Comparative Biochemical Characterization of L-Asparaginases from Four Species of Lactic Acid Bacteria
Kodchakorn Phetsri1, Makoto Furukawa1, Risa Yamashiro1, Yuka Kawamura1, Junji Hayashi2, Ryuta Tobe1, Yosuke Toyotake1, Mamoru Wakayama1*
1Collage of Life Sciences, Ritsumeikan University, Kusatsu, Shiga, Japan
2Faculty of Bioscience and Bioindustry, Tokushima University, Tokushima, Japan
*Corresponding Author: Mamoru Wakayama, Collage of Life Sciences, Ritsumeikan University, Kusatsu, Shiga 525-8577, Japan
Received: 10 September 2019; Accepted: 25 September 2019; Published: 30 September 2019
Article Information
Citation:
Kodchakorn Phetsri, Makoto Furukawa, Risa Yamashiro, Yuka Kawamura, Junji Hayashi, Ryuta Tobe, Yosuke Toyotake, Mamoru Wakayama. Comparative Biochemical Characterization of L-Asparaginases from Four Species of Lactic Acid Bacteria. Journal of Biotechnology and Biomedicine 2 (2019): 112-124.
DOI: 10.26502/jbb.2642-91280015
View / Download Pdf Share at FacebookAbstract
L-Asparaginase (ASNase; EC 3.5.1.1) is an enzyme that catalyzes the hydrolysis of L-asparagine to L-aspartic acid and ammonia. Generally, ASNases from Escherichia coli and Erwinia chrysanthemi are used for the treatment of acute lymphoblastic leukemia. However, few studies focusing on ASNase from lactic acid bacteria (LAB) have been reported. The aim of this study is to characterize ASNase genes from four LAB strains: Streptococcus thermophiles, Lactobacillus plantarum, L. acidophilus, and L. sakei. ASNase genes from each strain amplified by polymerase chain reaction PCR were inserted into NdeI and XhoI sites of pET28a-(+) and cloned in E. coli BL21(DE3). Recombinant ASNases were purified using nickel-nitrilotriacetic acid column chromatography. Among the four strains, the purified recombinant ASNase from S. thermophilus exhibited the highest specific activity of 113.0 U/mg and specificity for L-asparagine. The pH and temperature ranges for S. thermophilus ASNase were pH 8.0-9.0 and 30°C-50°C, respectively. The activity of the enzyme was significantly inhibited by Ni2+. Km and kcat values were 2.91 mM and 1.53 × 102 s–1, respectively. In this study, we described the biochemical properties of ASNases from four LAB and demonstrated that ASNase from S. thermophilus has potential applications in food processing.
Keywords
<p>L-asparaginase, Lactic acid bacteria, Characterization, Cloning</p>
Article Details
1. Introduction
L-Asparaginase (ASNase; EC 3.5.1.1.) is an enzyme that catalyzes the hydrolysis of L-asparagine to L-aspartic acid and ammonia. Over the past 30 years, bacterial ASNases have received considerable attention for their anticancer activity in the treatment of acute lymphoblastic leukemia and lymphosarcoma after the first report was published in 1967 by Hill JM et al. [1]. This therapy is based on the ability of ASNase to catalyze the conversion of L-asparagine to L-aspartic acid. Absence of L- asparagine causes tumor cells to die [2]. Among the bacterial ASNase resources, ASNases from Pseudomonas pseudoalcaligenes JHS-71 [3], Bacillus sp. [4], Staphylococcus [5] and Serratia marcescens [6] have been studied. However, their intrinsic glutaminase activity, which could cause serious side effects like neurotoxicity, hepatitis and other dysfunctions, restricts their clinical applications [7]. In contrast, enzymes from E. coli [8] and E. carotovora [9] the most effective for this purpose. This is because ASNases from these species acquire a strong preference to asparagine over glutamine, leading to less severe immunoreactive side effects.
Apart from the significant role of ASNases in the healthcare sector, ASNases have attracted the attention of the food processing industry for reducing acrylamide [10]. Acrylamide is a carcinogenic substance that is synthesized in large quantities during the Maillard reaction, occurring during normal food processing [11]. The accumulation of acrylamide during the Maillard reaction is especially problematic during the processing of cereals and potato-based products that are fried, baked, or roasted [12]. ASNase selectively hydrolyzes L-asparagine, which is an essential precursor for the formation of acrylamide [13]. The available commercial preparations of ASNase in the food industry are based on enzymes from recombinant strains of Aspergillus niger or A. oryzae [12]. Nevertheless, the present commercial applications of ASNase are still limited.
