Expanding the Clinical and Molecular Spectrum of PSMD12-Related Neurodevelopmental Syndrome: An Additional Patient and Review
Pietro Palumbo1#, Orazio Palumbo1#*, Ester Di Muro1,2, Maria Pia Leone1, Stefano Castellana3, Tommaso Biagini3, Tommaso Mazza3, Valentina Leotta4, Massimo Carella1 and Nenad Bukvic4*
1Division of Medical Genetics, Fondazione IRCCS Casa Sollievo della Sofferenza, Viale Padre Pio, 71013 San Giovanni Rotondo (FG), Italy
2Department of Cellular Biotechnologies and Hematology, University of Roma “La Sapienza”, Roma, Italy
3Bioinformatics Unit, Fondazione IRCCS Casa Sollievo della Sofferenza, Viale Padre Pio, 71013 San Giovanni Rotondo (FG), Italy
4Medical Genetics Unit, Azienda Ospedaliero Universitaria Consorziale Policlinico di Bari, Piazza Giulio Cesare 11, 70124 Bari, Italy
#These authors contributed equally to this work
*Corresponding Authors: Orazio Palumbo, Division of Medical Genetics, Fondazione IRCCS Casa Sollievo della Sofferenza, Viale Padre Pio, 71013 San Giovanni Rotondo (FG), Italy
Nenad Bukvic, Medical Genetics Unit, Azienda Ospedaliero Universitaria Consorziale Policlinico di Bari, Piazza Giulio Cesare 11, 70124 Bari, Italy
Received: 18 June 2019; Accepted: 04 July 2019; Published: 11 September 2019
Article Information
Citation: Pietro Palumbo, Orazio Palumbo, Ester Di Muro, Maria Pia Leone, Stefano Castellana, Tommaso Biagini, Tommaso Mazza, Valentina Leotta, Massimo Carella and Nenad Bukvic. Expanding the Clinical and Molecular Spectrum of PSMD12-Related Neurodevelopmental Syndrome: An Additional Patient and Review. Archives of Clinical and Medical Case Reports 3 (2019): 250-260.
View / Download Pdf Share at FacebookAbstract
Degradation of proteins by the ubiquitin-proteasome system (UPS) is an essential biological process that eukaryotic cells use to regulate their functions and coordinate their signaling networks. In particular, it plays a crucial role in neuronal development regulating very important functional and morphological interplays of neurons, such as synaptic plasticity, neurotransmitter release and morphogenesis of axons. Pathogenic alterations of genes involved in the proteolysis by UPS have been associated with several neurodevelopmental disorders (NDDs) and human diseases, highlighting the importance of this regulatory mechanism to developmental processes and neurogenesis. Here, we describe a 19 years old male patient showing a syndromic form of NDD. The main clinical features are intellectual disability/speech delay, congenital anomalies and facial dysmorphisms. Through a targeted resequencing approach (TRS), we identified a missense variant in PSMD12, a gene recently associated to an emerging syndromic form of NDD, which encodes for the non-ATPase subunit of the 19S regulator of 26S proteasome complex. The variant described herein, inherited from the father with apparently borderline cognitive ability, is useful to expand the molecular spectrum of heterozygous PSMD12 mutations and to provide insight into the molecular pathogenesis of this new condition since it is, to the best of our knowledge, the first missense substitution to date reported in medical literature. More importantly, our study highlight once again the utility of next generation sequencing in establishing an etiological basis in clinically and genetically heterogeneous conditions such as NDDs, thus allowing a better diagnosis, counseling and management of affected patients and their families.
