Advancing quality control of food samples by Next Generation Sequencing compared to culture-dependent techniques
Maria-Eleni Dimitrakopoulou, Chrysoula Kotsalou,Venia Stavrou, Apostolos Vantarakis*
Department of Medicine, University of Patras, Patra, Greece
*Corresponding Author: Apostolos Vantarakis, Department of Medicine, University of Patras, Patra, Greece
Received: 16 April 2021; Accepted: 27 April 2021; Published: 21 May 2021
Article Information
Citation:
Maria-Eleni Dimitrakopoulou, Chrysoula Kotsalou,Venia Stavrou, Apostolos Vantarakis. Advancing quality control of food samples by Next Generation Sequencing compared to culture-dependent techniques. Journal of Food Science and Nutrition Research 4 (2021): 118-130.
DOI: 10.26502/jfsnr.2642-11000066
View / Download Pdf Share at FacebookAbstract
Food quality and safety had become an increasing issue for quality authorities, food scientists, manufacturers, producers and consumers as well. In the present work, food quality control using bacterial identification isolated from food samples was performed. Greek olives (raw drupes), currants (dry fruits), roe (fish eggs), bacon and “feta” cheese were collected for this purpose. The aim of this study was to investigate the quality of traditional Greek food types and to compare bacterial population observed of each food matrix according to analysis method. Therefore, culture based and molecular methods were performed in parallel. Total bacterial DNA isolated from food samples was verified by PCR and PCR-RAPD amplification and then analyzed by Next Generation Sequencing. Results showed that NGS analysis can provide an advanced and detailed look at food microbiota in comparison to culture-dependent techniques. Climate conditions, environmental factors, even production or processing procedures can result an abundant bacterial diversity isolated from each food product, so NGS approaches can be suitable to elucidate the whole picture of bacteria populations. NGS analysis regarding food safety management can be applied in food industries and therefore quality and safety of food products can be reassured.
Keywords
<p>NGS, Bacteria, Food quality, Food safety, Quality control</p>
Article Details
1. Introduction
Both food safety and food quality consitute essential need for food products for consumers, food industries, producers and quality authorities as well. According to consumers’ requirements regarding food quality, food safety considered to be crucial [1]. A major and important area of challenges regarding food safety, is microbiological safety of foodstuff [2]. In 2010, 600 million incidents of foodborne infections were reported by WHO which lead to 420,000 deaths [3]. Thus, nowdays, due to the increasing interest of consumers, food quality systems tend to be more and more demanding, following new regulations and techniques [4]. The most popular and common method for food quality audit is a culture-dependent method, which allows analysis of live microbial cells after their selective isolation from microbial population by inoculating media [5]. However, this standard method of food analysis face up some limitations such as low sensitivity, low accuracy or time-consuming process [6,7]. Moreover, due to the presence of viable but non- culturable cells , culture-dependent methods can not be considered as an appropriate tool for analyse and study total microorganisms’ population of samples [8]. Therefore, culture-independent techniques for food analysis have been developed. In 1999, for the first time, culture-independent approaches presented in fermented food analysis, in terms of food microbiology [9]. Molecular PCR-based techniques are already used to characterize microorganisms in food, regarding fermentation, spoilage and contamination [10]. As far as these PCR-based techniques concern, real time-PCR, reverse transcription-PCR, Loop Mediated Isothermal Amplification (LAMP) approaches have currently widely reported as tools for food quality and safety issues [11,12]. Notably, a new research tool, known as Next Generation Sequencing, has been widely used in a variety of fields such as microbiology, diagnostics, forsenics etc. By Next Generation Sequencing method, total microbiome of a biological sample can be identified and sequenced. In the field of foodomics, genomic analysis of food samples by NGS considered to be high accurate, rapid and much more informative compared with traditional culture depenedent techniques [13]. Next Generation Sequencing applications offer in depth taxonomic identification of total microorganism in a food sample, regardless matrix’ complexity. Moreover, by metagenomic analysis of a food sample, actions such as recalls due to a foodborn pathogen detection or characterization of viable but non-culturable microorganisms, can easily be performed [14]. Food pathogens such as Escherichia coli (E. coli), Campylobacter jejuni (C. jejuni), Listeria monocytogenes (L. monocytogenes), Shigella spp., Salmonella spp., Staphylococccus aureus, Vibrio spp, Bacillus cereus, or, food spoilage microorganisms such as Acinetobacter spp., Pseudomonas spp., Botrytis spp. can be detected in a variety of food products and are responsible for major foodborn illnesses [15-17]. Therefore, there are publications that already have utilized NGS for understanding the ecology of food fermentation, for biodiversity analysis of spoilage microorganisms in poultry meat, or studying microbiota of wine and soya beans paste [18-21]. The aim of this study was to examine and characterize bacterial diversity of five complex and different food types in terms of food quality and safety aspect. For this purpose, we compare and evaluate standard culture-dependent methods for bacteria analysis of each food matrix with a molecular technique, Next Generation Sequencing.