Lactic acid bacteria (LAB) represent some of the most extensively studied microorganisms and have been widely used in food fermentation worldwide. During fermentation, the metabolic activity of LAB can change the nutritional properties of food matrices [14] with possible beneficial effects on human health. Particularly, LAB may be used as probiotics, which can help maintain good health and counteract diseases [15]. Fermentation using LAB can offer one or more organoleptic, nutritional, or health advantages because of the formation of pleasant substrates and bioactive end-products [16]. However, there is little information regarding ASNases from LAB. Obtaining biochemical and structural information about ASNases from LAB will help promote the development of potential applications in the food industry. The aim of the present study is to comparatively investigate the expression and biochemical properties of ASNases from the following four strains of LAB: S. thermophilus, L. plantarum, L. acidophilus, and L. sakei.
2. Materials and Methods
2.1 Bacterial strains
LAB were selected based on their potential applications in the food industry and phylogenetic data from the Kyoto Encyclopedia of Genes and Genomes database (KEGG). Once the four strains of LAB were selected, the ASNase activity of the each selected strain was examined. The genetic information regarding ASNases from LAB was obtained from the KEGG. The phylogenetic tree analyses of ASNase sequences were conducted using a robust online software program (http://www.phylogeny.fr/). Multiple sequence alignment was performed using BioEdit version 7.2.5. All LAB used in the present study (S. thermophilus NBRC 13957, L. plantarum NBRC 14711, L. acidophilus NBRC 13951, and L. sakei NBRC 15893) were obtained from the National Institute of Technology and Evaluation (NBRC), Japan. E. coli JM109 and E. coli BL21(DE3) were used for plasmid amplification and expression, respectively.
2.2 Cloning and expression of ASNase genes
The genomic DNAs from S. thermophilus, L. plantarum, L. acidophilus, and L. sakei were prepared for templates for the polymerase chain reaction (PCR) according to the general method [17]. The oligo primers were designed based on the nucleotide sequences of ASNases obtained from KEGG and synthesized by FASMAC Co., Japan. These primers contained NdeI and Xhol restriction sites, respectively. All the primer sequences are shown in Table 1 of Supplementary Data. The open reading frame of each ASNase was amplified by PCR using KOD-Plus-Neo Kit (TOYOBO CO., LTD (Osaka, Japan)) under the following conditions; initial denaturation at 94°C for 2 min, denaturation at 94°C for 10 s, annealing for 30 s, and the optimal temperature for all primers and extension at 68°C for 60 s for 35 cycles. The PCR fragment (approximately 1,000 bp) was digested with NdeI and Xhol (Takara Bio Inc.), and then cloned into the same sites of pET-28a(+). The expression plasmid was extracted by using FastGene® Plasmid Mini Kits (NIPPON GENE CO., LTD.) from the transformed E. coli JM109. E. coli BL21(DE3) was transformed by the expression plasmid for production of ASNase.
2.3 Purification of the recombinant ASNases
All cultivations were conducted in Luria-Bertani (LB) broth containing 20 mg/ml kanamycin. At OD600 of 0.4-0.6, 0.4 mM of isopropyl β-D-thiogalactopyranoside (IPTG) was added to the main culture and incubated for 20 h to induce gene expression. The cells were collected by centrifugation at 10,000 rpm at 4°C for 10 min. The pellets were re-suspended in 20 mM Tris-HCl (pH 8.0). Cells were disrupted by sonication and precipitation was removed by centrifugation. The supernatant was collected as crude extract and loaded into a nickel-nitrilotriacetic acid column. The column was washed with the buffer (20 mM Tris-HCl (pH 8.0), 150-200 mM NaCl, and –400 mM imidazole). The bound protein was eluted with 200 mM imidazole. The proteins in the eluted fractions were detected using SDS-PAGE, and protein concentrations were determined using a Lowry protein assay with bovine serum albumin as the standard. Molecular masses of the recombinant ASNases were determined using ProteoSEC 11/30 6-600 HR SEC Gel filtration columns (ProteinArk-The Innovation Centre, Sheffield, UK). The column was pre-equilibrated with a buffer containing 20 mM sodium phosphate (pH 8.0) and 150 mM sodium chloride. The standard curve was constructed using the following standard molecular mass proteins: thyroglobulin (669 kDa), γ-globulin (440 kDa), IgG (158 kDa), and Ovalbumin (44 kDa).