Keywords
Ubiquitin-proteasome system, PSMD12, Neurodevelopmental disorders, Next Generation Sequencing, Missense variant
Ubiquitin-proteasome system articles Ubiquitin-proteasome system Research articles Ubiquitin-proteasome system review articles Ubiquitin-proteasome system PubMed articles Ubiquitin-proteasome system PubMed Central articles Ubiquitin-proteasome system 2023 articles Ubiquitin-proteasome system 2024 articles Ubiquitin-proteasome system Scopus articles Ubiquitin-proteasome system impact factor journals Ubiquitin-proteasome system Scopus journals Ubiquitin-proteasome system PubMed journals Ubiquitin-proteasome system medical journals Ubiquitin-proteasome system free journals Ubiquitin-proteasome system best journals Ubiquitin-proteasome system top journals Ubiquitin-proteasome system free medical journals Ubiquitin-proteasome system famous journals Ubiquitin-proteasome system Google Scholar indexed journals PSMD12 articles PSMD12 Research articles PSMD12 review articles PSMD12 PubMed articles PSMD12 PubMed Central articles PSMD12 2023 articles PSMD12 2024 articles PSMD12 Scopus articles PSMD12 impact factor journals PSMD12 Scopus journals PSMD12 PubMed journals PSMD12 medical journals PSMD12 free journals PSMD12 best journals PSMD12 top journals PSMD12 free medical journals PSMD12 famous journals PSMD12 Google Scholar indexed journals Neurodevelopmental disorders articles Neurodevelopmental disorders Research articles Neurodevelopmental disorders review articles Neurodevelopmental disorders PubMed articles Neurodevelopmental disorders PubMed Central articles Neurodevelopmental disorders 2023 articles Neurodevelopmental disorders 2024 articles Neurodevelopmental disorders Scopus articles Neurodevelopmental disorders impact factor journals Neurodevelopmental disorders Scopus journals Neurodevelopmental disorders PubMed journals Neurodevelopmental disorders medical journals Neurodevelopmental disorders free journals Neurodevelopmental disorders best journals Neurodevelopmental disorders top journals Neurodevelopmental disorders free medical journals Neurodevelopmental disorders famous journals Neurodevelopmental disorders Google Scholar indexed journals health articles health Research articles health review articles health PubMed articles health PubMed Central articles health 2023 articles health 2024 articles health Scopus articles health impact factor journals health Scopus journals health PubMed journals health medical journals health free journals health best journals health top journals health free medical journals health famous journals health Google Scholar indexed journals Next Generation Sequencing articles Next Generation Sequencing Research articles Next Generation Sequencing review articles Next Generation Sequencing PubMed articles Next Generation Sequencing PubMed Central articles Next Generation Sequencing 2023 articles Next Generation Sequencing 2024 articles Next Generation Sequencing Scopus articles Next Generation Sequencing impact factor journals Next Generation Sequencing Scopus journals Next Generation Sequencing PubMed journals Next Generation Sequencing medical journals Next Generation Sequencing free journals Next Generation Sequencing best journals Next Generation Sequencing top journals Next Generation Sequencing free medical journals Next Generation Sequencing famous journals Next Generation Sequencing Google Scholar indexed journals treatment articles treatment Research articles treatment review articles treatment PubMed articles treatment PubMed Central articles treatment 2023 articles treatment 2024 articles treatment Scopus articles treatment impact factor journals treatment Scopus journals treatment PubMed journals treatment medical journals treatment free journals treatment best journals treatment top journals treatment free medical journals treatment famous journals treatment Google Scholar indexed journals Chromosomal Microarray Analysis articles Chromosomal Microarray Analysis Research articles Chromosomal Microarray Analysis review articles Chromosomal Microarray Analysis PubMed articles Chromosomal Microarray Analysis PubMed Central articles Chromosomal Microarray Analysis 2023 articles Chromosomal Microarray Analysis 2024 articles Chromosomal Microarray Analysis Scopus articles Chromosomal Microarray Analysis impact factor journals Chromosomal Microarray Analysis Scopus journals Chromosomal Microarray Analysis PubMed journals Chromosomal Microarray Analysis medical journals Chromosomal Microarray Analysis free journals Chromosomal Microarray Analysis best journals Chromosomal Microarray Analysis top journals Chromosomal Microarray Analysis free medical journals Chromosomal Microarray Analysis famous journals Chromosomal Microarray Analysis Google Scholar indexed journals patient articles patient Research articles patient review articles patient PubMed articles patient PubMed Central articles patient 2023 articles patient 2024 articles patient Scopus articles patient impact factor journals patient Scopus journals patient PubMed journals patient medical journals patient free journals patient best journals patient top journals patient free medical journals patient famous journals patient Google Scholar indexed journals disease articles disease Research articles disease review articles disease PubMed articles disease PubMed Central articles disease 2023 articles disease 2024 articles disease Scopus articles disease impact factor journals disease Scopus journals disease PubMed journals disease medical journals disease free journals disease best journals disease top journals disease free medical journals disease famous journals disease Google Scholar indexed journals phenotype articles phenotype Research articles phenotype review articles phenotype PubMed articles phenotype PubMed Central articles phenotype 2023 articles phenotype 2024 articles phenotype Scopus articles phenotype impact factor journals phenotype Scopus journals phenotype PubMed journals phenotype medical journals phenotype free journals phenotype best journals phenotype top journals phenotype free medical journals phenotype famous journals phenotype Google Scholar indexed journals