2. Materials and Methods
2.1 Samples collection
For this experinment, five greek food products from totally different category of food types and process level were chosen. A raw fruit, a dry fruit, a dairy, an animal product and a seafood were examined. In particular, fresh olives (Olea europaea L.cv Kalamon), dry vine products (Corinthian currants), cheese (“feta” PDO), smoked bacon and fish eggs (avgotaracho Mesolonghiou PDO) were purchased from local producers. Natural Greek black table olives from Kalamon variety considered to be the most famous variety and characterized by their high nutritional value [22]. Corinthian currants(Vitis Vinifera L., var. Apyrena) are cultivated in Southern Greece and are essential component in Mediterenian diet, due to their high antioxidant content [23]. “Feta” is a traditional Greek industrial cheese produced by pasteurized sheep’s and goat’s milk (up to 30%) in well-equipped cheese dairies, using rennin enzyme and commercial lactic acid cultures as starters [24]. Bacon from selected fine pork pancetta has a rich flavour and it was tradiotionally smoked with beech wood. Avgotaracho Mesolonghiou is known as “Greek caviar” and has a high commercial value, due to its unique aroma and flavor [25]. “Feta” cheese and bacon were processed and packaged, while the three other food products were collected raw.
2.2 Culture-dependent approach
Ten grams of each food sample were homogenized with 90ml sterile Buffer Peptone Water (10-1 dilution). A series of 5 fold dilutions were prepared and spread plate technique was followed on appropriate selective media. Medium for bacterial growth were chosen according to ISO 17025:2017. For detection of Salmonella species 25 ml of the food sample were diluted with 225 ml of sterile Buffer Peptone Water and mixed well. Microbiological analysis included enumeration and identification of potential pathogens conducted according to standard procedures for the number of LAB, E coli, Listeria monocytogenes, Enterobacteriaceae, Staphylococcus, Salmonella and Total mesophilic flora. Appropriate dilutions were then enumerated for these bacterial categories. All plates were incubated under aerobic conditions except from MRS and VRBG plates, that need an extra layer after spread plating, for anaerobic conditions. The mean number of colonies counted was expressed as colony forming units (cfu)/gr, according to each ISO barring Salmonella and Listeria, whose boundaries were determined by their presence or absence.
