Kouhpayeh HR, Tabasi F, Dehvari M, et al. Association between angiotensinogen (AGT), angiotensin-converting enzyme (ACE) and angiotensin-II receptor 1 (AGTR1) polymorphisms and COVID-19 infection in the southeast of Iran: a preliminary case-control study. Transl Med Commun 6(1)(2021):26.
- Setti L, Passarini F, De Gennaro G et al. SARS-Cov-2RNA found on particulate matter of Bergamo in Northern Italy: First evidence. Environ Res 188(2020):109754.
- Muscat Baron Y, Camilleri L. The emergence of ten SARS-CoV-2 variants and airborne PM2.5. Virology:current research 5(6) (2021): 1-12.
- Muscat Baron Y. The Emergence of the Omicron variant and the Airborne Pollutant PM2.5 in the Gauteng Province, South Africa 2021 Malta Public Health symposium: COVID-19 a Life Changer. Impact, Challenges, Wellbeing Way Forward (2021).
- Muscat Baron Y. The Emergence of the Stealth Omicron variant and the Airborne Pollutant PM2.5 in the Kolkata. India Researchgate 158(2021).
- Giani P, Castruccio S, Anav A, et al. Short-term and long-term health impacts of air pollution reductions from COVID-19 lockdowns in China and Europe: a modelling study. Lancet Planet Health 4(10) (2020): 474-482.


 Impact Factor: * 3.6
Impact Factor: * 3.6 Acceptance Rate: 76.49%
Acceptance Rate: 76.49%  Time to first decision: 10.4 days
Time to first decision: 10.4 days  Time from article received to acceptance: 2-3 weeks
Time from article received to acceptance: 2-3 weeks 