Article Details
1. Introduction
Neurodevelopmental disorders (NDDs) are a group of clinically and genetically heterogeneous disorders, diagnosed during early childhood, including intellectual disability (ID), autism spectrum disorders (ASD), communication and learning disorders, attention deficit/hyperactivity disorders (ADHD) and motor disorders. They have an estimated prevalence of approximately 1% and represent one of the major challenges in medicine being the most frequent causes of disability in children (nearly 3 of 100 babies are affected) and the main reason for referral in clinical genetic centers. Furthermore, incomplete penetrance and variable expressivity can influence their inheritance, and thus represents a challenge for genetic counselling [1]. In the last fifteen years [2], the identification of underlying genetic defects and risk factors for NDD have increased significantly due to the development of new diagnostic technologies like Next Generation Sequencing (NGS) and Chromosomal Microarray Analysis (CMA). This allowed many affected patients, who had previously undergone many invasive, expensive, and uninformative tests, to receive a definitive genetic diagnosis. Up to date, mutations at over 1000 loci have been associated with NDDs [3, 4]. Among them, the activity impairments of genes involved in the proteolysis by the ubiquitin-proteasome system (UPS) has been demonstrated for some distinctive clinical entities such as Angelman Syndrome (UBE3A, MIM:601623) [5], Kaufman oculocerebrofacial syndrome (UBE3B, MIM:608047) [6], X-linked Turner-type syndromic mental retardation (HUWE1, MIM:300697) [7], chromosome 16p13.2 deletion syndrome (USP7, MIM: 602519) [8], mental retardation, X-linked 99 and mental retardation, X-linked 99, syndromic female-restricted (USP9X, MIM: 30072) [9]. More recently another gene, PSMD12 (Proteosome 26S Subunit Non-ATPase, 12; OMIM 604450), encoding for a non-ATPase subunit of the 19S regulator of the 26S proteasome complex, has been implicated in the etiology of an emerging syndromic form of NDD characterized by ID, motor and speech delay, abnormal behavior, hypotonia, craniofacial, cardiac, renal, skeletal and genital abnormalities. Other clinical features, less frequently reported, are seizures and brain structural abnormalities [10, 11].
Herein, we report on the first patient with a missense variant in PSMD12, who showed ID, speech delay, abnormal behavior, and dysmorphic features. Modeling its impact on the three-dimensional structure demonstrate disturbance on the protein stability. We compared the clinical features of our patient to the previous reports founding strong clinical correlation in the phenotype.
2. Case Presentation
The patients was a 19-year-old male referred for genetic counseling due to ID, congenital malformation, facial dysmorphisms with delayed speech and language development. The proband was the second male child of apparently healthy, non-consanguineous Italian couple with not known family history of neurological abnormalities, recurrent miscarriages, other birth defects and/or genetic illnesses. The proband was delivered at term (38 weeks of gestation) by programed cesarean section. The pregnancy was characterized by contractions at 7th month of gestation, otherwise passed uneventful. The birth weight of 3,800 kg (90th centile), length of 50 cm (75th centile), and head circumference of 32 cm (5 -10th centile). APGAR scores were 9/10 at 1st and 5th min, respectively. Important data from anamnestic history: at age of 5 years the patient had chirurgical intervention for hypospadias; at age of 12 years the patient had dental chirurgical intervention for multiple unerupted teeth with unusual position (12 palatal teeth). Furthermore, during dentist controls misalignment of teeth and gemination were observed; treatment for scoliosis and flatfoot have been undertaken during development; ophthalmological correction for strabismus has been done; presence of multiple nevus on whole body is under dermatological control and regular follow up; no treatment for short stature. He also had a history of development delay (the first steps – walking at 18 months; first words at 24 months), learning problems and still requires learning support at school. The proband was evaluated in our medical genetics unit during the first visit at the age of 19 years. On examination his height was 185 cm (75-90th centile), weight was 65 Kg (25-50th centile) and occipital frontal circumference (OFC) was 57 cm (90th centile). A distinctive facial appearance was noted with elongated and hatchet face, bitemporal hollowing, thick eyebrow with synophrys, down slanting palpebral fissures and eyelids ptosis, long and prominent nasal bridge, low-set and posteriorly rotated ears, high arched palate, triangular chin with micrognathia, webbed neck, low hairline at the back of the neck (Figure 1). In addition, the patient showed pectus excavatum, prominent scoliosis, relative joint hyper extensibility and bilateral pes planus.