2.3 Bacterial DNA extraction-PCR amplification
Bacterial DNA extraction protocol was performed by boiling method as described by Ribiero[26].Bacterial Genomic DNA extracted from each food sample, was used as template for PCR amplification. To amplify 240bs from V3 region of 16s rRNA gene, the following universal primers: 338F (5′ACTCCTACGGGGGCAGCAG,Sigma,France), 518R(5′ATTACCGCGGCTGCTGG, Sigma, France) were used [25]. The PCR reaction was carried out in a final volume of 25μl, containing 100mg DNA template, 5μL 5xbuffer C Mg free, 1,5μL MgCl2(25mM), 0,5μL dNTPs(10mM), 5μL primers(1μΜ) and 0.1μL Taq polymerase (Kapa TAq PCR kit) (Sigma, France). PCR amplification was performed using Thermocycler (Biorad). The amplification program was carried out as follows: initial denaturation 95°C for 3mins, 30 cycles of denaturing at 95°C for 1min, annealing at 55°C for 1min, extension at 72°C for 1min with a final extension at 72°C for 10mins. The resulting amplicons of PCR were visualised by electrophoresis in 2% (w/v) agarose gel and were stained with 8μL of GelRed(Biotium).
2.4 PCR- Random Aplification of Polymorphic DNA
After verifying the amplicon (240bs) from each food product, extracted DNA were amplified by PCR-RAPD in order to obtain information about the bacteria communities each product. The following primers were tested for PCR-RAPD amplification: M13: 5’-GAGGGTGGCGGTTCT-3’, 1247:5’-AAGAGCCCGT-3’, 1290: 5’- GTGGATGCGA-3’, OPA10: 5’ GTGATCGCAG-3’, OPA15: 5’- TTCCGAACCC-3’[27-30]. PCR mixtures contained 1x PCR buffer, 5mmoll Mgcl2, 200μmoll-1 dNTPs, 2μmol primer, 1.25 Taq polymerase (Life Technology) and 50 ng bacterial DNA. PCR amplification was performed with a thermal cycler under following conditions : initial denaturation at 94°C for 2 min, followed by 35 cycles of denaturation at 94°C for 1 min, annealing at 45°C for 40s, elongation at 72°C for 2 min and final extension at 72°C for 10 min. PCR products were separated by electrophoresis 2% agarose gel and DNA ladder was used as DNA molecular weight marker.
2.5 Next Generation Sequencing
Bacteria community profiles of food samples were assessed by 16S rRNA gene sequencing using the Illumina technology .More specific, bacterial DNA extracted from food samples were sequenced using an Illumina MiSeq (Illumina RTA v1.17.28; MCS v2.2). Sequences observed from these samples were downloaded from the BaseSpace website.
3. Results and Discussion
3.1 Microbial cultures
Culture-dependet approach as a gold standard method for quality control of food products was performed (Table 1). shows the results from this gold standard method according to ISO17025:2017. Listeria monocytogynes and Salmonella spp. were not detected in any of food products. However, enamuration of total mesophilic flora in food products consider to be essential, as it is an indicator of products’ production, storage or transportation conditions [31].
3.2 PCR-RAPD
PCR-RAPD analysis for each food product was performed in order to examine whether there was a diversity in bacteria communities among samples. Therefore, five different primers were examined for this purpose. Primer OPA15 resulted the most informative PCR-RAPD band pattern profile (Figure 1).
3.3 Next Generation Sequencing
Within each different food type, abundance precentages for dominant OTUs in the species classification are presented. For instance, dominant bacteria strain identified in fish eggs samples were Acinetobacter venetianus, while in bacon were Photobacterium phosphoreum. As far as fruits concern, Pantoea agglomerans was detected in olives and Bacilus cereus in currants. Regarding “feta” cheese, appeared to be dominated by Lactococcus sp. (Figures 2-5).
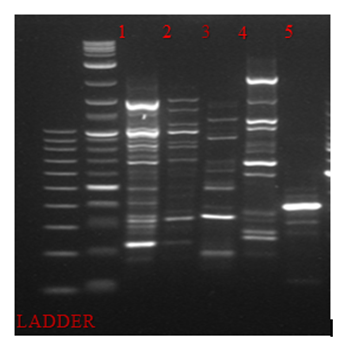
Figure 1: PCR-RAPD analysis of 16srRNA of bacterial DNA isolated from 1.fish eggs 2.olives 3.bacon 4.feta cheese 5.currants

Figure 2: Next generation sequencing analysis of 16srRNA of bacteria communities of fish eggs. Red highlighted : Dominant species Yellow highlighted: Next most abundant species.