2.4 Enzyme activity assay
ASNase activity was evaluated by measuring the quantity of ammonia released during the ASNase reaction. The standard reaction mixture contained 100 mM potassium phosphate buffer (pH 7.0), 30 mM of L-asparagine, and enzyme for a final volume of 0.5 ml. After incubation at 30°C for 30 min, the reaction mixture was boiled for 4 min to quell the reaction. The supernatant was collected by centrifugation at 10,000 rpm for 4 min. Glutamate dehydrogenase (GlDH) was used to determine the quantity of released ammonia in the ASNase reaction mixture [18]. The reaction mixture contained 100 mM Tris-HCl buffer (pH 8.0), 10 mM 2-oxoglutarate, 0.24 mM nicotinamide adenine dinucleotide (NADH), 5 U GlDH, and 200 µl of the supernatant of ASNase reaction for a final volume of 1 ml. The change of absorbance at 340 nm was measured for the quantification of ammonia. One unit of enzyme activity (1 U) was defined as the amount of an enzyme required to release 1 µmol of ammonia per minute. Protein concentrations were determined using the Lowry method using crystalline bovine serum albumin as the standard [19].
2.5 Characterization of the recombinant ASNases
Properties of the purified recombinant enzymes were investigated. For the optimum temperature, the enzyme activity was measured at temperatures ranging from 30°C to 60°C. For thermal stability, the purified enzyme was incubated at temperatures varying from 20°C to 60°C for 15 min prior to performing the enzyme activity assay. For the determination of the optimum pH and pH stability of the enzymes, the following buffers were used; citrate-NaOH (pH 5.0-6.0), potassium phosphate (pH 6.0-8.0), Tris-HCl (pH 8.0-9.0), and n-cyclohexyl-2-aminoethanesulfonic acid (CHES) (pH 9.0-10.0). The enzyme activity was examined after incubating the enzyme reaction mixture in these buffers at 4°C for 20 h before performing the enzyme assay. All data are cexpressed as the average results of at least three independent experiments. The effects of metal ions, EDTA, and DTT on ASNase activity were examined in the presence of 1 mM of the following: KCl, NaCl, CaCl2, NiCl2, CuSO4, CoCl2, MgCl2, FeSO4, FeCl3, ZnCl2, MnCl2, EDTA and DTT. After incubation for 15 min at 30°C, the enzyme reaction was terminated by adding 0.4 M trichloroacetic acid solution. The supernatant solution was separated by centrifugation and the amount of released ammonia was spectrophotometrically detected at 480 nm using Nessler’s reagent.
2.6 Substrate specificity and kinetic parameters
To investigate the substrate specificity of each ASNase, the following substrates were added to the reaction mixture and incubated for 15 min at 30°C: L-asparagine, D-asparagine, L-glutamine, D-glutamine, and succinic acid. The supernatant from the enzyme mixture was used for the quantification of ammonia by using Nessler’s reagent. The experiments were conducted in triplicate. The reaction mixtures without substrate and without enzyme were prepared as controls. Km and Vmax values for each recombinant ASNase toward L-asparagine were determined by measuring the rate of ammonia released during the hydrolysis reaction using GlDH assay. The experimental data were calculated using Lineweaver-Burk plot at substrate concentration ranging from 1 to 150 mM in tris hydrochloride buffer (pH 8.0).
3. Results
3.1 Selection of bacterial strains
Thus, 16 genera of LAB were selected by considering their potential application in food processing and phylogenetic data from the KEGG. Phylogenetic analysis using the amino acid sequences of ASNases from the selected LAB strains was performed using the KEGG databases (Figure 1). ASNase activities from each strain were examined (data not shown). In the present study, ASNases from four strains were based on the phylogenetic relation and ASNase activities of the original strain: S. thermophilus, L. plantarum, L. acidophilus, and L. sakei. The compared alignments of deduced amino acid sequences of ASNases from the four strains are shown in Figure 2. The productivity of ASNases from the selected strains was evaluated (Table 1). The results showed that the supernatant from S. thermophilus (St-ASNase) had the highest specific activity for ASNase (1.29 × 10-2 U/mg). ASNase activity was detected only in the precipitate from St-ASNase (1.00 × 10-2 U/mg) and La-ASNase (3.00 × 10-2 U/mg).