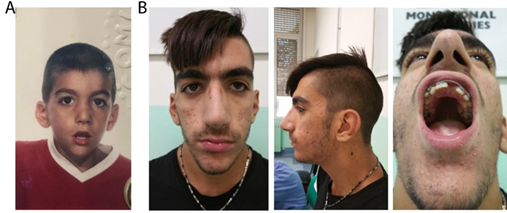
Figure 1: Patient with PSMD12 p.(Arg289Pro) at 6 years (A) and at 19 years of age (B). Detailed clinical description is reported in the text.
The proband had no reported seizures. In medical documentation, presented during genetic counseling, the results of routine hemato-chemical laboratory examination (hepatic, renal, hormonal, martial) were normal. ORL examination within normal limits. Ophthalmological examination: OO +1.75 sf. Strabismus. Normal male karyotype (46, XY) was observed; high resolution SNP array analysis, performed by using CytoScan HD Array (Affymetrix, Santa Clara, CA) as previously described [12] did not identify any pathogenic copy number variations (microdeletions or microduplications) in the patient; 5’UTR FMR1 resulted in normal allele (31 ± 1CGG ripetitions). On the basis of genetic counseling, objective examination, taking in consideration all possibilities of differential diagnosis and clinicians suspect was decided to perform further genetic tests by implementation of NDD gene panel, after pre-test counseling and obtaining of written informed consent.
3. Materials and Methods
A NDDs gene panel, containing 95 genes related to both syndromic and non-syndromic forms, was defined according to data obtained from scientific literature (Supplementary material). Probes were designed by using Agilent SureDesign Custom design tool (https://earray.chem.agilent.com/suredesign/): the regions of interest of this panel included all exons plus 25 bp flanking intron regions. The total amplicons number were 18.137 and the target size was 360.002 Kbp with a theoretical coverage of 99.87% spanning our targeted regions. Peripheral blood samples were taken from each patient and their parents and genomic DNA was isolated using Bio Robot EZ1 (Quiagen, Solna, Sweden). The quality of DNA was tested on 1% electrophorese agarose gel and the concentration was quantified with Nanodrop 2000 C spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA). A library of all coding regions of 95 genes was obtained using the Haloplex target enrichment kit (Agilent Technologies, Santa Clara, CA, USA) according to the manufacturer’s instructions. The libraries were pooled, and NGS sequencing was performed on a MiSeq sequencer (Illumina, San Diego, CA, USA) using a MiSeq Reagent kit V3 300 cycles flow cell. Variants were called by means of the HaplotypeCaller tool of GATK ver. 3.4 [13] and were annotated with ANNOVAR, using RefSeq gene and transcript annotations (updated to July 2015) [14]. Variants were sought in the most important public collections of variants, such as dbSNP ver. 152 [15], ExAC ver. 0.3 (http://exac.broadinstitute.org) [16], Exome Variant Server (http://evs.gs.washington.edu/EVS, accessed at December 2016), HRC [17], Kaviar [18] and ClinVar [19]. Missense variants were further annotated by dbNSFP ver. 3.2 from which we retrieved pre-computed pathogenicity predictions and evolutionary conservation measures [20].