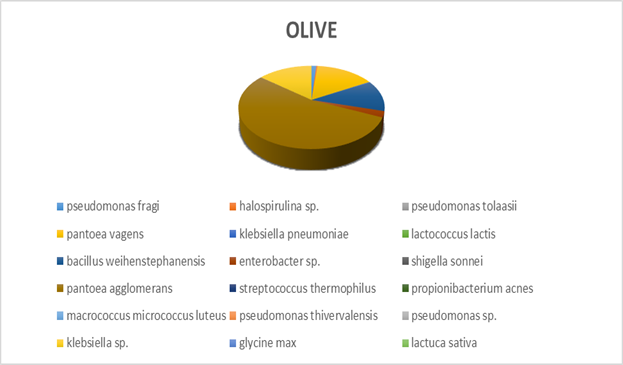
Figure 3: Next generation sequencing analysis of 16srRNA of bacteria communities of olive. Red highlighted: Dominant species Yellow highlighted: Next most abundant species
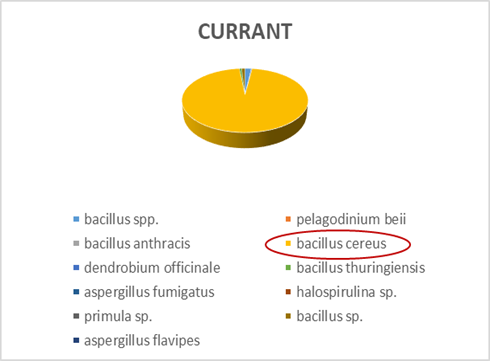

Figure 4: Next generation sequencing analysis of 16srRNA of bacteria communities of currants. Red highlighted: Dominant species

Figure 5: Next generation sequencing analysis of 16srRNA of bacteria communities of “feta” cheese. Red highlighted: Dominant species
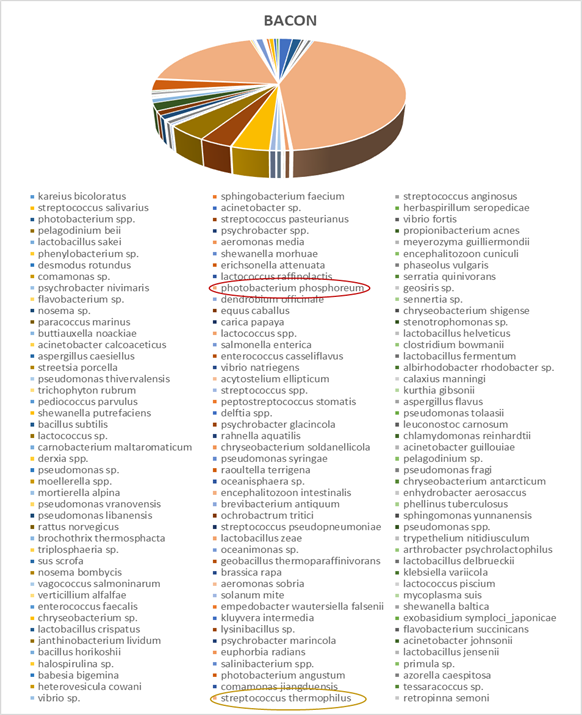
Figure 6: Next generation sequencing analysis of 16srRNA of bacteria communities of bacon. Red highlighted: Dominant species Yellow highlighted: Next most abundant species
4. Discussion
This study provides a comparison of culture-dependent analysis to NGS survey in order to characterize existing bacterial population isolated from each food type. At first, gold standard method for quality control of food samples was conducted. From the results obtained, in none of these food products examined, Salmonella spp. and Listeria Monocytogenes were detected. The total concentration of mesophilic flora ranged from 104 to 108 in all food samples. Except bacon sample, Enterobacteriaceae species were abundant too, in all food products. The highest percentage of LAB bacteria (108) was detected in “feta”cheese, as expected. PCR-RAPD amplification following, revealed high biodiversity in all food samples examined. RAPD fingerprinting pattern of currant had less strain variation compared with the strain variation obtained from the other food products. RAPD-PCR analysis of all food samples produced patterns with products ranging from approximately 100 bp to more than 1500 bp. However, NGS analysis enabled in depth identification of total bacteria communities of each food product, at the level of species.More specific, the dominant precentage of bacteria species found in fish eggs samples, at 86%, was Acinetobacter venetianus. Genus Acinetobacter has been reported as a pathogen bacteria for seafood, such as shrimps, catfsih or snout bream [32-34]. Acinetobacter species such as A. beijerinckii, A. gyllenbergii, A. haemolyticus, A. junii, A. parvus, A. tjernbergiae and A. venetianus, can induce hemolytic reactions and lyse mammalian red blood cells [35,36]. Macrococcus micrococcus luteus observed by fish eggs’ bacteria analysis, is present due to dietary control of fishes [37]. The highest precentage of bacteria communities analysis in olives belongs to Pantoea agglomerans or Enterobacter agglomerans. Pantoea agglomerans is a gram-negative aerobic bacillus of the family Enterobacteriaceae. There are publications that have associated presence of Pantoea agglomerans with