3.2 Cloning of the ASNase genes and protein expression from four strains of LAB
The approximately 1,000-bp DNA fragments corresponding to ASNase genes from the four strains were obtained and cloned into the pET28a (+) vector and expressed into E. coli BL21(DE3) cells. The overexpression of recombinant ASNase was induced by adding 0.4 mM IPTG and this mixture was incubated for 20 h at 37°C. All the recombinant ASNases were attached with a 6 × His-tag at the C and N-terminals to facilitate purification using nickel column affinity chromatography. The recombinant protein was eluted with 0.2 M imidazole. The purification process for the four ASNases is summarized in Table 2 ((a)-(d)). After active fractions were collected and dialyzed, the purified enzymes were subjected to SDS-PAGE. SDS-PAGE showed that all the purified enzymes were approximately 37 kDa in size (Supplementary Data in Figure 1). Molecular masses of the enzymes estimated by size exclusion chromatography were as follows: 122 kDa (St-ASNase), 137 kDa (Lp-ASNase), 89 kDa (La-ASNase), and 159 kDa (Ls-ASNase). These results indicated that St-ASNase, Lp-ASNase, and Ls-ASNase consisted of four identical subunits while La-ASNase was dimeric. The specific activities of St-ASNase, La-ASNase, Ls-ASNase, and Lp-ASNase were 113.0, 13.50, 4.71, and 1.14 U/mg, respectively (Table 2). The specific activity of recombinant St-ASNase was significantly higher than the others.
3.3 Effect of metal ions on enzyme activity
Enzyme activities were inhibited by some of the metal ions in Figure 3. Conspicuously, Ni2+ ion strongly inhibited St-ASNase and Lp-ASNase activities and reduced the relative activities up to 35% and 32%, respectively. In contrast, Zn2+ ion inhibited Ls-ASNase activity up to 71% and Co2+ reduced the La-ASNase activity by 76%.
3.4 Effect of temperature and pH on enzyme activity
The effect of temperature on the enzyme activity was examined at different temperatures ranging from 20°C to 70°C (Figure 4A). The maximum activities for La-ASNase, Lp-ASNase, and Ls-ASNase were obtained at 40°C. However, the optimal temperature of St-ASNase was 50°C. No enzymes showed activity at 70°C. The temperature stability of the purified ASNase indicated that all enzymes except for Ls-ASNase were stable between 30°C and 50°C, while Ls-ASNase began to show decreased activity at 40°C, when the percentage of relative activity was 84% (Figure 4B). These enzymes were most active between pH 8.0 and 9.0 (Figure 4C). The pH profiles of enzyme stability are shown in Figure 4D. All enzymes were stable within the range of pH 8.0-9.0. Interestingly, the activity of La-ASNase was lower in the acidic range while compared with the other ASNases.
3.5 Substrate specificity and kinetic parameters
The substrate specificities for ASNases from the four strains of LAB were assessed. The activity toward the L-asparagine was considered as 100% activity (Figure 5). All the purified recombinant ASNases showed very low activity toward D-asparagine, L-glutamine, D-glutamine, and succinic acid. The kinetic parameters were calculated based on the constructed Lineweaver-Burk plots using the purified recombinant ASNases. The Km and Vmax values are shown in Table 3. Km values were not very different among the four enzymes, but Vmax values varied between the four ASNases. The Km and Vmax values for St-ASNase were 2.34 mM and 1.74×102 mM/min, respectively. St-ASNase had the high catalytic efficiency and a slightly higher affinity toward L-Asn compared with the other purified enzymes. Moreover, the Vm values for Lp-ASNase, La-ASNase, and Ls-ASNase were significantly lower than those of St-ASNase (Table 3).
|
Strain |
Cell weight* (mg) |
Cell-free extract |
Cell debris |
||
|
Total protein (mg) |
Specific activity (U/mg) |
Total Protein (mg) |
Specific activity (U/mg) |
||
|
Streptococcus thermophilus |
1.26 ± 0.12 |
1.61 |
1.29 × 10-2 |
1.39 |
1.00 × 10-3 |
|
Lactobacillus plantarum |
1.39 ± 0.01 |
2.54 |
3.33 × 10-3 |
5.74 |
n.d.** |
|
Lactobacillus acidophilus |
1.36 ± 0.32 |
5.98 |
2.78 × 10-3 |
2.29 |
3.00 × 10-3 |
|
Lactobacillus sakei |
1.50 ± 0.25 |
2.05 |
3.49 × 10-3 |
7.43 |
n.d.** |
One unit of enzyme activity (1 U) was defined as the amount of an enzyme required to release 1 µmol of ammonia per min; *per 100 mL of MRS medium; **n.d.; non-detected
Table 1: Comparison of ASNases from four selected lactic acid bacterial strains.