The stability of the PSMD12 protein upon mutation was investigated thermodynamically through the FoldX algorithm (see Results). This study was conducted on the wild-type protein model obtained from the Protein Data Bank [21]. It was mutated in-silico through UCSC Chimera [22], yielding a second protein model. FoldX computed the total energy of PSMD12, wild-type and mutated, as a proxy of their overall stability and the Van der Waals interresidue clashes, as energy penalization factors. Thus, both pdb structures were minimized, before assessing their stability, namely all the side chains were slightly moved in order to reduce the Van der Waals’ clashes, before being analyzed by FoldX. The standalone version of FoldX is downloadable from http://foldx.crg.es. It was run with standard parameters and used to compute the total energy values of the wild-type and mutated models of PSMD12. These values were used to predict the overall protein stability, with and without mutation.
4. Results
Next generation sequencing revealed a missense variant in heterozygous state in the exon 8 of the PSMD12 gene (OMIM 604427) (PSMD12:NM_002816) c.866G>C resulting in a p.(Arg289Pro) substitution (GRCh37/hg19). The variant was detected with a depth of coverage greater than 300x, and with elevate quality scores (i.e., Phread quality > 3,000 and genotype quality = 99). This amino acid substitution was known but very rare (dbSNP ID: rs779733116; 8.2565e-6 estimated frequency by the ExAC database and 4.061e-06 estimated by gnomAD), affects a highly conserved residue (GERP++_RS score = 5.09; phyloP100way_vertebrate score = 7.523; phyloP100way_mammalian score = 0.892; phast-Cons100way_vertebrate score = 1.000; SiPhy_29way_logOdds = 18.496), and it is predicted as deleterious or probably damaging by several software tools, including SIFT (score = 0.001), PolyPhen2 (score = 1.0), LRT (score = 0), MutationTaster (score = 1), Provean (score = -6.14), CADD (score = 34), and M-CAP (score = 0.090) respectively. The variant was confirmed by Sanger sequencing, and parental DNA analysis showed its paternal inheritance (Figure 2).
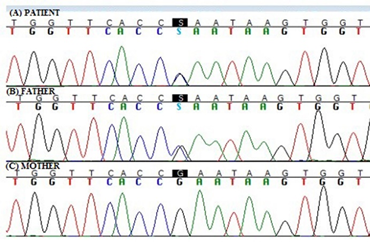
Figure 2: Sanger sequencing showing the c.866G>C nucleotide change in PSMD12. (A) Electropherogram of the patient. (B) Electropherogram of the father. (C) Electropherogram of the mother.
Furthermore, the free energy calculations for this variant were ΔGmut = 231.2182 kcal/mol and ΔGwt = 229.9657 kcal/mol, from which ΔΔG = ΔGmut - ΔGwt = +1.39 kcal/mol. The difference in free energy resulted to be positive and in the range to classify the p.(Arg289Pro) PSMD12 variant as destabilizing; the localization of the variant is depicted in Figure 3.

Figure 3: Location of the Arg289Pro substitution in the PSMD12 protein / Ribbon diagram of PSMD12, with Arg289Pro colored in red.
The variant has been submitted to LOVD (https://databases.lovd.nl/shared/variants/0000478178; patient ID #00234320).
5. Discussion
In this study, we describe a patient carrier of a missense mutation, the first to date reported in medical literature, of PSMD12, a gene involved in the degradation of proteins by the ubiquitin-proteasome system and recently associated to NDDs. The variant c.866G>C was predicted to be likely pathogenic [20] and involve the high conserved residue p.(Arg289Pro). To date, there have been two separate publications describing postnatal cases with PSMD12 pathogenic variants (point mutations or copy number variations). In Table 1 are summarized and compared the clinical and molecular findings on patients reported to date, including our, with a total of twenty affected patients.