Pseudomonas savastanoi pv. Savastanoi, which is responsible for olive knot diseace [38,39]. Dominant species detected in currants, were Bacilus cereus. Bacilus cereus strains considerd to be an emerging foodborn pathogen, as it can cause gastrointestinal diseases and is characterised by its spores resistance in heat and acid [40]. Bacilus cereus , which can be present in a variety of foodstuff , is a a gram-positive bacterium and at 103-105 concentration produces an emetic toxin known as cereulide [41]. As far as “feta” cheese concerns, Lactococcus sp. apperared in the most reads in NGS analysis. The microbial composition of “Feta” cheese was dominated by LAB as expected due to their presence in sheep and goat milk. Moreover, Lactococcus sp. is used as a starter culture for acid development during cheese production [42]. In bacon sample analysis, Photobacterium phosphoreum strain was abundant. Genus Photobacterium belongs to the family Vibrionaceae and is known as a fish spoilage, as it is usually isolated from aquaculture environment. Recently, Photobacterium phosphoreum was characterized as a spoilage bacterium, common in modified packaged meat [43,44]. NGS analysis revealed identification of more existing bacteria in food samples examined. According to NGS results obtained, it is essential need to implement this appoach for food quality control, as the existing bacteria strains identified were emerging pathogens. However, based on Greek standards for food quality and safety predominantly followed in food industry, main pathogen microorganisms responsible for food illnesses are tested by culture-dependent techniques. The existing protocols for microbiological audits by standard culture-dependent methods face up some limitations. NGS can overcome these limitations and provide detailed look at the microorganisms extracted from food samples. Interestingly, our findings are in agreement with other publications , such as NGS analysis for determination of fish flesh or milk microbiota, which report the superiority of NGS over culture-dependent methods [45,46]. Moreover, “omics” technologies can have a potential impact not only in food quality and safety, but also in food traceability and authentication. By NGS analysis, data for enhancement of “food traceability” or “food authentication” issues can be also collected [47,48]. Nevertheless, NGS application can provide information not only regarding food industry but also for agricultural practices such as sustainability, plant diseases and irrigation [49]. NGS approaches can be an additional tool to obtain information for marine environments, as well [50]. Thus, advancing NGS technologies could address important issues related to a variety of fields.