|
Purification steps |
Total protein (mg) |
Total activity (U) |
Specific activity (U mg-1) |
Yield (%) |
Purification (fold) |
|
Purification of the recombinant ASNase from S. thermophilus (St-ASNase) |
|||||
|
Crude extract |
82.9 |
168 |
2.03 |
100 |
1.0 |
|
Ni-Sepharose 6 FF |
1.03 |
117 |
113 |
69.5 |
56.0 |
|
Purification of the recombinant ASNase from L. acidophilus (La-ASNase) |
|||||
|
Crude extract |
109.0 |
625.72 |
5.74 |
100.0 |
1.0 |
|
Ni-Sepharose 6 FF |
2.52 |
14.05 |
13.50 |
5.44 |
2.35 |
|
Purification of the recombinant ASNase from L. sakei (Ls-ASNase) |
|||||
|
Crude extract |
165.86 |
135.05 |
0.81 |
100.0 |
1.0 |
|
Ni-Sepharose 6 FF |
3.40 |
16.03 |
4.71 |
11.9 |
5.80 |
|
Purification of the recombinant ASNase from L. plantarum (Lp-ASNase) |
|||||
|
Crude extract |
250 |
17.4 |
0.07 |
100 |
1 |
|
Ni-Sepharose 6 FF |
1.92 |
2.20 |
1.14 |
12.6 |
16.4 |
Table 2: Purification of recombinant ASNases from four lactic acid bacteria.
|
The recombinant L-ASNase |
Km (mM) |
Vmax (mM min-1) |
kcat (sec-1) |
kcat/Km (mM-1sec-1) |
|
St-ASNase |
2.34 |
1.74 × 102 |
1.01 × 102 |
43.0 |
|
Lp-ASNase |
7.85 |
0.406 |
2.33 |
0.30 |
|
La-ASNase |
6.23 |
0.651 |
0.51 |
8.27×10-2 |
|
Ls-ASNase |
7.64 |
7.10 |
4.09 |
0.54 |
One unit of enzyme activity (1 U) was defined as the amount of an enzyme required to release 1 µmol of ammonia per min.
Table 3: Comparison of kinetic parameters of recombinant ASNase from four lactic acid bacteria.
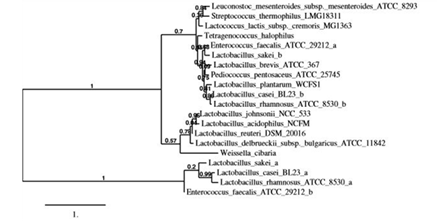
Figure 1: Phylogenetic tree of L-asparaginases from lactic acid bacteria. Phylogenetic tree was constructed based on the phylogenetic analysis of ASNase sequences of selected lactic acid bacteria by using a robust online software (http://www.phylogeny.fr/). Complete amino acid sequences deposited in KEGG database (https://www.genome.jp/kegg/kegg_ja.html) were used.

Figure 2: Comparative multiple sequence alignment of amino acid sequences from ASNases from four lactic acid bacteria at 70% of shade threshold based on KEGG database; (1) Streptococcus thermophilus CNRZ1066: str1724; (2) Lactobacillus acidophilus 30S: LAC30SC_09125; (3) Lactobacillus sakei LCA_0347; (4) Lactobacillus plantarum WCFS1: lp_2738, using BioEdit sequence alignment editor ver 7.2.5.
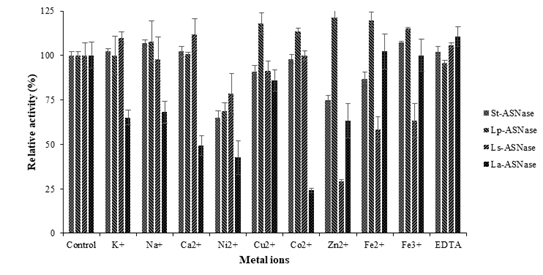
Figure 3: The effect of metal ions on ASNases activity from four lactic acid bacteria.
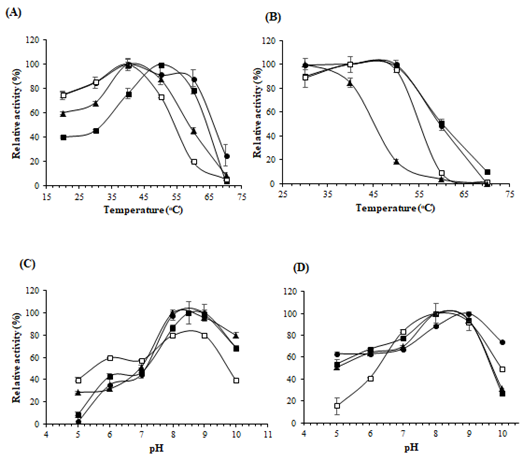
Figure 4: Effects of pH and temperature on enzyme activity and stability. (A) Optimum temperature; (B) Thermostability; (C) Optimum pH; (D) Stability of pH of the enzyme. The symbols used are as follows: St-ASNase (filled squares), Lp-ASNase (filled circles), La-ASNase (no filled squares), and Ls-ASNase (filled triangles)
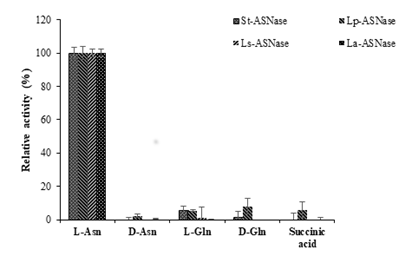
Figure 5: Comparison of substrate specificities of ASNases from four lactic acid bacteria.