|
|
Present case |
Khalil et al., 2018 |
Küry et al., 2017 |
|
PSMD12 Variant Type
|
c.866G>Cp.(Arg289Pro) (missense) |
9 SNVs (nonsense) |
4 SNVs (nonsense) 6 CNVs (microdeletions) |
|
Gender |
M |
6 F/3 M |
4 F/6 M |
|
Age of last evaluation |
19 years |
6 to 42 years |
21 months to 19 years |
|
Weight at birth (g) |
3,800 |
ND |
1,570 to 3,800 |
|
Length at birth (cm) |
50 |
ND |
43,5 to 50 |
|
OFC at birth (cm) |
32 |
ND |
30 to 34,9 |
|
Developmental delay/Intellectual disability |
+ |
8/9 (1 ND) |
10/10 |
|
Speech delay |
+ |
5/9 (3 ND) |
9/10 |
|
Abnormal behavior |
+ |
5/9 (1 ND) |
7/10 (2 ND) |
|
Motor delay |
- |
0/9 |
7/10 |
|
Seizures |
- |
0/9 |
3/10 |
|
Hypotonia |
- |
1/9 (8 ND) |
6/10 |
|
Brain abnormalities |
- |
1/9 (8 ND) |
2/10 (3 ND) |
|
Dysmorphic features |
+ |
4/9 (4 ND) |
9/10 (1 ND) |
|
Ophthalmological abnormalities |
+ |
2/9 (4 ND) |
8/10 |
|
Genital abnormalities |
+ |
0/9 (4 ND) |
6/10 |
|
Skeletal abnormalities |
+ |
1/9 (4 ND) |
5/10 |
|
Other abnormalities
|
Multiple (12) unerupted theeth Short stature |
IUGR and premature delivery (1/9) |
Cardiac Abnormalities: 5/10 Renal Abnormalities: 6/10 (1 ND) |
Abbreviations: M = male; F = female; ND = non determined; IUGR = intrauterine growth retardation; SNV = single nucleotide variations; CNV = copy number variations; + = present; - = absent.
Table 1: Clinical characteristics of individuals with PSMD12 variants.
The first study which provide a review of copy number variations (CNVs), nonsense variants and splice variant of PSMD12, was performed by Küry and colleagues [10] that reported clinical and molecular data of ten patients (six males and four females) affected by NDDs. They described three patients carriers of nonsense variant [c.367C>T p.(Arg123*), c.1274T>G p.(Leu425*), c.601C>T p.(Arg201*)] one patient carrier of a splice variant [c.909-2A>G, p.(?)] and six patient carriers of microdeletions in the 17q23.3q24.2 chromosome region, ranging in size from 0.62 Mb to 4.06 Mb, encompassing the PSMD12 gene. The only clinical trait shared among all patients was developmental delay (DD)/ID, while speech delay was present in 9/10 patient, motor delay and abnormal behavior was present in 7/10 patient.
More recently, Khalil and colleagues reported further nine patients (six females and three males) affected by NDDs carriers of nonsense mutations in PSMD12 [11]. They described two non-consanguineous families, one with eight affected individuals carrying the same variant presented in the affected father, and one with a single patient with a de novo mutation. In this cohort, from a molecular point of view, all patients were carriers of nonsense mutations in PSMD12: the p.(Arg123*) in 8/9 cases and the p.(Arg210*) in the remaining one. From a clinical point of view, in this study the clinical trait more frequently observed was developmental delay or the intellectual disability, present in 8/9 patient (in one case was not determined), while speech delay was present in 5/9 patients (in three cases was not determined), and abnormal behavior was present in 5/9 patient. As for the cases reported by Küry and colleagues, about other clinical traits, there is some variability among patients, even if they have the same genetic anomaly. This finding is quite common in NDDs and could be explained by variable expressivity and/or incomplete penetrance or differential effects on the transcript by different molecular events such as deletions and/or point mutations. Taken together, analysis of the data shows that consistent findings are DD/ID (19/20, 95%), speech delay (15/20, 75%), behavioral problems (13/20, 65%). Additional features also noted amongst the patients included ophthalmological problems (11/20, 55%), motor delay (7/20, 35%), hypotonia (7/20, 35%), genital and skeletal anomalies (7/20, 35%), functional and structural renal alterations (6/20, 30%), cardiac anomalies (5/20, 25%). Although not as common, other clinical features reported are seizure (3/20, 15%) and brain structural anomalies (3/20, 15%).