5. Conclusion
This study presents a first look at the microflora of five greek food products using Next generation Sequencing in correlation to gold standard methods. NGS approach resulted to the rapid identificiation of more strains of bacteria species than traditional methods.NGS application can provide knowledge of total microorganisms isolated from food products, regarding incidents of spoilage, foodborn diseases or fermentation processes. In this study, results from NGS analysis validated the usefullness of this approach in order to detect and identify total microbiota of a sample, in comparison to standard culture-dependent methods. Thus, NGS analysis can be implemented in food industry or established as the main protocol of microbiological quality control for quality authorities. NGS can also enhanche agricultural and aquacultural treatments, as the information obtained from NGS are associated with all the stages in a supply chain.
Acknowledgments
The present research received fund from the Single State Action Aid for Research, Technological Development & Innovation «INVESTIGATE - CREATE - INNOVATE" project "Trust Trace" T1EDK-04028. We also acknowledge support of this work by the project “Synthetic Biology: From omics technologies to genomic engineering (OMIC-ENGINE)” (MIS 5002636) which is implemented under the Action “Reinforcement of the Research and Innovation Infrastructure”, funded by the Operational Program "Competitiveness, Entrepreneurship and Innovation" (NSRF 2014-2020) and co-financed by Greece and the European Union (European Regional Development Fund).
Conflicts of Interest
The authors declare no conflict of interest.
References
- Peri C. The Universe of Food Quality. Food Qual. Prefer 17 (2006): 3-8.
- Fung F, Wang HS, Menon S. Food Safety in the 21st Biomed J 41 (2018): 88-95.
- World Health Organization (WHO). Food-Borne Disease Burden Epidemiology Reference Group. Encycl. Parasitol (2015): 1-1.
- Kotsanopoulos KV, Arvanitoyannis IS. The role of auditing, food safety, and food quality standards in the food industry: A Review. Compr Rev Food Sci Food Saf 16 (2017): 760-775.
- Davis C. Enumeration of Probiotic Strains: Review of Culture-Dependent and Alternative Techniques to Quantify Viable Bacteria. J Microbiol. Methods 103 (2014): 9-17.
- Adewumi GA, Oguntoyinbo FA, Keisam S, et al. Combination of Culturf-Ndependent and Culture-Dependent Molecular Methods for the Determination of Bacterial Community of Iru, a Fermented Parkia Biglobosa Seeds. Front Microbiol 3 (2012).
- Lee HW, Yoon SR, Kim SJ. et al. Identification of Microbial Communities, with a Focus on Foodborne Pathogens, during Kimchi manufacturing process using culture-independent and dependent analyses. Lwt 81 (2017): 153–159.
- Zhao X, Zhong J, Wei C et al. Current Perspectives on viable but non-culturable state in foodborne pathogens. Front Microbiol 8 (2017).
- Ampe F, Ben Omar N, Moizan C. et al. Polyphasic study of the spatial distribution of microorganisms in mexican pozol, a fermented maize dough, demonstrates the need for cultivation-independent methods to investigate traditional fermentations. Appl Environ Microbiol 65 (1999): 5464-5473.
- Justé A, Thomma BP, Lievens B. Recent advances in molecular techniques to study microbial communities in food-associated matrices and processes. Food Microbiol 25 (2008): 745-761.
- Postollec F, Falentin H, Pavan S. et al. Recent advances in quantitative PCR (QPCR) applications in food microbiology. Food Microbiol 28 (2011): 848-861.
- Hameed S, Xie L, Ying Y. Conventional and emerging detection techniques for pathogenic bacteria in food science: A Review. Trends Food Sci. Technol 81 (2018): 61-73.
- Jagadeesan B, Gerner-Smidt P, Allard MW, et al. The use of next generation sequencing for improving food safety: Translation into practice. Food Microbiol 79 (2019): 96-115.
- Leonard SR, Mammel MK, Lacher DW, et al. Application of metagenomic sequencing to food safety: detection of shiga toxin-producing Escherichia Coli on fresh bagged spinach. Appl Environ Microbiol 81 (2015): 8183-8191.