4. Discussion
ASNases could be derived from various resources such as eukaryotic and prokaryotic organisms. Microbial ASNases, especially those derived from E. coli [20] or E. carotovora [21] have been effectively used in pharmaceutical industries. Nevertheless, the hypersensitivity reactions have been observed because of the production of anti-asparaginase antibody. Antibody production has been observed in up to 60% of patients at some time during ASNase therapy [22]. In the food industry, commercially used ASNase products for acrylamide reduction are obtained from A. niger and A. oryzae under the brand names PreventASeTM and Acrylaway®, respectively [23]. Interestingly, LAB also play an important role in the food, agricultural and pharmaceutical industries. The EFSA’s “Panel on Biological Hazards” has reported that LAB, which are associated with fermented food, are not associated with any clinical problems [24]. This may be because of their safe metabolic activity in foods and production of organic acids and other beneficial metabolites. Moreover, their general occurrence in foods along with widespread use means that they are Generally Recognized as Safe for human consumption [25]. Therefore, LAB may become the new safe and beneficial source of ASNases for industrial applications, especially in food processing. In the present study, we cloned ASNase genes from four strains of LAB and characterized the recombinant enzymes to determine the properties of ASNases. LAB strains were selected based on their phylogenetic analysis and activities of the ASNase in the original strains. Because of the selected bacterial strains, only activities specificcally from St-ASNase and La-ASNase were slightly detected in the precipitate. ASNase is presumably located in the periplasmic region or cytosol [26]. The detected activity may be derived from the intact cells, which may have remained due to incomplete sonication.
Because the expression plasmid contained the gene for ASNase fused to N- and C-terminal His-6 tag, the purification was completed in a one-step procedure using affinity chromatography. All the ASNase genes have been expressed using the exactly same regulation system for expression, i.e. inserting the coding region of ASNase into NdeI and XhoI sites of pET-28a(+) in E. coli BL21(DE3). However, the productivity of the enzyme varied according to the strains. A moderate amount of St-ASNase and Lp-ASNase proteina was observed in the soluble fraction. However, in the case of Ls-ASNase, a small amount of protein was found in the soluble fraction. La-ASNase was highly detected in the soluble as well as insoluble fraction. Because the size of each subunit is approximately 40 kDa, the difference in productivity of the enzyme may not be derived from the size of the enzyme, but from the other factors such as amino acid composition and sequence. In the present study, most of these recombinant ASNases had a tetrameric structure, which is probably classified into type II ASNase group, which includes the such as periplasmic enzyme from E. coli. In contrast, the molecular mass of La-ASNase was approximately 89 kDa. La-ASNase presumably has a dimeric structure that puts it in the type I ASNase group, which contains the cytosolic enzyme from E. coli [26].
The biochemical properties of the recombinant of ASNases from four LAB strains were comparatively evaluated. The effect of temperature on the activities of the enzymes was observed from 20°C to 70°C. Among the ASNases studied, St-ASNase showed a maximum activity at 50°C. This value is higher than the reported optimum temperature ASNase from L. reuteri DSM 20016, which showed a reduction in activity after 40°C and no activity above this temperature [27]. St-ASNase showed almost the same thermal stability as the recombinant ASNase from B. subtilis (BsAIID49M) [28] and type II ASNase from E. coil EcAII [29]. The thermal stabilities of St-ASNase, La-ASNase, and Lp-ASNase were in the range of 40°C to 50°C. These enzymes can maintain stability at the higher temperatures compared with the recombinant ASNase from Rhizomucor miehei [29] which showed no activity following incubation at 45°C and P. fluorescens, which was stable at 37°C [30].