Interesting to note, all patients to date described in medical literature have been identified by whole exome sequencing or whole genome array analysis, suggesting that the phenotype associated with PSMD12 point mutations or microdeletions is not yet easily recognizable. In fact, although facial dysmorphisms were observed in 14/20 (70%), they were variable, tipically very mild or not described in detail. Our patient is the one with the most detailed dysmorphic characterization (elongated and hatchet face, bitemporal hallowing, thick eyebrow with synophrys, down slanting palpebral fissures and eyelids ptosis, long and prominent nasal bridge, low-set and posteriorly rotated ears, high arched palate, triangular chin with micrognathia, webbed neck, low hairline at the back of the neck) (Figure 1), and for this reason useful to start defining a typical facial gestalt that addresses the diagnosis. Obviously, additional cases are needed to expand the spectrum of clinical features associated with this emerging syndrome and define the core phenotype that make it recognizable or suspected.
Our patient inherited the genetic variation from the father, who did not underwent to date psychological evaluation of cognitive capability (recommended during genetic counseling). However, the variable expressivity of some genetic alteration is an evidence, not uncommon in NDDs, which represents a great challenge in genetic counselling. As an example, microdeletions or point mutations of NRXN1, a gene responsible of ASD, ADHD, ID, seizure, schizophrenia can be inherited form apparently healthy parents [23, 24]. It is possible that other polygenic, epigenetic or multifactorial effects such as synergic heterozygosity, modifier gene effect, could play a role in the penetrance and expression of some genetic variants. Obviously, although the variant affects a highly conserved residue and its impact is predicted to be deleterious or likely deleterious by most of the predictors consulted and by in silico modeling, functional studies are needed to confirm these emerging bioinformatics evidences.
The haploinsufficiency of PSMD12 is the mechanism proposed by Küry et al. [10], and Khalil et al. [11] as causative of clinical phenotype observed in their patients. The stop mutation or a whole gene deletion may certainly cause the haploinsufficiency of the gene, and this could be the reason why this kind of alteration arise as de novo. A missense mutation may have a variable effect on phenotype, and this kind of genetic variation of PSMD12 may results more susceptible to genetic interactions listed previously (i.e. synergic heterozygosity, modifier gene effect), generating the variability in clinical manifestations.
Obviously, being our patient the first to date reported with a missense mutation, further clinical and functional evidences are needed to corroborate this hypothesis, expanding the knowledge regarding the clinical effect of the different molecular alterations of PSMD12. At this moment the above reported observations are the only one possible without further functional investigations and clinical reports of affected patients with missense mutations in PSMD12 gene.
6. Conclusion
In conclusion, we described the first missense mutation of PSMD12, detected in a patient affected by an emerging syndromic form of NDD. Our finding confirm the recent disease association of PSMD12, expand the molecular spectrum of heterozygous PSMD12 mutations and provide insight into molecular pathogenesis of this new condition. Ascertainment of more cases with PSMD12 mutations is required to further delineate the phenotypic spectrum of PSMD12-related neurodevelopmental disorder.
Acknowledgments
We thank the family for their cooperation. This study was supported by a grant of the Italian Ministry of Health (Ricerca Corrente 2019) to Massimo Carella.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Author Contributions
PP, OP, MC and NB conceived the study, designed experiments and wrote the manuscript. NB collected the patients’ clinical information. EDM, MPL and VL performed the molecular analysis. SC, TB and TM performed the analysis of TRS data and modeling the impact of the reported variant on the three-dimensional structure of the protein. PP, OP, MC and NB interpreted the results. PP and OP contributed equally to this work. OP and NB are the corresponding authors. All authors reviewed and approved the final manuscript.
Consent for Pubblication
This study was carried out following the recommendations of the Declaration of Helsinki and the Italian law for biomedical experimentation with written informed consent from all subjects. The study was approved by the local Ethics committee of the Fondazione IRCCS Casa Sollievo della Sofferenza, San Giovanni Rotondo (FG).
References
- Mitchell KJ. The genetics of neurodevelopmental disease. Curr Opin Neurobiol 21 (2011): Curr Opin Neurobiol 21 (2011): 197-203.
- Vissers LE, Gilissen C, Veltman JA. Genetic studies in intellectual disability and related disorders. Nat Rev Genet 17 (2016): 9-18.
- Rosenfeld JA, Patel A. Chromosomal Microarrays: Understanding Genetics of Neurodevelopmental Disorders and Congenital Anomalies. J Pediatr Genet 6 (2017): 42-50.
- T?rlungeanu DC, Novarino G. Genomics in neurodevelopmental disorders: an avenue to personalized medicine. Exp Mol Med 50 (2018): 100.