- Maruši? A. Food Safety and Security: What Were Favourite Topics for Research in the Last Decade? J Glob Health 1 (2011): 72-78.
- Al-Kharousi ZS, Guizani N, Al-Sadi AM, et al. Hiding in fresh fruits and vegetables: Opportunistic pathogens may cross geographical barriers. Int J Microbiol (2016).
- Bintsis T. Foodborne Pathogens. AIMS Microbiol 3 (2017): 529-563.
- Christoph, Heinrich, Candela P. Next-Generation Approaches to the Microbial Ecology of Food Fermentations. Brisk Bin. Robust Invariant Scalable Keypoints 45 (2014).
- Lee HS, Kwon M, Heo S, et al. Characterization of the biodiversity of the spoilage microbiota in chicken meat using next generation sequencing and culture dependent approach. Korean J Food Sci Anim Resour 37 (2017): 535-541.
- Nam Y, Lee SY, Lim S. Microbial community analysis of korean soybean pastes by next-generation sequencing. Int J Food Microbiol 155 (2015): 36-42.
- Bokulich NA, Joseph CML, Allen G, et al. Next-generation sequencing reveals significant bacterial diversity of botrytized wine. Plos one 7 (2012): 3-12.
- Papadelli M, Zoumpopoulou G, Georgalaki M, et al. Evaluation of two lactic acid bacteria starter cultures for the fermentation of natural black table olives (Olea Europaea L Cv Kalamon). Polish J. Microbiol 64 (2015): 265-271.
- Yanni AE, Efthymiou V, Lelovas P, et al. Effects of Dietary Corinthian Currants (Vitis Vinifera L., Var. Apyrena) on atherosclerosis and plasma phenolic compounds during prolonged hypercholesterolemia in New Zealand White Rabbits. Food Funct 6 (2015): 963-971.
- Manolopoulou E, Sarantinopoulos P, Zoidou E, et al. Evolution of microbial populations during traditional feta cheese manufacture and ripening. Int J Food Microbiol 82 (2003): 153-161.
- Dimitriou E, Katselis G, Moutopoulos DK, et al. Description of the processing stages of a protected designation of origin fish product: The Greek Caviar “Avgotaracho Messolongiou.” Agric Econ Rev 17 (2016): 50-62.
- Ribeiro JC, Tamanini R, Soares BF, et al. Efficiency of boiling and four other methods for genomic DNA extraction of deteriorating spore-forming bacteria from milk. Semin Agrar 37 (2016): 3069-3078.
- Huey B, Hall J. Hypervariable DNA fingerprinting in Escherichia Coli: Minisatellite Probe from Bacteriophage M13. J Bacteriol 171 (1989): 2528-2532.
- Venieri D, Vantarakis A, Komninou G, et al. Differentiation of faecal escherichia coli from human and animal sources by random amplified polymorphic DNA-PCR (RAPD-PCR). Water Sci. Technol 50 (2004): 193-198.
- Thomas PC. Genetic characterization of aeromonas hydrophila using protein profiling and RAPD PCR. Asian Fish Sci 22 (2009): 763-771.
- Koche M, Gade R, Kothikar R, et al. Biochemical studies on genotypic characterization of pseudomonas fluorescens isolates by PCR-RAPD Analysis. Int J Chem Stud 8 (2020): 2915-2917.
- Mcmahon W, Mills J, Mohnke F, et al. In Foods: Collaborative Study 54 (2009): 165-174.
- Huang X, Gu Y, Zhou H, et al. Acinetobacter Venetianus, a potential pathogen of red leg disease in freshwater-cultured whiteleg shrimp penaeus vannamei. Aquac Reports 18 (2020).
- Mires D. Acinetobacter Lwoffii: An Emerging pathogen for red head disease in farmed channel catfish ictalurus punctatus (2010).