The effect of pH on enzyme activity and stability was analyzed in the range of 5.0-10.0. All the enzymes showed similar pH dependency in activity and stability in the pH range of 8.0-9.0. The range of the pH value is closer to that of Thermococcus kodakarensis KOD1 [31], Enterobacter cloacae [32], P. aeruginosa [33] and E. coli [34] which had ASNases with optimal activity at pH 8.0. For the ASNases studied in the present study, no remarkable effect of most metal ions on enzyme activity was observed. However, all the enzymes from LAB described here were inhibited by Ni2+.
5. Conclusion
ASNases are widely distributed across various microorganisms but ASNases from LAB have not been widely studied. This study examines and characterizes the biochemical properties of ASNases from four different LAB strains. Among them, St-ASNase may be a good candidate for food processing, specifically for reducing acrylamide in food products. However, this study on ASNase from LAB merely a preliminary step. There is still little information on the biochemical properties and functions of ASNases in LAB. Much information regarding ASNase from LAB will increase the feasibility of the application of this enzyme in human health and food industries.
Acknowledgments
This work was financially supported by The Japan Food Chemical Research Foundation and The Ministry of Education, Culture, Sports, Science and Technology in Japan (MEXT).
References
- Hill JM, Roberts J, Loeb E, et al. L-Asparaginase therapy for leukemia and other malignant neoplasms. Remission in human leukemia JAMA 202 (1967): 882-888.
- Kotzia GA, Labrou NE. L-Asparaginase from Erwinia Chrysanthemi 3937: cloning, expression and characterization. Journal of Biotechnology 127 (2007): 657-669.
- Arastoo BD. L-Asparaginase production in Pseudomonas pseudoalcaligenes strain JHS-71 isolated from Jooshan Hot-spring. Molecular Biology Research Communications 5 (2016): 1-10.
- Vidhya M, Aishwarya R, Alagarsamy S, et al. Production, purification and characterization of extracellular L-asparaginase from a soil isolate of Bacillus sp. African Journal of Microbiology Research 4 (2010): 1862-1867.
- Han S, Jung J, Park W. Biochemical characterization of L-asparaginase in NaCl-tolerant Staphylococcus sp. OJ82 isolated from fermented seafood. Journal of Microbiology and Biotechnology 24 (2014): 1096-1104.
- John WB, Arthur WP. Purification and properties of L-asparaginase from Serratia marcescens. Journal of Bacteriology 106 (1971): 578-587.
- Jamie K, James M, Stefan F. Safety, efficacy, and clinical utility of asparaginase in the treatment of adult patients with acute lymphoblastic leukemia. Onco Targets and Therapy 10 (2017): 1413-1422.
- Khushoo A, Pal Y, Singh BN, et al. Extracellular expression and single step purification of recombinant Escherichia coli L-asparaginase II. Protein Expression and Purification 38 (2004): 29-36.
- Warangkar SC, Khobragade CN. Purification, characterization, and effect of thiol compounds on activity of the Erwinia carotovora L-asparaginase. Enzyme research (2010).
- Onishi Y, Prihanto AA, Yano S, et al. Effective treatment for suppression of acrylamide formation in fried potato chips using L-asparaginase from Bacillus subtilis. 3 Biotech 5 (2015): 783-789.
- Fernanda F, Gonçalves D, Stanislau BJ, et al. Acrylamide mitigation in French fries using native L-asparaginase from Aspergillus oryzae CCT 3940. Food Science and Technology (2017): 222-229.
- Claus A, Weisz GM, Schieber A. Acrylamide in cereal products: a review. Journal of Cereal Science 47 (2008): 118-133.
- Thomas MA, Schobachler B, Escher F, et al. Acrylamide in gingerbread: critical factors for formation and possible ways for reduction. Journal of Agricultural and Food Chemistry 51 (2014): 4282-4288.
- Zieli?ska D, Kolo?yn-Krajewska D. Review Aritcle; Food-origin lactic acid bacteria may exhibit probiotic properties. BioMed Research International 159 (2018).
- Markowiak P, ?li?ewska K. Review: Effects of probiotics, prebiotics, and symbiotics on human health. Nutrients 9 (2017): 1021.
- Sanilier N, Gokcen BB, Sezgin AC. Health benefits of fermented foods. Critical Reviews in Food Science and Nutrition 59 (2019): 506-527.
- Saito H, Miura K. Preparation of transforming deoxyribonucleic acid by phenol treatment. Biochimica et Biophysica Acta 72 (1963): 619-629.
- Yano S, Minato R, Thongsanit J, et al. Overexpression of type I asparaginase of Bacillus subtilis in Escherichia coli, rapid puri?cation and characterization of recombinant type I L-asparaginase. Annals Microbiology 58 (2008): 711-716.