- Kishino T, Lalande M, Wagstaff J. UBE3A/E6-AP mutations cause Angelman syndrome. Nat Genet 15 (1997): 70–73.
- Basel-Vanagaite L, Dallapiccola B, Ramirez-Solis R, et al. Deficiency for the ubiquitin ligase UBE3B in a blepharophimosis-ptosis-intellectual-disability syndrome. Am J Hum Genet 91 (2012): 998-1010.
- Froyen G, Corbett M, Vandewalle J, et al. Submicroscopic duplications of the hydroxysteroid dehydrogenase HSD17B10 and the E3 ubiquitin ligase HUWE1 are associated with mental retardation. Am J Hum Genet 82 (2008): 432–443.
- Hao YH, Fountain MD Jr, Fon Tacer,K, et al. USP7 Acts as a Molecular Rheostat to Promote WASH-Dependent Endosomal Protein Recycling and Is Mutated in a Human Neurodevelopmental Disorder. Mol Cell 59 (2015): 956-969.
- Homan CC, Kumar R, Nguyen LS, et al. Mutations in USP9X are associated with X-linked intellectual disability and disrupt neuronal cell migration and growth. Am J Hum Genet. 94 (2014): 470–478.
- Küry S, Besnard T, Ebstein F, et al. De Novo Disruption of the Proteasome Regulatory Subunit PSMD12 Causes a Syndromic Neurodevelopmental Disorder. Am J Hum Genet 100 (2017): 352-363.
- Khalil R, Kenny C, Hill RS, et al. PSMD12 haploinsufficiency in a neurodevelopmental disorder with autistic features. Am J Med Genet B Neuropsychiatr Genet 177 (2018): 736-745.
- Palumbo P, Accadia M, Leone MP, et al. Clinical and molecular characterization of an emerging chromosome 22q13.31 microdeletion syndrome. Am J Med Genet A 176 (2018): 391-398.
- McKenna A, Hanna M, Banks E, et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20 (2010): 1297-1230.
- Wang K, Li M, Hakonarson H. ANNOVAR: Functional annotation of genetic variants from next-generation sequencing data. Nucleic Acids Res 38 (2010): e164.
- Sherry ST, Ward MH, Kholodov M, et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res 29 (2001): 308-311.
- Lek M, Karczewski KJ, Minikel EV, et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536 (2016): 285-291.
- McCarthy S, Das S, Kretzschmar W, et al. Haplotype Reference Consortium. A reference panel of 64,976 haplotypes for genotype imputation. Nat Genet 48 (2016): 1279-1283.
- Glusman G, Caballero J, Mauldin DE, et al. KAVIAR: an accessible system for testing SNV novelty. Bioinformatics 27 (2011): 3216-3217.
- Landrum MJ, Lee JM, Riley GR, et al. ClinVar: public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res 42 (2014): D980-985.
- Liu X, Wu C, Li C, et al. dbNSFP v3.0: A One-Stop Database of Functional Predictions and Annotations for Human Non-synonymous and Splice Site SNVs. Human Mutation 37 (2016): 235-241.
- Berman HM, Battistuz T, Bhat TN, et al. The Protein Data Bank. Acta Crystallogr D58 (2002): 899-907.
- Pettersen EF, Goddard TD, Huang CC, et al. UCSF Chimera, A Visualization System for Exploratory Research and Analysis. J Comput Chem 25 (2004): 1605-1612.
- Viñas-Jornet M, Esteba-Castillo S, Gabau E, et al. A common cognitive, psychiatric, and dysmorphic phenotype in carriers of NRXN1 deletion. Mol Genet Genomic Med 2 (2014): 512-521.
- Dabell MP, Rosenfeld JA, Bader P, et al. Investigation of NRXN1 deletions: clinical and molecular characterization. Am J Med Genet A 161A (2013): 717-731.


 Impact Factor: * 5.31
Impact Factor: * 5.31 Acceptance Rate: 75.63%
Acceptance Rate: 75.63%  Time to first decision: 10.4 days
Time to first decision: 10.4 days  Time from article received to acceptance: 2-3 weeks
Time from article received to acceptance: 2-3 weeks 