- Cao H, Yu L, Ou R, et al. Acinetobacter Johnsonii: An emerging pathogen for cultured blunt snout bream megalobrama amblycephala. Isr J Aquac Bamidgeh 69 (2017).
- Gerner-Smidt P, Tjernberg I, Ursing J. Reliability of phenotypic tests for identification of Acinetobacter Species. J Clin Microbiol 29 (1991): 277-282.
- Vaneechoutte M, Nemec A, Musílek M, et al. Description of Acinetobacter Venetianus Ex Di Cello et Al. 1997 Sp. Nov. Int. J. Syst. Evol. Microbiol 59 (2009): 1376-1381.
- Askarian F, Zhou Z, Olsen RE, et al. Culturable Autochthonous gut bacteria in Atlantic Salmon (Salmo Salar L.) Fed Diets with or without Chitin. Characterization by 16S RRNA gene sequencing, ability to produce enzymes and in vitro growth inhibition of four fish pathogens. Aquaculture 9 (2012): 326-329.
- Marchi G, Sisto A, Cimmino A, et al. Interaction between Pseudomonas Savastanoi Pv. Savastanoi and Pantoea Agglomerans in Olive Knots. Plant Pathol 55 (2006): 614-624.
- Quesada JM, García A, Bertolini E, et al. Recovery of Pseudomonas Savastanoi Pv. Savastanoi from symptomless shoots of naturally infected Olive Trees. Int Microbiol 10 (2007): 77-84.
- Dietrich R, Jessberger N, Ehling-Schulz M, et al. The food poisoning toxins of Bacillus Cereus. Toxins (Basel) 13 (2021).
- Agata N, Ohta M, Yokoyama K. Production of Bacillus Cereus Emetic Toxin (Cereulide) in various foods. Int J Food Microbiol 73 (2002): 23-27.
- Rantsiou K, Urso R, Dolci P, et al. Microflora of feta cheese from four Greek manufacturers. Int J Food Microbiol 126 (2008): 36-42.
- Hilgarth M, Fuertes-Pèrez S, Ehrmann M, et al. An adapted isolation procedure reveals Photobacterium Spp. as common spoilers on modified atmosphere packaged meats. Lett Appl Microbiol 66 (2018): 262-267.
- Hauschild P, Hilgarth M, Vogel RF. Hydrostatic Pressure- and Halotolerance of Photobacterium Phosphoreum and P. Carnosum Isolated from Spoiled Meat and Salmon. Food Microbiol 10 (2020): 103679.
- Tsironi T, Lougovois V, Simou VN, et al. Next Generation Sequencing (NGS) for the determination of fish flesh microbiota. J Food Res 8 (2019): 101.
- Sattin E, Andreani NA, Carraro L, et al. A Multi-Omics approach to evaluate the quality of milk whey used in ricotta cheese production. Front. Microbiol 7 (2016): 1-13.
- Haynes E, Jimenez E, Pardo, MA, et al. The future of NGS (Next Generation Sequencing) analysis in testing food authenticity. Food Control 101 (2019): 134-143.
- Muñoz-Colmenero M, Martínez JL, Roca A, et al. NGS tools for traceability in candies as high processed food products: ion torrent pgm versus conventional PCR-Cloning. Food Chem 214 (2017): 631-636.
- Esposito A, Colantuono C, Ruggieri V, et al. Bioinformatics for agriculture in the Next-Generation Sequencing era. Chem Biol Technol Agric 3 (2016): 1-12.
- Yue GH, Wang L Current status of genome sequencing and its applications in aquaculture. Aquaculture 468 (2017): 337-347.


 Impact Factor:
* 3.8
Impact Factor:
* 3.8
 Acceptance Rate:
77.96%
Acceptance Rate:
77.96%
 Time to first decision: 10.4 days
Time to first decision: 10.4 days
 Time from article received to acceptance:
2-3 weeks
Time from article received to acceptance:
2-3 weeks