- Lowry OH, Rosebrough NJ, Farr AL, et al. Protein measurement with the Folin phenol reagent. The Journal of Biological Chemistry 193 (1951): 265-275.
- Wang Y, Qian S, Meng G, et al. Cloning and expression of L-asparaginase gene in E. coli. Applied Biochemistry and Biotechnology 95 (2001): 93-101.
- Kotzia GA, Labrou NE. Cloning, expression and characterization of Erwinia carotovora L-asparaginases, Journal of Biotechnology 119 (2005): 309-323.
- Panosyan EH, Seibel NL, Martin AS Children’s Cancer Group Study CCG-1961. Asparaginase antibody and asparaginase activity in children with higher-risk acute lymphoblastic leukemia: Children's Cancer Group Study CCG-1961. Journal of pediatric hematology oncology 26 (2004): 217-226.
- Xu F, Oruna-Concha MJ, Elmore JS. The use of asparaginase to reduce acrylamide levels in cooked food. Food Chemistry 210 (2016): 163-171.
- Andreoletti O, Budka H, Buncic S, et al. Foodborne antimicrobial resistance as a biological hazard. European Food Safety Authority (2008).
- Bourdichon F, Casaregola S, Farrokh C, et al. Food fermentations: Microorganisms with technological beneficial use. International Journal of Food Microbiology 154 (2012): 87-97.
- Cedar H, Schwartz JH. Localization of two L-asparaginases in anaerobically grown Escherichia coli. The Journal of Biological Chemistry 243 (1967): 3753-3755.
- Susan AS, Iyappan S, Vijaya LK, et al. In silico analysis, molecular cloning, expression and characterization of L-asparaginase gene from Lactobacillus reuteri DSM 20016. 3 Biotech 7 (2017): 348.
- Onishi Y, Yano S, Thongsanit J, et al. Expression in Escherichia coli of a gene encoding type II L-asparaginase from Bacillus subtilis, and characterization of its unique properties. Annals of Microbiology 61 (2011): 517-524.
- Huang L, Liu Y, Sun Y, et al. Biochemical characterization of a novel L-asparaginase with low glutaminase activity from Rhizomucor miehei and its application in food safety and leukemia treatment. Applied and Environmental Microbiology 80 (2014): 1561-1569.
- Sindhu R, Manonmani HK. Expression and characterization of recombinant l -asparaginase from Pseudomonas fluorescens. Protein Expression and Purification 143 (2018): 83-91.
- Hong SJ, Lee YH, Khan AR, et al. Cloning, expression, and characterization of thermophilic L-asparaginase from Thermococcus kodakarensis KOD1. Journal of Basic Microbiology 54 (2014): 500-508.
- Husain I, Sharma A, Kumar S, et al. Purification and characterization of glutaminase free asparaginase from Enterobacter cloacae: In-vitro evaluation of cytotoxic potential against human myeloid leukemai HL-60 cells. PLoS ONE (2016).
- Nuzhath F, Mohd MK, Imran AK. L-Asparaginase produced from soil isolates of Pseudomonas aeruginosa shows potent anti-cancer activity on HeLa cells. Saudi Journal of Biological Sciences 26 (2019): 1146-1153.
- Derst C, Henseling J, Rohm KH. Engineering the substrate specificity of Escherichia coli asparaginase II. Selective reduction of glutaminase activity by amino acid replacements at position 248. Protein Science 9 (2000): 2009-2017.
- Bansal S, Gnaneswari D, Mishra P, et al. Structural stability and functional analysis of L-asparaginase from Pyrococcus furiosus. Biochemistry (Mosc) 75 (2010): 375-381.
- Pritsa AA, Kyriakidis DA. L-Asparaginase of Thermus thermophilus: Purificaion, properties and identification of essential amino acids for its catalytic activity. Molecular and Cellular Biochemistry 216 (2001): 93-101.
- Alam S, Pranaw K, Tiwari R, et al. Recent development in the uses of asparaginase as food enzyme. Green Bio-Processes (2018): 55-81.
- Jia M, Xu M, He B, et al. Cloning, expression, and characterization of l-asparaginase from a newly isolated Bacillus subtilis B11-06. Journal of Agricultural and Food Chemistry 61 (2013): 9428-9434.


 Impact Factor: * 5.3
Impact Factor: * 5.3 Acceptance Rate: 75.63%
Acceptance Rate: 75.63%  Time to first decision: 10.4 days
Time to first decision: 10.4 days  Time from article received to acceptance: 2-3 weeks
Time from article received to acceptance: 2-3 weeks 